Abstract
DNA 5-methylcytosine is a dynamic epigenetic mark with important roles in development and disease. In the Tet-Tdg demethylation pathway, methylated cytosine is iteratively oxidized by Tet dioxygenases, and unmodified cytosine is restored via thymine DNA glycosylase (Tdg). Here we show that human NEIL1 and NEIL2 DNA glycosylases coordinate abasic-site processing during TET-TDG DNA demethylation. NEIL1 and NEIL2 cooperate with TDG during base excision: TDG occupies the abasic site and is displaced by NEILs, which further process the baseless sugar, thereby stimulating TDG-substrate turnover. In early Xenopus embryos, Neil2 cooperates with Tdg in removing oxidized methylcytosines and specifying neural-crest development together with Tet3. Thus, Neils function as AP lyases in the coordinated AP-site handover during oxidative DNA demethylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Schübeler, D. Function and information content of DNA methylation. Nature 517, 321–326 (2015).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. Int. Edn Engl. 50, 7008–7012 (2011).
Kohli, R.M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013).
Wu, S.C. & Zhang, Y. Active DNA demethylation: many roads lead to Rome. Nat. Rev. Mol. Cell Biol. 11, 607–620 (2010).
Maiti, A. & Drohat, A.C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338 (2011).
Cortázar, D. et al. Embryonic lethal phenotype reveals a function of TDG in maintaining epigenetic stability. Nature 470, 419–423 (2011).
Cortellino, S. et al. Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146, 67–79 (2011).
Shen, L. et al. Genome-wide analysis reveals TET- and TDG-dependent 5-methylcytosine oxidation dynamics. Cell 153, 692–706 (2013).
Song, C.X. et al. Genome-wide profiling of 5-formylcytosine reveals its roles in epigenetic priming. Cell 153, 678–691 (2013).
Raiber, E.A. et al. Genome-wide distribution of 5-formylcytosine in embryonic stem cells is associated with transcription and depends on thymine DNA glycosylase. Genome Biol. 13, R69 (2012).
Zhu, B. et al. 5-methylcytosine-DNA glycosylase activity is present in a cloned G/T mismatch DNA glycosylase associated with the chicken embryo DNA demethylation complex. Proc. Natl. Acad. Sci. USA 97, 5135–5139 (2000).
Fromme, J.C. & Verdine, G.L. Base excision repair. Adv. Protein Chem. 69, 1–41 (2004).
Hosfield, D.J. et al. DNA damage recognition and repair pathway coordination revealed by the structural biochemistry of DNA repair enzymes. Prog. Nucleic Acid Res. Mol. Biol. 68, 315–347 (2001).
Prasad, R., Shock, D.D., Beard, W.A. & Wilson, S.H. Substrate channeling in mammalian base excision repair pathways: passing the baton. J. Biol. Chem. 285, 40479–40488 (2010).
Hazra, T.K. et al. Identification and characterization of a human DNA glycosylase for repair of modified bases in oxidatively damaged DNA. Proc. Natl. Acad. Sci. USA 99, 3523–3528 (2002).
Hazra, T.K. et al. Identification and characterization of a novel human DNA glycosylase for repair of cytosine-derived lesions. J. Biol. Chem. 277, 30417–30420 (2002).
Spruijt, C.G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–1159 (2013).
Müller, U., Bauer, C., Siegl, M., Rottach, A. & Leonhardt, H. TET-mediated oxidation of methylcytosine causes TDG or NEIL glycosylase dependent gene reactivation. Nucleic Acids Res. 42, 8592–8604 (2014).
Bandaru, V., Sunkara, S., Wallace, S.S. & Bond, J.P. A novel human DNA glycosylase that removes oxidative DNA damage and is homologous to Escherichia coli endonuclease VIII. DNA Repair (Amst.) 1, 517–529 (2002).
Liu, M. et al. The mouse ortholog of NEIL3 is a functional DNA glycosylase in vitro and in vivo. Proc. Natl. Acad. Sci. USA 107, 4925–4930 (2010).
Morland, I. et al. Human DNA glycosylases of the bacterial Fpg/MutM superfamily: an alternative pathway for the repair of 8-oxoguanine and other oxidation products in DNA. Nucleic Acids Res. 30, 4926–4936 (2002).
Takao, M. et al. A back-up glycosylase in Nth1 knock-out mice is a functional Nei (endonuclease VIII) homologue. J. Biol. Chem. 277, 42205–42213 (2002).
Krokeide, S.Z. et al. Human NEIL3 is mainly a monofunctional DNA glycosylase removing spiroimindiohydantoin and guanidinohydantoin. DNA Repair (Amst.) 12, 1159–1164 (2013).
Dou, H., Mitra, S. & Hazra, T.K. Repair of oxidized bases in DNA bubble structures by human DNA glycosylases NEIL1 and NEIL2. J. Biol. Chem. 278, 49679–49684 (2003).
Krokan, H.E. & Bjørås, M. Base excision repair. Cold Spring Harb. Perspect. Biol. 5, a012583 (2013).
Liu, M., Doublié, S. & Wallace, S.S. Neil3, the final frontier for the DNA glycosylases that recognize oxidative damage. Mutat. Res. 743-744, 4–11 (2013).
Arab, K. et al. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol. Cell 55, 604–614 (2014).
Steinacher, R. & Schär, P. Functionality of human thymine DNA glycosylase requires SUMO-regulated changes in protein conformation. Curr. Biol. 15, 616–623 (2005).
Waters, T.R., Gallinari, P., Jiricny, J. & Swann, P.F. Human thymine DNA glycosylase binds to apurinic sites in DNA but is displaced by human apurinic endonuclease 1. J. Biol. Chem. 274, 67–74 (1999).
Hardeland, U., Bentele, M., Jiricny, J. & Schär, P. Separating substrate recognition from base hydrolysis in human thymine DNA glycosylase by mutational analysis. J. Biol. Chem. 275, 33449–33456 (2000).
Hardeland, U., Steinacher, R., Jiricny, J. & Schär, P. Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J. 21, 1456–1464 (2002).
Fitzgerald, M.E. & Drohat, A.C. Coordinating the initial steps of base excision repair: apurinic/apyrimidinic endonuclease 1 actively stimulates thymine DNA glycosylase by disrupting the product complex. J. Biol. Chem. 283, 32680–32690 (2008).
Wiederhold, L. et al. AP endonuclease-independent DNA base excision repair in human cells. Mol. Cell 15, 209–220 (2004).
Dawlaty, M.M. et al. Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev. Cell 29, 102–111 (2014).
Xu, Y. et al. Tet3 CXXC domain and dioxygenase activity cooperatively regulate key genes for Xenopus eye and neural development. Cell 151, 1200–1213 (2012).
LaBonne, C. & Bronner-Fraser, M. Neural crest induction in Xenopus: evidence for a two-signal model. Development 125, 2403–2414 (1998).
Wilson, D.M. III & Barsky, D. The major human abasic endonuclease: formation, consequences and repair of abasic lesions in DNA. Mutat. Res. 485, 283–307 (2001).
Mokkapati, S.K., Wiederhold, L., Hazra, T.K. & Mitra, S. Stimulation of DNA glycosylase activity of OGG1 by NEIL1: functional collaboration between two human DNA glycosylases. Biochemistry 43, 11596–11604 (2004).
Pascucci, B. et al. Reconstitution of the base excision repair pathway for 7,8-dihydro-8-oxoguanine with purified human proteins. Nucleic Acids Res. 30, 2124–2130 (2002).
Banerjee, D. et al. Preferential repair of oxidized base damage in the transcribed genes of mammalian cells. J. Biol. Chem. 286, 6006–6016 (2011).
Lee, J. et al. AP endonucleases process 5-methylcytosine excision intermediates during active DNA demethylation in Arabidopsis. Nucleic Acids Res. 42, 11408–11418 (2014).
Li, Y. et al. An AP endonuclease functions in active DNA demethylation and gene imprinting in Arabidopsis. PLoS Genet. 11, e1004905 (2015).
Williams, K., Christensen, J. & Helin, K. DNA methylation: TET proteins-guardians of CpG islands? EMBO Rep. 13, 28–35 (2012).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011).
Vella, P. et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol. Cell 49, 645–656 (2013).
Dawlaty, M.M. et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev. Cell 24, 310–323 (2013).
Gu, T.P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Tini, M. et al. Association of CBP/p300 acetylase and thymine DNA glycosylase links DNA repair and transcription. Mol. Cell 9, 265–277 (2002).
Canugovi, C. et al. Endonuclease VIII-like 1 (NEIL1) promotes short-term spatial memory retention and protects from ischemic stroke-induced brain dysfunction and death in mice. Proc. Natl. Acad. Sci. USA 109, 14948–14953 (2012).
Regnell, C.E. et al. Hippocampal adult neurogenesis is maintained by Neil3-dependent repair of oxidative DNA lesions in neural progenitor cells. Cell Reports 2, 503–510 (2012).
Sejersted, Y. et al. Endonuclease VIII-like 3 (Neil3) DNA glycosylase promotes neurogenesis induced by hypoxia-ischemia. Proc. Natl. Acad. Sci. USA 108, 18802–18807 (2011).
Chakraborty, A. et al. Neil2-null mice accumulate oxidized DNA bases in the transcriptionally active sequences of the genome and are susceptible to innate inflammation. J. Biol. Chem. 290, 24636–24648 (2015).
Hashimoto, H. et al. Structure of a Naegleria Tet-like dioxygenase in complex with 5-methylcytosine DNA. Nature 506, 391–395 (2014).
Onizuka, K., Yeo, J., David, S.S. & Beal, P.A. NEIL1 binding to DNA containing 2′-fluorothymidine glycol stereoisomers and the effect of editing. ChemBioChem 13, 1338–1348 (2012).
Gawantka, V., Delius, H., Hirschfeld, K., Blumenstock, C. & Niehrs, C. Antagonizing the Spemann organizer: role of the homeobox gene Xvent-1. EMBO J. 14, 6268–6279 (1995).
Villanueva, S., Glavic, A., Ruiz, P. & Mayor, R. Posteriorization by FGF, Wnt, and retinoic acid is required for neural crest induction. Dev. Biol. 241, 289–301 (2002).
Bradley, L., Wainstock, D. & Sive, H. Positive and negative signals modulate formation of the Xenopus cement gland. Development 122, 2739–2750 (1996).
Sive, H.L., Grainiger, R.M. & Harland, R.M. Early Development of Xenopus laevis: A Laboratory Manual. Cold Spring Harbor Laboratory Press, (2000).
Saka, Y. & Smith, J.C. Spatial and temporal patterns of cell division during early Xenopus embryogenesis. Dev. Biol. 229, 307–318 (2001).
Hensey, C. & Gautier, J. Programmed cell death during Xenopus development: a spatio-temporal analysis. Dev. Biol. 203, 36–48 (1998).
Kellner, S. et al. Absolute and relative quantification of RNA modifications via biosynthetic isotopomers. Nucleic Acids Res. 42, e142 (2014).
Ravanat, J.L. et al. Cellular background level of 8-oxo-7,8-dihydro-2′-deoxyguanosine: an isotope based method to evaluate artefactual oxidation of DNA during its extraction and subsequent work-up. Carcinogenesis 23, 1911–1918 (2002).
Acknowledgements
We thank U. Stapf (IMB) for technical assistance, A. Rao (Division of Signaling and Gene Expression, La Jolla Institute for Allergy and Immunology), P. Schär (Department of Biomedicine, University of Basel) and Y. Zheng (New England BioLabs) for reagents and the IMB core facilities for technical support. This work was supported by an European Research Council Senior Investigator Grant to C.N. ('DNAdemethylase'). M.U.M. was supported by a Natural Sciences and Engineering Research Council of Canada Postdoctoral Fellowship (NSERC-PDF 403829-2011).
Author information
Authors and Affiliations
Contributions
L.S. and A.v.S. carried out biochemical assays. D.H. conducted Xenopus experiments. M.U.M. performed LC-MS/MS measurements. K.A. performed experiments on HNO387 cells. L.S. and S.K. carried out protein-protein interaction assays. All authors analyzed and discussed the data. L.S. and C.N. conceived the study, designed experiments and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 DNA demodification and glycosylase activities are sequence-context independent.
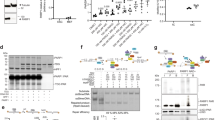
(a) Scheme of the substrates used for the assays. Left, sequence context 1, right, sequence context 2. The sequence context difference flanking the HpaII recognition site is shown. (b, c) DNA demodification assay (b) and DNA glycosylase assay (c) using HeLa cell extract (50 µg in a 50 µl assay) and 20 nM 5mC, 5hmC, 5fC and 5caC containing 160 bp dsDNA of the respective sequence context (compare also Figure 1b and Figure 2b). Product peaks of demodification- (+ HpaII treatment) and glycosylase activities (79mers; arrows) are shown with efficiencies in % of 79 nt peak integral relative to the total fluorescent signal per electropherogram. Note the similar glycosylase and demodification activities of the extract toward both oligos.
Supplementary Figure 2 Representative electropherograms of DNA glycosylase and DNA demodification assays.
(a,b) Primary data electropherograms from DNA glycosylase assays from (a) Figure 2c and (b) Figure 2d. Arrows indicate cleavage products (79 nt) with amounts of products shown as percentages above. (c) Primary data electropherograms from DNA demodification assay of Figure 2e. Arrows indicate HpaII cleavage products (79 nt) with amounts of products shown as percentages (arrow).
Supplementary Figure 3 Recombinant NEIL1 and NEIL2 possess 5hU glycosylase and AP lyase activity.
(a) Coomassie-stained SDS-PAGE of the indicated purified recombinant proteins used in this study. NgTet1: Naegleria gruberi Tet1. His6: N- or C-terminal hexahistidin tag. Relative molecular weights of marker proteins (M) [x103] are indicated on the left. (b-d) Electropherograms of reaction products from DNA glycosylase assays with ds (5hU:G; AP:G) and ss (5hU) oligonucleotide substrates. (b) NEIL1 and NEIL2 process ds and ssDNA containing 5-hydroxyuracil. Arrows indicate reaction products with efficiencies (%). Note: N-terminally hexahistidin-tagged NEIL1 or NEIL2 (H6-NEIL1, H6-NEIL2) are inactive and serve as negative controls. (c) NEIL1 and NEIL2 process AP sites. Arrows indicate reaction products with efficiencies (%). Note that spontaneous AP site hydrolysis under mock conditions is 25% (background). Synthesis of AP site containing oligonucleotides is described in ‘Online Methods’ section. (d) Recombinant NEIL1 and NEIL2 harbor AP lyase activity. A 79mer marker oligonucleotide was mixed with cleavage products shown in (c). The APEX1 cleavage product containing a 3’-hydroxyl co-migrated with the 79mer marker (blue peak). The cleavage products of NEIL1 or NEIL2 migrate slightly faster (arrows), indicative of a 3’-phosphate residue (containing two additional negative charges) after AP lyase reaction (β,δ-elimination).
Supplementary Figure 4 TDG stimulation controls with SMUG1 and APEX1 and effects of APEX1 on 5fC and 5caC processing in HeLa cells.
(a) DNA glycosylase assay of TDG and SMUG1 alone or in combination (100-fold molar excess SMUG1 (total protein) over TDG) on 5fC and 5caC containing ds oligonucleotides. Product peaks of glycosylase activities (79mers) are highlighted in red with efficiencies shown in % and -fold stimulations. Note: SMUG1 had no detectable glycosylase activity toward 5fC and 5caC substrates and was not able to stimulate base excision of TDG. S1, SMUG1. (b) DNA glycosylase assay as in (a) but with purified APEX1 instead of SMUG1 using a 5-fold molar excess (total protein) over TDG. A1, APEX1. (c) LC-MS/MS quantification of genomic cytosine modifications as in Figure 2f from HeLa cells siRNA depleted of the indicated genes. (d) DNA glycosylase assay on 5fC and 5caC containing oligonucleotides using HeLa extracts as in c. (e) Quantification of DNA glycosylase activities shown in d. Error bars, s.d. (n = 3 assay repetitions). **P < 0.01, ***P < 0.005 by two-tailed unpaired Student’s t-test. (f) Demodification assay on 5fC and 5caC containing oligonucleotides using HeLa extracts siRNA depleted of the indicated genes. Repair efficiencies are shown as %.
Supplementary Figure 5 NEIL1 and NEIL2 interact with TDG.
(a,b) Microscale thermophoresis binding assays. (a) Binding of fluorescently-labeled NEIL1 and NEIL2 to non-labeled TDG (reverse labeling as compared to binding assay shown in Figure 5c). (b) Left: Fluorescently-labeled TDG and SMUG1 do not interact in vitro. Right, positive control: High affinity binding of GFP to αGFP antibody. Fitted curves for each binding experiment (n = 3 binding assays), normalized fluorescence timetraces and calculated Kd-values are shown. Kd errors are calculated technical errors derived from the curve fittings. Fnorm (‰), normalized fluorescence per mill.
Supplementary Figure 6 Characterization of neil1, neil2, tdg and tet3 in X. laevis embryogenesis.
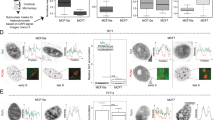
(a) qPCR expression profiles of neil1, neil2, tdg and tet3 during different stages of X. laevis development. Highest expression values per profiled gene were arbitrarily set to 100. Error bars, s.d. (n = 3 embryo batches each consisting of at least 5 embryos). (b) TUNEL (apoptosis) assay of unilaterally morpholino injected embryos (lineage traced by lacZ, light blue speckles) and quantification of TUNEL signal (dark blue speckles; n = 25, 32, 36, 36 embryos from left to right). Note that only tdg MO induces substantive apoptosis. (c) Phospho-histone H3 (cell proliferation) assay of unilaterally morpholino injected embryos as in (b). Note: No obvious differences in proliferation were detected (n >30 embryos each). (d) Phenotype of stage 34 embryos resulting from neil2 MO and neil2 MO2 injections. Note: Both morpholinos induced the same phenotype. (e), Relative expression levels of neil2 (normalized to h4) in control MO and neil2 MO injected embryos at stage 16 and stage 23. Scale bars, 200 µm.
Supplementary Figure 7 Relative abundance of X. laevis genomic DNA modifications.
(a) Quantification of total genomic 5mC, 5hmC, 5fC and 5caC levels in X. laevis whole embryos at stage 32, human HEK293T cells and mouse ES cells (mESCs) by LC-MS/MS. Error bars, s.d. (n = 3 X. laevis embryo batches consisting each of at least 5 embryos and 3 cell cultures of HEK293T and mESCs, respectively). (b) LC-MS/MS quantification as in (a) but in control and neil2 morphant Xenopus animal cap explants including measurements of 8oxoG. Error bars, s.d. (n = 3 explant batches each consisting of 20 animal cap explants). n.s., not significant. *P < 0.05, ***P < 0.005 by two-tailed unpaired Student’s t-test.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 1168 kb)
Supplementary Data Set 1
Original uncropped gels (PDF 1441 kb)
Rights and permissions
About this article
Cite this article
Schomacher, L., Han, D., Musheev, M. et al. Neil DNA glycosylases promote substrate turnover by Tdg during DNA demethylation. Nat Struct Mol Biol 23, 116–124 (2016). https://doi.org/10.1038/nsmb.3151
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3151
This article is cited by
-
DNA methylation restricts coordinated germline and neural fates in embryonic stem cell differentiation
Nature Structural & Molecular Biology (2024)
-
NEIL1 and NEIL2 DNA glycosylases modulate anxiety and learning in a cooperative manner in mice
Communications Biology (2021)
-
Active turnover of genomic methylcytosine in pluripotent cells
Nature Chemical Biology (2020)
-
Reversal of nucleobase methylation by dioxygenases
Nature Chemical Biology (2020)
-
GADD45A binds R-loops and recruits TET1 to CpG island promoters
Nature Genetics (2019)