Abstract
Rif1 regulates replication timing and repair of double-strand DNA breaks. Using a chromatin immunoprecipitation–sequencing method, we identified 35 high-affinity Rif1-binding sites in fission yeast chromosomes. Binding sites tended to be located near dormant origins and to contain at least two copies of a conserved motif, CNWWGTGGGGG. Base substitution within these motifs resulted in complete loss of Rif1 binding and in activation of late-firing or dormant origins located up to 50 kb away. We show that Rif1-binding sites adopt G quadruplex–like structures in vitro, in a manner dependent on the conserved sequence and on other G tracts, and that purified Rif1 preferentially binds to this structure. These results suggest that Rif1 recognizes and binds G quadruplex–like structures at selected intergenic regions, thus generating local chromatin structures that may exert long-range suppressive effects on origin firing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Masai, H., Matsumoto, S., You, Z., Yoshizawa-Sugata, N. & Oda, M. Eukaryotic chromosome DNA replication: where, when, and how? Annu. Rev. Biochem. 79, 89–130 (2010).
Méchali, M., Yoshida, K., Coulombe, P. & Pasero, P. Genetic and epigenetic determinants of DNA replication origins, position and activation. Curr. Opin. Genet. Dev. 23, 124–131 (2013).
Aparicio, O.M. Location, location, location: it's all in the timing for replication origins. Genes Dev. 27, 117–128 (2013).
Matsumoto, S., Hayano, M., Kanoh, Y. & Masai, H. Multiple pathways can bypass the essential role of fission yeast Hsk1 kinase in DNA replication initiation. J. Cell Biol. 195, 387–401 (2011).
Knott, S.R. et al. Forkhead transcription factors establish origin timing and long-range clustering in S. cerevisiae. Cell 148, 99–111 (2012).
Yoshida, K., Poveda, A. & Pasero, P. Time to be versatile: regulation of the replication timing program in budding yeast. J. Mol. Biol. 425, 4696–4705 (2013).
Hayano, M. et al. Rif1 is a global regulator of timing of replication origin firing in fission yeast. Genes Dev. 26, 137–150 (2012).
Yamazaki, S. et al. Rif1 regulates the replication timing domains on the human genome. EMBO J. 31, 3667–3677 (2012).
Cornacchia, D. et al. Mouse Rif1 is a key regulator of the replication-timing programme in mammalian cells. EMBO J. 31, 3678–3690 (2012).
Hiraga, S. et al. Rif1 controls DNA replication by directing Protein Phosphatase 1 to reverse Cdc7-mediated phosphorylation of the MCM complex. Genes Dev. 28, 372–383 (2014).
Davé, A., Cooley, C., Garg, M. & Bianchi, A. Protein phosphatase 1 recruitment by Rif1 regulates DNA replication origin firing by counteracting DDK activity. Cell Reports 7, 53–61 (2014).
Mattarocci, S. et al. Rif1 controls DNA replication timing in yeast through the PP1 phosphatase Glc7. Cell Reports 7, 62–69 (2014).
Silverman, J., Takai, H., Buonomo, S.B.C., Eisenhaber, F. & De Lange, T. Human Rif1, ortholog of a yeast telomeric protein, is regulated by ATM and 53BP1 and functions in the S-phase checkpoint. Genes Dev. 18, 2108–2119 (2004).
Zimmermann, M., Lottersberger, F., Buonomo, S.B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715 (2013).
Chapman, J.R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013).
Escribano-Díaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1–RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Callen, E. et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell 153, 1266–1280 (2013).
Feng, L., Fong, K.W., Wang, J., Wang, W. & Chen, J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. J. Biol. Chem. 288, 11135–11143 (2013).
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006).
Dan, J. et al. Rif1 maintains telomere length homeostasis of ESCs by mediating heterochromatin silencing. Dev. Cell 29, 7–19 (2014).
Kanoh, J. & Ishikawa, F. spRap1 and spRif1, recruited to telomeres by Taz1, are essential for telomere function in fission yeast. Curr. Biol. 11, 1624–1630 (2001).
Bailey, T.L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Tanizawa, H. et al. Mapping of long-range associations throughout the fission yeast genome reveals global genome organization linked to transcriptional regulation. Nucleic Acids Res. 38, 8164–8177 (2010).
Bochman, M.L., Paeschke, K. & Zakian, V.A. DNA secondary structures: stability and function of G-quadruplex structures. Nat. Rev. Genet. 13, 770–780 (2012).
Burge, S., Parkinson, G.N., Hazel, P., Todd, A.K. & Neidle, S. Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res. 34, 5402–5415 (2006).
Fletcher, T.M., Sun, D., Salazar, M. & Hurley, L.H. Effect of DNA secondary structure on human telomerase activity. Biochemistry 37, 5536–5541 (1998).
Iida, K. et al. Fluorescent-ligand-mediated screening of G-quadruplex structures using a DNA microarray. Angew. Chem. Int. Ed. Engl. 52, 12052–12055 (2013).
Biffi, G., Tannahill, D., McCafferty, J. & Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 5, 182–186 (2013).
Uno, S., You, Z. & Masai, H. Purification of replication factors using insect and mammalian cell expression systems. Methods 57, 214–221 (2012).
Xu, D. et al. Rif1 provides a new DNA-binding interface for the Bloom syndrome complex to maintain normal replication. EMBO J. 29, 3140–3155 (2010).
Sukackaite, R. et al. Structural and biophysical characterization of murine Rif1 C terminus reveals high specificity for DNA cruciform structures. J. Biol. Chem. 289, 13903–13911 (2014).
Sengar, A., Hiddi, B. & Phan, A.T. Formation of G-quadruplexes in poly-G sequences: structure of a propeller-type parallel-stranded G-quadruplex formed by a G15 stretch. Biochemistry 53, 7718–7723 (2014).
Sabouri, N., Capra, J. & Zakian, V.A. The essential Schizosaccharomyces pombe Pfh1 DNA helicase promotes fork movement past G-quadruplex motifs to prevent DNA damage. BMC Biol. 12, 101 (2014).
Park, S. et al. Palmitoylation controls the dynamics of budding-yeast heterochromatin via the telomere-binding protein Rif1. Proc. Natl. Acad. Sci. USA 108, 14572–14577 (2011).
Cayrou, C. et al. New insights into replication origin characteristics in metazoans. Cell Cycle 11, 658–667 (2012).
Valton, A.L. et al. G4 motifs affect origin positioning and efficiency in two vertebrate replicators. EMBO J. 33, 732–746 (2014).
Shi, T. et al. Rif1 and Rif2 shape telomere function and architecture through multivalent Rap1 interactions. Cell 153, 1340–1353 (2013).
Katou, Y. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083 (2003).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Nicol, J.W., Helt, G.A., Blanchard, S.G., Raja, A. & Loraine, A.E. The Integrated Genome Browser: free software for distribution and exploration of genome-scale datasets. Bioinformatics 25, 2730–2731 (2009).
Machanick, P. & Bailey, T.L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Zheng, K.W., Chen, Z., Hao, Y.H. & Tan, Z. Molecular crowding creates an essential environment for the formation of stable -quadruplexe G-quadruplexes in long double-stranded DNA. Nucleic Acids Res. 38, 327–338 (2010).
Martin, C.D. et al. A simple vector system to improve performance and utilisation of recombinant antibodies. BMC Biotechnol. 6, 46 (2006).
Nishitani, H., Lygerou, Z., Nishimoto, T. & Nurse, P. The Cdt1 protein is required to license DNA for replication in fission yeast. Nature 404, 625–628 (2000).
Acknowledgements
We thank J. Horiuchi for critical reading of the manuscript and useful suggestions. We would like to thank K. Ohta and the members of his laboratory for assistance in ChIP-seq analyses, which were supported by Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research on Priority Areas ('non-coding DNA'). This work was supported by JSPS KAKENHI (Grant-in-Aid for Scientific Research (A) (grant nos. 23247031 and 26251004) and Grant-in-Aid for Scientific Research on Priority Areas ('non-coding RNA' and 'Genome Adaptation'; grant nos. 24114520 and 25125724, respectively) to H.M. and Grant-in-Aid for Scientific Research (C) (grant no. 24570205) to S.M., Grant-in-Aid for Scientific Research (A) (grant no. 15H02369) to K.S. and Grant-in-Aid for Young Scientists (B) (grant no. 26870575) to N. Kono) and by the Naito Foundation Continuation Subsidy for Outstanding Projects (to H.M.). We thank members of our laboratory for helpful discussion.
Author information
Authors and Affiliations
Contributions
Y.K. conducted ChIP-chip and ChIP-seq analyses, mutant construction and informatics analyses of the binding sequences, and designed primers for amplification of Rif1BSs. S.M. assisted in strain construction and characterization, data analysis and manuscript preparation. R.F. expressed and purified the full-length fission yeast Rif1 protein and assisted in plasmid construction. N. Kono generated the three-dimensional (3D) chromosome model and analyzed 3D locations of Rif1BS and origins. C.R.-G. and K.M. conducted informatics analyses of Rif1BSs and Rif1CSs. K.I. and K.N. provided L1BOD-7OTD and related information. K.S. supported the research. H.M. and N. Kakusho conducted in vitro analyses of the G4-like nature of Rif1BS and binding of Rif1 to these structures. Y.K. and H.M. conceived and designed the experiments and wrote the paper. S.M. and C.R.-G. provided input regarding the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 ChIP-seq data of Rif1 on the entire fission yeast chromosomes in wild type, taz1∆, and rap1∆ cells at G1 phase.
Thirty-five Rif1BS are indicated. Vertical axes show pileup aligned read distributions. The entire chromosome III is early-replicating and there is only one Rif1BS on this chromosome. Note that telomere regions are not shown.
Supplementary Figure 2 Correlation between the numbers of Rif1CSs and Rif1 binding strength.
(a) Peak intensities in ChIP-Seq data of Rif1BS are compared with regard to the numbers of Rif1CS (at least two motifs, one motif and no motif). For each position indicated, peaks and mocks (ChIP in the non-tagged strain) are shown in upper and lower panels, respectively. The positive and negative controls of Rif1-ChIP are telomere repeats and centromere dg or ars2004 sites, respectively. (b) Quantification of Rif1-binding at the indicated locations by ChIP-qPCR. Efficiency of immunoprecipitation (%: ChIP/Input) is shown (mean of three independent cell cultures; error bar, s.e.m).
Supplementary Figure 3 Features of two representative Rif1BSs investigated in this study.
ChIP-Seq profiles of Rif1 binding (in the wild-type), Mcm4 binding (in hsk1-89) and BrdU-incorporation in the presence of hydroxyurea (in the wild type and rif1∆) are shown. Vertical axes show pileup aligned reads. The bottom drawing shows expansion of a part of the genome region. Rif1BS and Mcm4BS are shown by green and blue, respectively. The genes and non-coding RNAs are shown by black and light blue bars, respectively. The sequences containing Rif1CS (colored residues) are also shown at the bottom.
Supplementary Figure 4 Statistical analyses of correlation between the features of Rif1BSs and the nature of replication origins.
(a) Rif1 binding sites and replication efficiency in the wild-type and rif1∆ cells across the chromosome I. Upper, Rif1 ChIP-Seq peaks on the chromosome I and the locations of Rif1BS shown by vertical blue bars. Lower, origin usage was determined by calculating fractions of active origins (out of potential origins) at each location (in 100 kb windows and 10 kb increment) in the presence of HU (based on the data in ref. 7) Blue, wild-type; red, rif1∆. Rif1BS, indicated by dotted vertical lines, are enriched in the regions where origin usage efficiency is low in the wild-type but increased in rif1∆. (b) The density profile of BrdU-IP/Input log2-ratio in the 1 kb segments centered around the 90 Rif1BSs determined by ChIP-Seq. Regions near Rif1BS are generally characterized by low origin usage efficiency in the wild-type (left), which significantly increased in rif1∆ (right). (c–f) Analyses on the distance between Rif1BS and origins. Distances from Rif1BS to closest origins (c,e) and those from origins to closest Rif1BS (d,f) were calculated. The entire ninety potential Rif1BSs were included for calculation (c,d), or they were split into two groups, one with two or more CS (Rif1-twoCS; 35 sites; left of e,f) and the other with only one or no CS (Rif1-woCS; 55 sites; right of e and f), and calculation was conducted for both groups separately. Origins were obtained from ref.7. In both analyses, Rif1BSs were close to late origins and especially to LE ori than to EE or EL ori, and this trend was more evident for Rif1BS-twoCS than Rif1BS-woCS. (g) 34 Rif1BS (excluding the one in the subtelomere segment) located on chromosome arms projected onto 3D model of the fission yeast24. The positions of the three chromosomes are indicated in the upper panels (Chr. I, II, and III). Red spheres, blue and green bars are Rif1BS, LE ori and LL ori, respectively. The projected model was drawn by PyMol, a protein visualization software (www.pymol.org).
Supplementary Figure 5 Binding of L1BOD-7OTD to G-quadruplex DNA and its stabilization.
A 40-mer oligonucleotide containing a typical G4 sequence (4G) or its mutant version (0G), as shown below the panels, was either untreated (–) or heat-treated (+) and mixed with L1BOD-7OTD of varied concentrations, as shown. The samples were run on 12% PAGE (1×TBE, 50 mM KCl and 10% glycerol) and UV-irradiated at 254 nm (for detection of the compound, upper). The same gel was stained with ethidium bromide, and DNA bands were detected at 302 nm (lower). L1BOD-7OTD bound specifically to the heat-treated oligonucleotide DNA but not to untreated DNA or mutant DNA. The amount of G4 DNA (shifted DNA in a complex with the compound) increased with addition of more L1BOD-7OTD. This probably reflects the ability of L1BOD-7OTD to stabilize G4 structures.
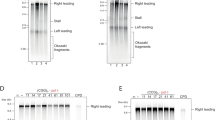
Supplementary Figure 6 Analyses of Rif1BS DNAs after heat denaturation-renaturation for potential formation of G4 conformation.
Thirty-three remaining Rif1BS was amplified and DNA fragments were isolated. DNA fragments (0.3 pmole) were either untreated (–) or heat-treated (+) and run on 8% PAGE (1× TBE, 40% PEG200 and 50 mM KCl), and DNA was detected by cybergreen.
Supplementary Figure 7 Sequences of Rif1BS-derived oligonucleotides used for gel shift, DMS, S1 nuclease and polymerase stop analyses; purification of Rif1 protein.
(a) Sequences of Rif1BS-containing DNA fragments (Rif1BSI:2663kb and Rif1BSII:4255kb) used for analyses. G-stretches or G-tract are indicated in orange. The underlined residues indicate the sequences for single-stranded oligonucleotides used for the analyses. (b) Purification of fission yeast Rif1 protein. Fission yeast Rif1 protein (1400 amino acids) were expressed on a mammalian expression vector with N-terminal 6× His tag and C-terminal 3× FLAG tag. After transfection into 15 15cm-plates of 293T cells, cells were harvested and Rif1 protein was purified by consecutive steps of anti-FLAG antibody and nickel affinity columns33. Peak fractions from the nickel column were pooled and further purified by monoQ column. Eluted fractions were analyzed on 5-20% gradient SDS-PAGE, and proteins were detected by silver staining. Fraction 13 was used for gel shift assays. The full-length Rif1 is shown by an arrow. An additional band at around 70 kDa is derived from the C-terminal segment, which can associate with the full length Rif1.
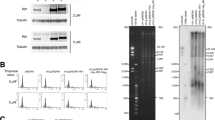
Supplementary Figure 8 Localization of the fission yeast Rif1 protein in nuclease and salt-insoluble fractions.
Fission yeast cells in which the endogenous Rif1 gene was FLAG-tagged at its C-terminus were fractionated. (a) A scheme for cellular fractionation of fission yeast cells. (b) Each fraction obtained in a was analyzed on SDS-PAGE and proteins indicated were detected by western blotting.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–4 and Supplementary Note (PDF 1845 kb)
Supplementary Data Set 1
Uncropped images (PDF 23884 kb)
Rights and permissions
About this article
Cite this article
Kanoh, Y., Matsumoto, S., Fukatsu, R. et al. Rif1 binds to G quadruplexes and suppresses replication over long distances. Nat Struct Mol Biol 22, 889–897 (2015). https://doi.org/10.1038/nsmb.3102
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3102
This article is cited by
-
Rif1 restrains the rate of replication origin firing in Xenopus laevis
Communications Biology (2023)
-
Dynamic alternative DNA structures in biology and disease
Nature Reviews Genetics (2023)
-
Bioactive nutraceuticals as G4 stabilizers: potential cancer prevention and therapy—a critical review
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)
-
Shedding light on the base-pair opening dynamics of nucleic acids in living human cells
Nature Communications (2022)
-
Chemical targeting of G-quadruplexes in telomeres and beyond for molecular cancer therapeutics
The Journal of Antibiotics (2021)