Abstract
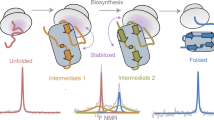
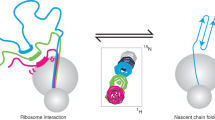
Although detailed pictures of ribosome structures are emerging, little is known about the structural and cotranslational folding properties of nascent polypeptide chains at the atomic level. Here we used solution-state NMR spectroscopy to define a structural ensemble of a ribosome–nascent chain complex (RNC) formed during protein biosynthesis in Escherichia coli, in which a pair of immunoglobulin-like domains adopts a folded N-terminal domain (FLN5) and a disordered but compact C-terminal domain (FLN6). To study how FLN5 acquires its native structure cotranslationally, we progressively shortened the RNC constructs. We found that the ribosome modulates the folding process, because the complete sequence of FLN5 emerged well beyond the tunnel before acquiring native structure, whereas FLN5 in isolation folded spontaneously, even when truncated. This finding suggests that regulating structure acquisition during biosynthesis can reduce the probability of misfolding, particularly of homologous domains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dobson, C.M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Onuchic, J.N., Luthey-Schulten, Z. & Wolynes, P.G. Theory of protein folding: the energy landscape perspective. Annu. Rev. Phys. Chem. 48, 545–600 (1997).
Dobson, C.M., Sali, A. & Karplus, M. Protein folding: a perspective from theory and experiment. Angew. Chem. Int. Ed. Eng. 37, 868–893 (1998).
Cabrita, L.D., Dobson, C.M. & Christodoulou, J. Protein folding on the ribosome. Curr. Opin. Struct. Biol. 20, 33–45 (2010).
Nicola, A.V., Chen, W. & Helenius, A. Co-translational folding of an alphavirus capsid protein in the cytosol of living cells. Nat. Cell Biol. 1, 341–345 (1999).
Zhang, G., Hubalewska, M. & Ignatova, Z. Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nat. Struct. Mol. Biol. 16, 274–280 (2009).
Frydman, J., Erdjument-Bromage, H., Tempst, P. & Hartl, F.U. Co-translational domain folding as the structural basis for the rapid de novo folding of firefly luciferase. Nat. Struct. Biol. 6, 697–705 (1999).
O'Brien, E.P., Christodoulou, J., Vendruscolo, M. & Dobson, C.M. New scenarios of protein folding can occur on the ribosome. J. Am. Chem. Soc. 133, 513–526 (2011).
Kaiser, C.M., Goldman, D.H., Chodera, J.D., Tinoco, I. Jr. & Bustamante, C. The ribosome modulates nascent protein folding. Science 334, 1723–1727 (2011).
Kim, S.J. et al. Protein folding. Translational tuning optimizes nascent protein folding in cells. Science 348, 444–448 (2015).
Schmeing, T.M. & Ramakrishnan, V. What recent ribosome structures have revealed about the mechanism of translation. Nature 461, 1234–1242 (2009).
Siller, E., DeZwaan, D.C., Anderson, J.F., Freeman, B.C. & Barral, J.M. Slowing bacterial translation speed enhances eukaryotic protein folding efficiency. J. Mol. Biol. 396, 1310–1318 (2010).
Deuerling, E., Schulze-Specking, A., Tomoyasu, T., Mogk, A. & Bukau, B. Trigger factor and DnaK cooperate in folding of newly synthesized proteins. Nature 400, 693–696 (1999).
Schaffitzel, C. et al. Structure of the E. coli signal recognition particle bound to a translating ribosome. Nature 444, 503–506 (2006).
Kramer, G., Boehringer, D., Ban, N. & Bukau, B. The ribosome as a platform for co-translational processing, folding and targeting of newly synthesized proteins. Nat. Struct. Mol. Biol. 16, 589–597 (2009).
Rutkowska, A. et al. Dynamics of trigger factor interaction with translating ribosomes. J. Biol. Chem. 283, 4124–4132 (2008).
Kaiser, C.M. et al. Real-time observation of trigger factor function on translating ribosomes. Nature 444, 455–460 (2006).
Ferbitz, L. et al. Trigger factor in complex with the ribosome forms a molecular cradle for nascent proteins. Nature 431, 590–596 (2004).
Knight, A.M. et al. Electrostatic effect of the ribosomal surface on nascent polypeptide dynamics. ACS Chem. Biol. 8, 1195–1204 (2013).
Kelkar, D.A., Khushoo, A., Yang, Z. & Skach, W.R. Kinetic analysis of ribosome-bound fluorescent proteins reveals an early, stable, cotranslational folding intermediate. J. Biol. Chem. 287, 2568–2578 (2012).
Clark, P.L. & King, J. A newly synthesized, ribosome-bound polypeptide chain adopts conformations dissimilar from early in vitro refolding intermediates. J. Biol. Chem. 276, 25411–25420 (2001).
Bhushan, S. et al. SecM-stalled ribosomes adopt an altered geometry at the peptidyl transferase center. PLoS Biol. 9, e1000581 (2011).
Bhushan, S. et al. α-Helical nascent polypeptide chains visualized within distinct regions of the ribosomal exit tunnel. Nat. Struct. Mol. Biol. 17, 313–317 (2010).
Lu, J. & Deutsch, C. Folding zones inside the ribosomal exit tunnel. Nat. Struct. Mol. Biol. 12, 1123–1129 (2005).
Woolhead, C.A., Johnson, A.E. & Bernstein, H.D. Translation arrest requires two-way communication between a nascent polypeptide and the ribosome. Mol. Cell 22, 587–598 (2006).
Nilsson, O.B. et al. Cotranslational protein folding inside the ribosome exit tunnel. Cell Rep. 12, 1533–1540 (2015).
Cabrita, L.D., Hsu, S.T., Launay, H., Dobson, C.M. & Christodoulou, J. Probing ribosome-nascent chain complexes produced in vivo by NMR spectroscopy. Proc. Natl. Acad. Sci. USA 106, 22239–22244 (2009).
Hsu, S.T. et al. Structure and dynamics of a ribosome-bound nascent chain by NMR spectroscopy. Proc. Natl. Acad. Sci. USA 104, 16516–16521 (2007).
Nakatogawa, H. & Ito, K. The ribosomal exit tunnel functions as a discriminating gate. Cell 108, 629–636 (2002).
Rosenzweig, R. & Kay, L.E. Bringing dynamic molecular machines into focus by methyl-TROSY NMR. Annu. Rev. Biochem. 83, 291–315 (2014).
Schanda, P., Kupce, E. & Brutscher, B. SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J. Biomol. NMR 33, 199–211 (2005).
Camilloni, C., Cavalli, A. & Vendruscolo, M. Replica-averaged metadynamics. J. Chem. Theory Comput. 9, 5610–5617 (2013).
Peterson, J.H., Woolhead, C.A. & Bernstein, H.D. The conformation of a nascent polypeptide inside the ribosome tunnel affects protein targeting and protein folding. Mol. Microbiol. 78, 203–217 (2010).
Frauenfeld, J. et al. Cryo-EM structure of the ribosome–SecYE complex in the membrane environment. Nat. Struct. Mol. Biol. 18, 614–621 (2011).
Hsu, S.T., Cabrita, L.D., Fucini, P., Dobson, C.M. & Christodoulou, J. Structure, dynamics and folding of an immunoglobulin domain of the gelation factor (ABP-120) from Dictyostelium discoideum. J. Mol. Biol. 388, 865–879 (2009).
Deuerling, E. et al. Trigger Factor and DnaK possess overlapping substrate pools and binding specificities. Mol. Microbiol. 47, 1317–1328 (2003).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
O'Brien, E.P., Ciryam, P., Vendruscolo, M. & Dobson, C.M. Understanding the influence of codon translation rates on cotranslational protein folding. Acc. Chem. Res. 47, 1536–1544 (2014).
Clarke, J., Cota, E., Fowler, S.B. & Hamill, S.J. Folding studies of immunoglobulin-like beta-sandwich proteins suggest that they share a common folding pathway. Structure 7, 1145–1153 (1999).
Kim, Y.E., Hipp, M.S., Bracher, A., Hayer-Hartl, M. & Hartl, F.U. Molecular chaperone functions in protein folding and proteostasis. Annu. Rev. Biochem. 82, 323–355 (2013).
Hartl, F.U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Wright, C.F., Teichmann, S.A., Clarke, J. & Dobson, C.M. The importance of sequence diversity in the aggregation and evolution of proteins. Nature 438, 878–881 (2005).
Sivashanmugam, A. et al. Practical protocols for production of very high yields of recombinant proteins using Escherichia coli. Protein Sci. 18, 936–948 (2009).
Rutkowska, A. et al. Large-scale purification of ribosome-nascent chain complexes for biochemical and structural studies. FEBS Lett. 583, 2407–2413 (2009).
Ferrage, F., Zoonens, M., Warschawski, D.E., Popot, J.L. & Bodenhausen, G. Slow diffusion of macromolecular assemblies by a new pulsed field gradient NMR method. J. Am. Chem. Soc. 125, 2541–2545 (2003).
Didenko, T., Boelens, R. & Rüdiger, S.G. 3D DOSY-TROSY to determine the translational diffusion coefficient of large protein complexes. Protein Eng. Des. Sel. 24, 99–103 (2011).
Waudby, C.A. & Christodoulou, J. An analysis of NMR sensitivity enhancements obtained using non-uniform weighted sampling, and the application to protein NMR. J. Magn. Reson. 219, 46–52 (2012).
Augustyniak, R., Ferrage, F., Damblon, C., Bodenhausen, G. & Pelupessy, P. Efficient determination of diffusion coefficients by monitoring transport during recovery delays in NMR. Chem. Commun. (Camb.) 48, 5307–5309 (2012).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Christodoulou, J. et al. Heteronuclear NMR investigations of dynamic regions of intact Escherichia coli ribosomes. Proc. Natl. Acad. Sci. USA 101, 10949–10954 (2004).
Timpe, L.C. & Peller, L. A random flight chain model for the tether of the Shaker K+ channel inactivation domain. Biophys. J. 69, 2415–2418 (1995).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Tribello, G.A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED 2: new feathers for an old bird. Comput. Phys. Commun. 185, 604–613 (2014).
Piana, S., Lindorff-Larsen, K. & Shaw, D.E. How robust are protein folding simulations with respect to force field parameterization? Biophys. J. 100, L47–L49 (2011).
Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, M.L. & Klein, L. Comparison of simple potential functions for stimulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Hess, B., Bekker, H., Berendsen, H.J.C. & Fraaije, J. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Acknowledgements
We thank J. Kirkpatrick for NMR technical assistance and useful discussions and B. Bukau (Ruprecht-Karls-Universität Heidelberg) for the kind gift of the anti-SecM antibody. J.C. acknowledges the use of the Biomolecular NMR Facility, University College London, and thanks T. Frenkiel and G. Kelly of the Medical Research Council Biomedical NMR Centre at the Crick Institute, London for the use of the facility. J.C. and T.W. acknowledge the use of the Advanced Research Computing High End Resource (ARCHER) UK National supercomputing service (http://www.archer.ac.uk/). L.D.C. is supported by the Wellcome Trust and by an Alpha-1 Foundation grant. A.L.R. is supported as a National Health and Medical Research Council (Australia) C.J. Martin Fellow. T.W. is supported as an European Molecular Biology Organization Long-Term Fellow and is also supported by the Wellcome Trust. The work of C.M.D. and M.V. is supported by a Wellcome Trust Programme Grant (094425/Z/10/Z to C.M.D. and M.V.). This work was supported by a Biotechnology and Biochemical Sciences Research Council New Investigators Award (BBG0156511 to J.C.) and a Wellcome Trust Investigator Award (097806/Z/11/Z to J.C.).
Author information
Authors and Affiliations
Contributions
L.D.C., C.M.D. and J.C. designed the research. J.C. supervised the overall project. L.D.C., A.M.E.C., H.M.M.L., C.A. Waudby, M.-E.K., A.S.W. and X.W. performed the research. A.L.R. and L.D.C. performed the PEGylation experiments, and C.A. Woodhead contributed valuable technical experience in in vitro–translation experiments. C.C., T.W., L.S.G. and M.V. performed the NMR-restrained molecular dynamics simulations. L.D.C., A.M.E.C., A.L.R., C.A. Waudby and J.C. analyzed the data. L.D.C., A.M.E.C., C.A. Waudby, M.V., C.M.D. and J.C. wrote the paper. All authors discussed the results and contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs.
(a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used in this study. Sequence boundaries are indicated. The isolated FLN constructs comprise: a range of FLN5 C-terminal truncations in which between 2 and 21 amino acids have been removed, “FLNΔX”, in which X refers to the extent of the truncation, the folding incompetent variant “FLN5 Y719E” harboring a destabilizing Glu mutation at the site indicated in orange, and “FLN5-6Δ18” in which FLN6 has the removal of 18 amino acids from its C-terminus (Hsu, S.T. et al. Proc Natl Acad Sci U S A 104, 16516-21, 2007). In SecM-stalled FLN5 RNCs, the FLN5 sequence is tethered to the PTC via increasing lengths of the FLN6 sequence and the SecM translational arrest motif; for simplicity this will be referred to here as the “linker”. The linker, ranging from 21 to 110 amino acids, therefore corresponds to the distance (in residues) between its most C-terminal residue, G750, and the PTC. Folding-incompetent RNCs have the Y719E mutation in FLN5 and are referred to as “FLN5+L Y719E RNCs”. For the poly(GS) RNCs, FLN6 is replaced with a repeating poly(GS) motif, GGGG/S, to generate equivalent linkers of between 21 to 67 residues in length, as present in the FLN5 RNCs. (b) Representative PAGE gel of a purified RNC (FLN5+42) and of 70S ribosomes visualized with Coomassie blue stain (left) and silver stain (right). The gels show a banding pattern characteristic of ribosomal proteins and show that the RNCs are essentially free of extraneous proteins. The bands corresponding to ribosomal proteins L1, L2, L6, S5, L29, L31, L35 are highlighted. (c) An anti-His western blot of FLN5+42 RNC and of 70S ribosomes, showing the tRNA-bound form of the RNC and the absence of any non-specific detection of ribosomal proteins in untranslating 70S ribosomes. The extent of nascent chain attachment to ribosomes was determined to be > 90% across all samples. (d) The amount of residual trigger factor (TF) and DnaK present within the purified RNC samples was assessed by densitometry analysis of anti-TF and anti-DnaK western blots, respectively. The purified samples are essentially free of the effects of TF and DnaK (≤ 1.2%). Highlighted in asterisks are the RNCs for which TF or DnaK levels were below levels of detection. (e) Representative western blots of RNC samples which were presented in a modified form in Fig. 2c are shown here for clarity, where the RNCs shown in Fig. 2c are marked by asterisks.
Supplementary Figure 2 Monitoring of the integrity of RNCs.
An integrated approach using NMR and biochemical analyses was used to determine the integrity of the RNCs over time. (a) For each of FLN5+21, 45, 47, 67 and 110 RNCs, the 1H-13C correlation spectrum is shown (identical to Fig. 3a), overlaid with that of isolated FLN5 (cyan). Resonances arising from labelled 70S proteins can be observed in spectra at ca. δH=0.8 p.p.m. (b) The integrity of these RNCs was monitored over time and the timeframe during which the nascent chain was determined to be attached in each RNC as derived by NMR methods is indicated by the shaded region: 13C-edited diffusion experiments (1H STE-1H,13C-HMQC) were used to determine the diffusion coefficient associated with the nascent chain in FLN5+67 and 110 RNCs (cyan). The changes in intensities of the cross-peaks observed in the correlation spectra were used to monitor the FLN5+45 and 47 RNCs (green) and the diffusion coefficients (from 13C-edited experiments) of the combined ribosome-derived resonances were monitored for FLN5+21 RNC (yellow). (c) Western blots against either the N-terminal His-tag or C-terminal SecM sequence report on both the tRNA-bound and released forms of the nascent chain. (d) Densitometry analysis of the tRNA-bound form over time as assessed by the anti-His western blot. In the analyses shown in panels b, c and d, the shaded region represents the timeframe corresponding to an exclusively ribosome-bound nascent chain and for which the 2D correlation spectra are summed and presented in panel a. (e) As a representative example for 15N-labelled RNCs, a 1H-15N SOFAST-HMQC spectrum (identical to that in Fig. 3b) of FLN5+21 RNC (black) is overlaid with a 2D correlation spectrum of isolated FLN5∆12 (blue) and a selection of unambiguously assigned resonances is indicated. (f) Diffusion coefficient for FLN5+21 RNC resonances assessed over time (calculated from 15N XSTE spectra; single spectrum shown on left). A signal attenuation (I95/I5) by a factor of > 0.62 corresponds to a diffusion coefficient of an intact ribosomal particle (D = 2.0 ± 0.3 10-11 m2 s-1) at 25°C. The timeframe during which the nascent chain is assessed as being intact is shaded in grey. (g) 15N XSTE spectra of FLN5+21 RNC recorded at gradient strengths 5% and 95% of the maximum gradient strength Gmax, 0.557 T m-1 for the timeframe during which the nascent chain was deemed to be intact. (h) Anti-His and anti-SecM western blot analyses of FLN5+21 RNC.
Supplementary Figure 3 U-15N–labeled spectra: assignments, chemical-shift analyses and linewidth measurements on FLN5 RNCs and isolated FLN5 variants.
1H-15N correlation spectra and resonance assignments of: (a) FLN5Δ12 (b) FLN5-6Δ18 (c) the 70S ribosome for which the ribosomal protein L7/L12 gives rise to most resonances that are observed (Christodoulou, J. et al. Proc Natl Acad Sci U S A 101, 10949-10954, 2004). (d) 1H-15N correlation spectrum of FLN+21 RNC. (e) Overlay of the 1H-15N correlation spectra of FLN5+21 RNC (black) and isolated FLN5Δ12 (blue), demonstrating that the disordered region of FLN5+21 RNC spectrum (7.9-8.6 ppm in 1H dimension) overlays closely with that of disordered FLN5. (f) Overlay of FLN5+21 RNC (black) and FLN5-6Δ18 (green); the resonances in the disordered region of the RNC spectrum do not correspond to those of unfolded FLN6. (g) Overlay of the spectra of FLN5+21 RNC (black) and of the 70S ribosome particle (magenta), the limited overlap with ribosomal protein L7/L12 resonances indicating an essentially negligible level of background signal of ribosomal proteins in the RNC. (h) 1H-15N correlation spectrum of FLN5+110 RNC. (i) Overlay of 1H-15N correlation spectra of FLN5+110 RNC (black) and FLN5Δ12 (blue); the resonances in the disordered region of the RNC spectrum clearly do not correspond to those of unfolded isolated FLN5. (j) Overlay of 1H-15N correlation spectra of FLN5+110 RNC (black) and FLN5-6Δ18 (green); the unfolded region of FLN5+110 RNC overlays closely with that of disordered FLN6. (k) Overlay of 1H-15N correlation spectra of FLN5+110 RNC (black) and 70S ribosomes (magenta); there is a negligible contribution of resonances arising from the presence of labelled 70S ribosomal proteins. (l) 1H-15N correlation spectrum of FLN5 Y719E (orange), overlaid with that of a C-terminally truncated variant, FLN5Δ12 (black). The close overlay demonstrates that the Y719E mutation (generated based upon predictions using the PoPMuSiC algorithm (Dehouck, Y., Kwasigroch, J.M., Gilis, D. & Rooman, M. BMC Bioinformatics 12, 151, 2011) results in an unfolded conformation that is highly comparable to that observed in FLN5Δ12. 15N R2 relaxation rates (at 277 K, data not shown) averaged 2.8 ± 0.6 s-1 for the 81 residues analysed (error taken as the standard deviation), indicating a highly disordered protein. The 1H-15N spectra of both proteins were completely assigned and the key chemical shift changes between the two are indicated with dotted lines. (m) Chemical shift differences between FLN5 Y719E and FLN5Δ12 are mapped against the amino acid sequence, as calculated by the formula  This shows that the chemical shifts are closely similar between the two constructs apart from at the mutation site (marked by an asterisk) and at the C-terminus of FLN5Δ12. (n) Comparison of linewidths between isolated FLN5Δ16 and FLN5+21, +31, +37 and +42 RNCs as measured for resonances Ala683, Ala694 and Val682. (o) 1H cross-sections of FLN5+21, 31, 37 and 42 RNC resonances used for lineshape fitting (as described in the Online Methods): Ala683 and Ala694 are fitted simultaneously to Lorentzian lineshapes. RNC linewidths (blue) are greater than those observed for isolated FLN5Δ16 (grey).
This shows that the chemical shifts are closely similar between the two constructs apart from at the mutation site (marked by an asterisk) and at the C-terminus of FLN5Δ12. (n) Comparison of linewidths between isolated FLN5Δ16 and FLN5+21, +31, +37 and +42 RNCs as measured for resonances Ala683, Ala694 and Val682. (o) 1H cross-sections of FLN5+21, 31, 37 and 42 RNC resonances used for lineshape fitting (as described in the Online Methods): Ala683 and Ala694 are fitted simultaneously to Lorentzian lineshapes. RNC linewidths (blue) are greater than those observed for isolated FLN5Δ16 (grey).
Supplementary Figure 4 PEGylation of FLN5 Y719E RNCs.
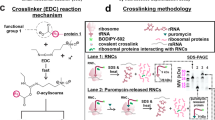
(a) Autoradiography of SDS-PAGE gels as presented in a cropped form in Fig. 4b is provided here for clarity. The arrows indicate the different forms of FLN5 nascent chains observed: tRNA-bound nascent chain (green), PEGylated tRNA-bound nascent chain (red) and released nascent chain (cyan), PEGylated released nascent chain (blue). Approximate migration of molecular weight standards is indicated on the left. (b) RNase A treated FLN5 Y719E RNCs, showing the PEGylation achievable on the released nascent chains (color coding as in a). Approximate migration of molecular weight standards is indicated on the left.
Supplementary Figure 5 1H-15N correlation spectra of FLN5 C-terminal truncations.
(a) Natively folded full-length FLN5 domain. (b) FLN5Δ8. The removal of eight C-terminal residues results in FLN5 sampling both unfolded and native-like conformational states. (c) FLN5Δ16, shows a characteristic spectrum of an unfolded polypeptide. Below, the regions removed by the C-terminal truncation are highlighted on ribbon diagrams of FLN5 (PDB: 1QFH), in magenta for FLN5Δ8 and blue for FLN5Δ16.
Supplementary Figure 6 FLN5 RNCs in which FLN6 is substituted with a poly(GS) linker.
(a) Schematic of the poly(GS)-linker RNCs, in which the FLN6 domain is substituted with a poly(GS) sequence. The FLN6 residues that have been substituted in two RNCs analysed by NMR, FLN5+31 and +42 to generate FLN5+31 GS and +42 GS, respectively, are shown in the highlighted box. (b) Anti-His western blot of FLN5+31 GS and 42 GS RNCs in their released and tRNA-bound forms, respectively. (c) The emergence of FLN5 from the exit tunnel in a series of poly(GS)-linker RNCs was monitored by PEGylation of G751C variants of the FLN5 RNCs in a manner similar to that shown in Fig. 4 for the FLN6 domain. (d) Overlay of 1H-15N correlation spectra of FLN5+42 GS RNC and FLN5+42 RNC. The latter was recorded at 950 MHz and with enhanced resolution, however FLN5+42 and +42 GS RNCs have similar intrinsic linewidths. The close overlay suggests shows the similar unfolded conformational preferences between the two nascent chains i.e. FLN5 is unfolded in the FLN5+42 GS RNC. (e) Overlay of 1H-15N correlation spectra FLN5+42 GS RNC when it is intact (dark green) and released (after ˜20 h of NMR acquisition, light green), which shows the appearance of additional glycine resonances arising from the poly(GS) linker in the released nascent chain. (f) Relative intensities of FLN5+42 (upper panel) and FLN5+31 (lower panel) GS RNCs compared to isolated, unfolded FLN5 (the green trace represents a 5-point moving average). 5-point moving average plots of the relative intensities of FLN5+42 (upper panel) and FLN5+31 (lower panel) RNCs compared to isolated unfolded FLN5 are also shown for comparison (pink). The close overlay of the two indicates that altering the linker does not seem to impart significantly on the conformation and/or ribosome interactions of unfolded FLN5. (g) Overlay of 1H-15N correlation spectra of FLN5+42 GS RNC after ~20 h of NMR acquisition and of natively folded FLN5: upon release from the ribosome, resonances corresponding to natively folded FLN5 are observed.
Supplementary Figure 7 Interactions between folded and unfolded isolated FLN5 variants with 70S ribosomes.
Relative peak intensities of: (a) FLN5 in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5 alone, at 5 μM (as presented in Fig. 6b) (b) FLN5Δ12 in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5Δ12 alone, at 5 μM. (c) FLN5 Y719E in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5 Y719E alone, at 8 µM (as presented in Fig. 6b). Shaded areas highlight the peak broadenings observed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1269 kb)
A structural ensemble of a ribosome-bound nascent chain
An NMR-restrained structural ensemble for FLN5+110 RNC during in vivo co-translational protein folding depicting the lowest free energy microstates accessible by the ribosome-bound FLN5+110 nascent chain. (MOV 22689 kb)
Rights and permissions
About this article
Cite this article
Cabrita, L., Cassaignau, A., Launay, H. et al. A structural ensemble of a ribosome–nascent chain complex during cotranslational protein folding. Nat Struct Mol Biol 23, 278–285 (2016). https://doi.org/10.1038/nsmb.3182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3182