Abstract
The inability to digest lactose, due to lactase nonpersistence, is a common trait in adult mammals, except in certain human populations that exhibit lactase persistence. It is not known how the lactase gene is dramatically downregulated with age in most individuals but remains active in some individuals. We performed a comprehensive epigenetic study of human and mouse small intestines, by using chromosome-wide DNA-modification profiling and targeted bisulfite sequencing. Epigenetically controlled regulatory elements accounted for the differences in lactase mRNA levels among individuals, intestinal cell types and species. We confirmed the importance of these regulatory elements in modulating lactase mRNA levels by using CRISPR–Cas9-induced deletions. Genetic factors contribute to epigenetic changes occurring with age at the regulatory elements, because lactase-persistence and lactase-nonpersistence DNA haplotypes demonstrated markedly different epigenetic aging. Thus, genetic factors enable a gradual accumulation of epigenetic changes with age, thereby influencing phenotypic outcome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Rasinperä, H. et al. Transcriptional downregulation of the lactase (LCT) gene during childhood. Gut 54, 1660–1661 (2005).
Sahi, T., Launiala, K. & Laitinen, H. Hypolactasia in a fixed cohort of young Finnish adults: a follow-up study. Scand. J. Gastroenterol. 18, 865–870 (1983).
Ingram, C.J., Mulcare, C.A., Itan, Y., Thomas, M.G. & Swallow, D.M. Lactose digestion and the evolutionary genetics of lactase persistence. Hum. Genet. 124, 579–591 (2009).
Enattah, N.S. et al. Identification of a variant associated with adult-type hypolactasia. Nat. Genet. 30, 233–237 (2002).
Poulter, M. et al. The causal element for the lactase persistence/non-persistence polymorphism is located in a 1 Mb region of linkage disequilibrium in Europeans. Ann. Hum. Genet. 67, 298–311 (2003).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Enattah, N.S. et al. Evidence of still-ongoing convergence evolution of the lactase persistence T-13910 alleles in humans. Am. J. Hum. Genet. 81, 615–625 (2007).
Itan, Y., Powell, A., Beaumont, M.A., Burger, J. & Thomas, M.G. The origins of lactase persistence in Europe. PLoS Comput. Biol. 5, e1000491 (2009).
Gamba, C. et al. Genome flux and stasis in a five millennium transect of European prehistory. Nat. Commun. 5, 5257 (2014).
Ranciaro, A. et al. Genetic origins of lactase persistence and the spread of pastoralism in Africa. Am. J. Hum. Genet. 94, 496–510 (2014).
Tishkoff, S.A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nat. Genet. 39, 31–40 (2007).
Jones, B.L. et al. Diversity of lactase persistence alleles in Ethiopia: signature of a soft selective sweep. Am. J. Hum. Genet. 93, 538–544 (2013).
Troelsen, J.T., Olsen, J., Møller, J. & Sjöström, H. An upstream polymorphism associated with lactase persistence has increased enhancer activity. Gastroenterology 125, 1686–1694 (2003).
Olds, L.C. & Sibley, E. Lactase persistence DNA variant enhances lactase promoter activity in vitro: functional role as a cis regulatory element. Hum. Mol. Genet. 12, 2333–2340 (2003).
Jensen, T.G. et al. The -14010*C variant associated with lactase persistence is located between an Oct-1 and HNF1α binding site and increases lactase promoter activity. Hum. Genet. 130, 483–493 (2011).
Fang, L., Ahn, J.K., Wodziak, D. & Sibley, E. The human lactase persistence-associated SNP -13910*T enables in vivo functional persistence of lactase promoter-reporter transgene expression. Hum. Genet. 131, 1153–1159 (2012).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Elliott, G. et al. Intermediate DNA methylation is a conserved signature of genome regulation. Nat. Commun. 6, 6363 (2015).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Bell, J.T. et al. Epigenome-wide scans identify differentially methylated regions for age and age-related phenotypes in a healthy ageing population. PLoS Genet. 8, e1002629 (2012).
De Jager, P.L. et al. Alzheimer's disease: early alterations in brain DNA methylation at ANK1, BIN1, RHBDF2 and other loci. Nat. Neurosci. 17, 1156–1163 (2014).
Gjoneska, E. et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer's disease. Nature 518, 365–369 (2015).
Weiser, M.M. Intestinal epithelial cell surface membrane glycoprotein synthesis. I. An indicator of cellular differentiation. J. Biol. Chem. 248, 2536–2541 (1973).
Kriukiene, E. et al. DNA unmethylome profiling by covalent capture of CpG sites. Nat. Commun. 4, 2190 (2013).
Fang, R., Olds, L.C. & Sibley, E. Spatio-temporal patterns of intestine-specific transcription factor expression during postnatal mouse gut development. Gene Expr. Patterns 6, 426–432 (2006).
Diep, D. et al. Library-free methylation sequencing with bisulfite padlock probes. Nat. Methods 9, 270–272 (2012).
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Gallego Romero, I. et al. Herders of Indian and European cattle share their predominant allele for lactase persistence. Mol. Biol. Evol. 29, 249–260 (2012).
Li, Y. et al. Genome-wide DNA methylome analysis reveals epigenetically dysregulated non-coding RNAs in human breast cancer. Sci. Rep. 5, 8790 (2015).
Di Ruscio, A. et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature 503, 371–376 (2013).
Magistri, M., Faghihi, M.A., St Laurent, G. III & Wahlestedt, C. Regulation of chromatin structure by long noncoding RNAs: focus on natural antisense transcripts. Trends Genet. 28, 389–396 (2012).
Yu, D.H. et al. Developmentally programmed 3′ CpG island methylation confers tissue- and cell-type-specific transcriptional activation. Mol. Cell. Biol. 33, 1845–1858 (2013).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Ong, C.T. & Corces, V.G. CTCF: an architectural protein bridging genome topology and function. Nat. Rev. Genet. 15, 234–246 (2014).
Sheaffer, K.L. et al. DNA methylation is required for the control of stem cell differentiation in the small intestine. Genes Dev. 28, 652–664 (2014).
Kaaij, L.T. et al. DNA methylation dynamics during intestinal stem cell differentiation reveals enhancers driving gene expression in the villus. Genome Biol. 14, R50 (2013).
Lewinsky, R.H. et al. T-13910 DNA variant associated with lactase persistence interacts with Oct-1 and stimulates lactase promoter activity in vitro . Hum. Mol. Genet. 14, 3945–3953 (2005).
Domcke, S. et al. Competition between DNA methylation and transcription factors determines binding of NRF1. Nature 528, 575–579 (2015).
Xiang, J.F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Hilton, I.B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Delaney, L. & Smith, J.P. Childhood health: trends and consequences over the life course. Future Child. 22, 43–63 (2012).
Belsky, D.W. et al. Polygenic risk and the development and course of asthma: an analysis of data from a four-decade longitudinal study. Lancet Respir. Med. 1, 453–461 (2013).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Platt, R.J. et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Yeh, K.Y., Yeh, M. & Holt, P.R. Intestinal lactase expression and epithelial cell transit in hormone-treated suckling rats. Am. J. Physiol. 260, G379–G384 (1991).
Goda, T., Yasutake, H., Tanaka, T. & Takase, S. Lactase-phlorizin hydrolase and sucrase-isomaltase genes are expressed differently along the villus-crypt axis of rat jejunum. J. Nutr. 129, 1107–1113 (1999).
Royce, T.E., Carriero, N.J. & Gerstein, M.B. An efficient pseudomedian filter for tiling microrrays. BMC Bioinformatics 8, 186 (2007).
Krueger, F. & Andrews, S.R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Garber, M. et al. Identifying novel constrained elements by exploiting biased substitution patterns. Bioinformatics 25, i54–i62 (2009).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
We thank A. Turinsky and L. Strug for statistical recommendations; T. Karvelis for help with sgRNA construction; M. Susic for technical assistance; A. Patterson, P. Kapranov and D. Swallow for comments on the manuscript; and F. Zhang (Broad Institute of MIT and Harvard University) for pX330 plasmid. This work was supported in part by the Canadian Institutes of Health Research (MOP-199170, MOP-119451 and MOP-77689), the US National Institutes of Health (MH088413 and DK085698), the Krembil Foundation and Brain Canada, to A.P. This work was also supported by the Canadian Centre for Computational Genomics (C3G), part of the Genome Innovation Network (GIN), funded by Genome Canada through Genome Quebec and Ontario Genomics, to M.B. A.P. is supported as the Tapscott Chair in Schizophrenia Studies, University of Toronto, Canada. E.K. was supported by a grant from the Research Council of Lithuania (MIP-045/2013). J.G. and K.K. were funded by a grant (MIP-14032) from the Research Council of Lithuania. V.L. was supported by a Canadian Institutes of Health Research Fellowship (200910MFE-211514-141430).
Author information
Authors and Affiliations
Contributions
The study was designed by V.L. and A.P., and directed and coordinated by V.L., M.B. and A.P. V.L., E.O., R.J. and C.P. planned and performed the experimental work. O.J.B. coordinated and performed the computational analysis. J.G. and K.K. contributed to the computational analysis of the microarrays. A.M., R.P., A.Ž., K.A. and L.K. collected the human jejunum surgical samples and other human tissues. G.G. and V.Š. prepared the CRISPR–Cas9n cell-line constructs, and R.J. and A.N. contributed to the CRISPR–Cas9n cell-line work. K.K. was involved in the haplotype-dependent epigenetic aging analysis. E.K. consulted on the mTAG approach. S.E., A.Z. and G.O. were involved in bisulfite padlock-probe design and preparation. The manuscript was written by V.L., O.J.B. and A.P., and was commented on by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Investigation of lactase-gene regulation in the human jejunum.
(a) Boxplots showing the age distribution of the human tissue samples. We attained jejunum surgical samples from 56 (cohort 1) and 59 (cohort 2) unrelated Caucasian individuals. We also attained 58 blood samples and 18 sperm samples from separate, unrelated Caucasian individuals. Error bars represent s.e.m. (b) A chelating method specifically removed enterocytes (epithelial cells) from intestinal samples. We optimized the chelating method to remove enterocytes specifically from the villi, where LCT expression is most abundant. Left panel: Jejunum tissue prior to the chelating method (H&E stain, 40X magnification). Arrows indicate examples of enterocytes. Middle panels: Enterocyte cell fraction isolated by chelating method (40X and 60X magnification). Right panel: Jejunum tissue after the chelating method shows that enterocytes primarily from villi have been isolated (40X magnification). (c) Steady-state LCT mRNA levels in the two cohorts investigated. A wide range of LCT mRNA levels was represented in both cohorts. Cohort 1 n = 56 individuals and cohort 2 n = 59 individuals. Data are means ± s.e.m.
Supplementary Figure 2 Chromosome-wide profiling of DNA modifications on chromosomes 9 and 19 associated with LCT mRNA levels.
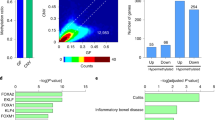
Tiling microarrays investigated DNA modifications in human enterocytes isolated from the jejunum (n = 56 individuals). Manhattan plots demonstrating that DNA modification within chromosomes that do not contain the LCT gene (chromosome 9 and 19; GRCh37/hg19) is not significantly associated with LCT mRNA (all sites FDR q ≥ 0.05). SLP refers to signed log10(p-value).
Supplementary Figure 3 Chromosome-wide analysis using human tiling microarrays to examine DNA modifications correlated with LCT mRNA levels in enterocytes and jejunum lacking enterocytes.
The Parsimonious Temporal Aggregation (PTA) algorithm was used to merge similar adjacent probes and assayed resulting regions with the highest (10%) inter-sample variability on chromosome 2 (GRCh37/hg19). Both enterocytes (in red) and the enterocyte-deficient jejunum (in blue) were examined for each individual (n = 56 individuals). (a) On chromosome 2 a significant positive association was found between density of unmodified cytosines in LCT exon 1–intron 1 and LCT mRNA levels in enterocytes (indicated by arrow; R = 0.65, chr2:136,593,890-136,594,100). The horizontal blue line shows the significance threshold. DNA modifications in the enterocyte-deficient jejunum were not significantly associated with LCT mRNA levels on chromosome 2. (b) Close-up of the region where lower DNA modification densities were correlated with increased LCT mRNA levels in enterocytes (highlighted by horizontal red block). DNA modifications did not significantly correlate with LCT mRNA levels in the enterocyte-deficient jejunum. The vertical dashed red line shows the transcription start of LCT.
Supplementary Figure 4 Epigenetically controlled regions associated with the age-dependent downregulation of Lct in mice.
(a) Steady-state Lct mRNA levels in mouse jejunum samples. Lct mRNA levels examined in the jejunum of adult (post-natal day 60) and infant (post-natal day 6) mice. ***p < 0.0001; two-tailed Student's t-test. Data are means ± s.e.m. (n = 11 and 7 mice). (b) Chromosome-wide scan of DNA modifications in the enterocytes of aging mice. Numerous DNA modification differences were detected on chromosome 1 of jejunal enterocytes from postnatal day 6 mice (infants) compared to day 60 (adults) mice (n = 14 and 15 mice, respectively). Within the extended Lct and Mcm6 region (100 kb), the largest DNA modification difference between the two mouse age groups were found at Lct intron 2, while smaller differences occurred at Lct intron 8–exon 9 and exon 10. Lct downregulation with age correlated with a gain in DNA modifications at these regions. SLP indicates the significance of correlation with age, with negative values signifying higher DNA modification levels in adult mice. The dashed line indicates the significance threshold after Benjamini-Hochberg multiple testing correction. (c) Schematic drawing of the regions containing DNA modifications associated with LCT mRNA regulation, as identified in a chromosome-wide scan (microarrays) and fine mapping investigation (bisulfite padlock probe technique). DNA modifications were investigated in jejunal enterocytes from human samples (cohorts 1 and 2) as well as infant and adult mice C57BL/6N inbred mice (post-natal day 6 vs 60). Peach bars indicate regions with high sequence homology between humans and mice. (d) Sequence alignment of MCM6 intron 13 region in human (GRCh37/hg19; chr2: 136608600-136608770) with corresponding region in mouse (C57BL/6 strain, GRCm38/mm10; chr1:128337434-128337972). This region was deleted in the mouse CRISPR-Cas9 experiment. MCM6 SNP C/T-13910 site is highlighted in yellow and * indicates DNA sequence homology between human and mouse.
Supplementary Figure 5 Comparison of DNA-modification densities at LCT regulatory regions in intestinal and nonintestinal tissue.
DNA modification profiles in enterocytes (C) differ from those of the enterocyte-deficient jejunum (T), and even more so from those of blood (BL). Principal components analysis was performed on the DNA modification levels for 262 samples across the seven key LCT regulatory regions identified in this study. This study found 42 CpG sites associated with LCT regulation, which clustered into the following seven regions: LCT intron 5, intron 3, intron 2, and exon 1 as well as MCM6 3′-UTR, exon 16 and intron 13–exon 13. For the principal components analysis there were 31 samples without a measurement for 1 region (2 blood, 13 enterocyte, 19 enterocyte-deficient jejunum), and 3 samples without a measurement 2 regions (3 enterocyte-deficient jejunum). Missing values were imputed using k-nearest-neighbors (k=10) using the "impute.knn" function within the R "impute" package. The first two principal components are plotted.
Supplementary Figure 6 Genetic factors associated with lactase persistence and nonpersistence and their effects on DNA modifications at LCT regulatory regions.
(a) LCT mRNA levels in the human jejunum relative to SNP genotypes. Individuals genotyped for the SNP in MCM6 intron 13 C/T(–13910), the SNP in MCM6 intron 9 G/A(–22018) and the deletion (no deletion = L, deletion = S) in LCT intron 1. Total sample size was n = 56 individuals in cohort 1 and n = 59 individuals in cohort 2. The haplotype containing T(–13910), A(–22018), and deletion in LCT intron 1 was associated with higher LCT mRNA levels. Analysis of variance (ANOVA) found a SNP C/T(–13910) haplotype effect in cohort 1 and 2 (F2,53 = 58.5, p < 10-13 and F2,56 = 206.2, p < 10-25, respectively). * p < 0.05 and ** p < 10-3 compared to CC/GG/LL genotype; ## p < 10-3 compared to TT/AA/SS genotype; Tukey's honest significant difference (HSD) post-hoc test. Error bars represent s.e.m. (b) Linkage disequilibrium in LCT and MCM6 region in individuals with European ancestry. For the intestinal samples (n =115 individuals), three genetic variants were examined: MCM6 intron 13 C/T(–13910) (labelled 1), the SNP in MCM6 intron 9 G/A(–22018) (labelled 2) and the deletion in LCT intron 1 (labelled 3). Left panel: Haplotype analysis revealed strong linkage disequilibrium (D' > 0.96) between these SNPs. D' values are shown in the boxes. Right panel: Haplotypes and their population frequencies. Haplotypes containing the C(–13910) are lactase non-persistent and haplotype with T(–13910) are lactase persistent. (c) DNA modification densities for LCT–MCM6 SNP genotypes. DNA modifications in enterocyte samples and enterocyte-deficient jejunal samples were examined for each genotype. The seven loci that demonstrated a significant correlation between DNA modifications and LCT mRNA levels were examined. In enterocytes, average DNA modification density significantly diverges between genotypes at MCM6 exon 13–intron 13 (p < 10-10) and at MCM6 exon 16 (for G/A(–22018) p < 0.01). In enterocyte-deficient jejunum, smaller DNA modification density differences are observed between genotypes at MCM6 intron 13–exon 13 (p < 10-5) as well as the 3′UTR region of MCM6 (p < 0.05). p-values are one-way ANOVA. Sample sizes of individuals for SNP C/T(–13910) (CC, CT, TT) n = 43, 43 and 14; SNP G/A(–22018) (GG, GA, AA) n = 43, 42 and 15; LCT intron 1 deletion (LL, LS, SS) n = 27, 46 and 27. Data are means ± s.e.m. DNA modification densities in enterocytes of SNP C/T(–13910) for each of the seven regions is displayed in Figure 3 of main text. * p < 0.05 comparisons between homozygote allele carriers by Tukey's HSD post-hoc test.
Supplementary Figure 7 Epigenetic modifications affecting lactase-gene regulation in mice.
(a) Mapping of DNA modifications in mice using the bisulfite padlock probe sequencing approach revealed regions involved in the age-dependent decline in Lct. Enterocytes of infant (post-natal day 6) and adult mice (day 60) were compared (n = 16 and 13 mice, respectively). DNA modification densities at Lct intron 2 and intron 8 significantly increased with age (FDR q < 0.05). Borderline significant associations between Lct mRNA levels and DNA modification density were also observed at Mcm6 exon 15 and intron 3. Dashed line indicates the significance threshold. (b) Chromatin modification profiles in the small intestine of an adult (60 days old) C57BL/6N mouse. Examination of ENCODE data revealed that Lct intron 2 contained an enhancer element (H3K4me1 and H3K27ac). There was also a promoter element (H3K4me3, H3K27ac, POLR2A) in Lct exon 1–intron 1 and an inactive enhancer in Mcm6 intron 13 (H3K4me1 only). The ENCODE sample code for the adult mouse small intestine was ENCBS157ENC (n = 1 mouse). To investigate the capacity for epigenetically-controlled regulatory regions to alter Lct mRNA levels, we used the CRISPR-Cas9 genome editing technique to induce deletions in mice (deleted regions highlighted in pink).
Supplementary Figure 8 CRISPR–Cas9-induced deletions in mice and a human cell line.
(a) Chromatograms showing mutations in Lct intron 1, Lct intron 2 and Mcm6 intron 13 generated by the CRISPR-Cas9 genome editing technique in C57BL/6N mice. (b) Off-target analysis for the CRISPR-Cas9-modified mice. A total of 46 sites were tested in the genetically modified mice and no off-target mutations were detected. Red, blue and green coordinates are the sites tested for the Mcm6 intron 13, Lct intron 1 and Lct intron 2 sgRNA guides, respectively. The orange arrow indicates the Lct–Mcm6 locus. (c) Off-target analysis for the CRISPR-Cas9n-modified human Caco-2 cell line. A total of 172 sites were tested in the genetically-modified cells and 2 off-target mutations were detected in the LCT intron 2 line (highlight in bold, top right insert). The off-target activity was at chr2:185,963,907 (11 bp deletion) and chr2:240,525,653 (21 bp insertion). These off-target deletions were both over 49 Mb from the LCT locus and unlikely to explain the differences in LCT mRNA in CRISPR-Cas9n-modified cells. Red, blue and green coordinates are the sites tested for the MCM6 intron 13, LCT intron 1 and LCT intron 2 sgRNA guides, respectively. The orange arrow indicates LCT–MCM6 locus. (d) The CRISPR-Cas9 genome editing technique generated three mouse lines (837, 1002, 1004) with the same deletion in Lct intron 2. Similar decreases in LCT mRNA were observed in each line. Mouse sample size for 837, 1002, 1004: WT n = 5, 3 and 5; homozygote mutant n = 8, 4 and 7. Two-way ANOVA showed a significant genotype effect (F1,26 = 173.0, p < 10-14). *p < 0.05, **p < 0.01, ***p < 10-3 between each homozygote mutant and respective wild-type group; Tukey's HSD post-hoc test. Data are means ± s.e.m.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1 and 2 and Supplementary Notes 1 and 2 (PDF 2543 kb)
Supplementary Table 3
sgRNAs and primer sequences used for the CRISPR-Cas9 deletions in mice and a human intestinal cell line. (XLSX 43 kb)
Supplementary Table 4
Bisulfite padlock probe sequences and barcode primers used for the high resolution DNA modification analysis at lactase gene region in humans and mice. (XLSX 68 kb)
Supplementary Data Set 1
Chromosome-wide scan of jejunal enterocytes identifies significant DNA modification differences between infant and adult mice. (XLSX 752 kb)
Rights and permissions
About this article
Cite this article
Labrie, V., Buske, O., Oh, E. et al. Lactase nonpersistence is directed by DNA-variation-dependent epigenetic aging. Nat Struct Mol Biol 23, 566–573 (2016). https://doi.org/10.1038/nsmb.3227
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3227
This article is cited by
-
Genetic and cultural adaptations underlie the establishment of dairy pastoralism in the Tibetan Plateau
BMC Biology (2023)
-
Epigenetic inactivation of the autophagy–lysosomal system in appendix in Parkinson’s disease
Nature Communications (2021)
-
Origins of human disease: the chrono-epigenetic perspective
Nature Reviews Genetics (2021)
-
Plasticity-Led Evolution and Human Culture
Integrative Psychological and Behavioral Science (2021)
-
Lactose Intolerance—Old and New Knowledge on Pathophysiological Mechanisms, Diagnosis, and Treatment
SN Comprehensive Clinical Medicine (2021)