Abstract
Argonautes and their small-RNA cofactors form the core effectors of ancient and diverse gene-silencing mechanisms whose roles include regulation of gene expression and defense against foreign genetic elements. Although Argonautes generally act within multisubunit complexes, what governs their assembly into these machineries is not well defined. Here, we show that loading of small RNAs onto Argonaute is a checkpoint for Argonaute's association with conserved GW-protein components of silencing complexes. We demonstrate that the Argonaute small interfering RNA chaperone (ARC) complex mediates loading of small RNAs onto Ago1 in Schizosaccharomyces pombe and that deletion of its subunits, or mutations in Ago1 that prevent small-RNA loading, abolish the assembly of the GW protein–containing RNA-induced transcriptional silencing (RITS) complex. Our studies uncover a mechanism that ensures that Argonaute loading precedes RITS assembly and thereby averts the formation of inert and potentially deleterious complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Ghildiyal, M. & Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108 (2009).
Malone, C.D. & Hannon, G.J. Small RNAs as guardians of the genome. Cell 136, 656–668 (2009).
Moazed, D. Small RNAs in transcriptional gene silencing and genome defence. Nature 457, 413–420 (2009).
Olovnikov, I., Chan, K., Sachidanandam, R., Newman, D.K. & Aravin, A.A. Bacterial Argonaute samples the transcriptome to identify foreign DNA. Mol. Cell 51, 594–605 (2013).
Hutvagner, G. & Simard, M.J. Argonaute proteins: key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 9, 22–32 (2008).
Jinek, M. & Doudna, J.A. A three-dimensional view of the molecular machinery of RNA interference. Nature 457, 405–412 (2009).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nat. Rev. Genet. 14, 447–459 (2013).
Hammond, S.M., Bernstein, E., Beach, D. & Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–296 (2000).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Colmenares, S.U., Buker, S.M., Buhler, M., Dlakić, M. & Moazed, D. Coupling of double-stranded RNA synthesis and siRNA generation in fission yeast RNAi. Mol. Cell 27, 449–461 (2007).
Reinhart, B.J. & Bartel, D.P. Small RNAs correspond to centromere heterochromatic repeats. Science 297, 1831 (2002).
Yu, R., Jih, G., Iglesias, N. & Moazed, D. Determinants of heterochromatic siRNA biogenesis and function. Mol. Cell 53, 262–276 (2014).
Buker, S.M. et al. Two different Argonaute complexes are required for siRNA generation and heterochromatin assembly in fission yeast. Nat. Struct. Mol. Biol. 14, 200–207 (2007).
Matranga, C., Tomari, Y., Shin, C., Bartel, D.P. & Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Rand, T.A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Bayne, E.H. et al. Stc1: a critical link between RNAi and chromatin modification required for heterochromatin integrity. Cell 140, 666–677 (2010).
Gerace, E.L., Halic, M. & Moazed, D. The methyltransferase activity of Clr4Suv39h triggers RNAi independently of histone H3K9 methylation. Mol. Cell 39, 360–372 (2010).
Hong, E.-J.E., Villén, J., Gerace, E.L., Gygi, S.P. & Moazed, D.A. Cullin E3 ubiquitin ligase complex associates with Rik1 and the Clr4 histone H3-K9 methyltransferase and is required for RNAi-mediated heterochromatin formation. RNA Biol. 2, 106–111 (2005).
Volpe, T.A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Zhang, K., Mosch, K., Fischle, W. & Grewal, S.I.S. Roles of the Clr4 methyltransferase complex in nucleation, spreading and maintenance of heterochromatin. Nat. Struct. Mol. Biol. 15, 381–388 (2008).
Bernard, P. et al. Requirement of heterochromatin for cohesion at centromeres. Science 294, 2539–2542 (2001).
Ellermeier, C. et al. RNAi and heterochromatin repress centromeric meiotic recombination. Proc. Natl. Acad. Sci. USA 107, 8701–8705 (2010).
Probst, A.V., Dunleavy, E. & Almouzni, G. Epigenetic inheritance during the cell cycle. Nat. Rev. Mol. Cell Biol. 10, 192–206 (2009).
Azevedo, J., Cooke, R. & Lagrange, T. Taking RISCs with Ago hookers. Curr. Opin. Plant Biol. 14, 594–600 (2011).
Braun, J.E., Huntzinger, E. & Izaurralde, E. The role of GW182 proteins in miRNA-mediated silencing. Adv. Exp. Med. Biol. 768, 147–163 (2013).
El-Shami, M. et al. Reiterated WG/GW motifs form functionally and evolutionarily conserved ARGONAUTE-binding platforms in RNAi-related components. Genes Dev. 21, 2539–2544 (2007).
Till, S. et al. A conserved motif in Argonaute-interacting proteins mediates functional interactions through the Argonaute PIWI domain. Nat. Struct. Mol. Biol. 14, 897–903 (2007).
Braun, J.E., Huntzinger, E., Fauser, M. & Izaurralde, E. GW182 proteins directly recruit cytoplasmic deadenylase complexes to miRNA targets. Mol. Cell 44, 120–133 (2011).
Chekulaeva, M. et al. miRNA repression involves GW182-mediated recruitment of CCR4–NOT through conserved W-containing motifs. Nat. Struct. Mol. Biol. 18, 1218–1226 (2011).
Fabian, M.R. et al. miRNA-mediated deadenylation is orchestrated by GW182 through two conserved motifs that interact with CCR4–NOT. Nat. Struct. Mol. Biol. 18, 1211–1217 (2011).
Partridge, J.F. et al. Functional separation of the requirements for establishment and maintenance of centromeric heterochromatin. Mol. Cell 26, 593–602 (2007).
Pontier, D. et al. NERD, a plant-specific GW protein, defines an additional RNAi-dependent chromatin-based pathway in Arabidopsis. Mol. Cell 48, 121–132 (2012).
Behm-Ansmant, I. et al. mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes Dev. 20, 1885–1898 (2006).
Baillat, D. & Shiekhattar, R. Functional dissection of the human TNRC6 (GW182-related) family of proteins. Mol. Cell. Biol. 29, 4144–4155 (2009).
Giner, A., Lakatos, L., García-Chapa, M., López-Moya, J.J. & Burgyán, J. Viral protein inhibits RISC activity by argonaute binding through conserved WG/GW motifs. PLoS Pathog. 6, e1000996 (2010).
Eulalio, A., Helms, S., Fritzsch, C., Fauser, M. & Izaurralde, E. A C-terminal silencing domain in GW182 is essential for miRNA function. RNA 15, 1067–1077 (2009).
Liu, J., Valencia-Sanchez, M.A., Hannon, G.J. & Parker, R. MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat. Cell Biol. 7, 719–723 (2005).
Woolcock, K.J. et al. RNAi keeps Atf1-bound stress response genes in check at nuclear pores. Genes Dev. 26, 683–692 (2012).
Halic, M. & Moazed, D. Dicer-independent primal RNAs trigger RNAi and heterochromatin formation. Cell 140, 504–516 (2010).
Marasovic, M., Zocco, M. & Halic, M. Argonaute and Triman generate Dicer-independent priRNAs and mature siRNAs to initiate heterochromatin formation. Mol. Cell 52, 173–183 (2013).
Ma, J.-B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005).
Ma, J.-B., Ye, K. & Patel, D.J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004).
Schirle, N.T. & MacRae, I.J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Iida, T., Nakayama, J. & Moazed, D. siRNA-mediated heterochromatin establishment requires HP1 and is associated with antisense transcription. Mol. Cell 31, 178–189 (2008).
Irvine, D.V. et al. Argonaute slicing is required for heterochromatic silencing and spreading. Science 313, 1134–1137 (2006).
Martinez, N.J. & Gregory, R.I. Argonaute2 expression is post-transcriptionally coupled to microRNA abundance. RNA 19, 605–612 (2013).
Smibert, P., Yang, J.-S., Azzam, G., Liu, J.-L. & Lai, E.C. Homeostatic control of Argonaute stability by microRNA availability. Nat. Struct. Mol. Biol. 20, 789–795 (2013).
Frank, F., Sonenberg, N. & Nagar, B. Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 465, 818–822 (2010).
Bühler, M., Spies, N., Bartel, D.P. & Moazed, D. TRAMP-mediated RNA surveillance prevents spurious entry of RNAs into the Schizosaccharomyces pombe siRNA pathway. Nat. Struct. Mol. Biol. 15, 1015–1023 (2008).
Gregory, R.I., Chendrimada, T.P., Cooch, N. & Shiekhattar, R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123, 631–640 (2005).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Maniataki, E. & Mourelatos, Z. A human, ATP-independent, RISC assembly machine fueled by pre-miRNA. Genes Dev. 19, 2979–2990 (2005).
Boland, A., Huntzinger, E., Schmidt, S., Izaurralde, E. & Weichenrieder, O. Crystal structure of the MID-PIWI lobe of a eukaryotic Argonaute protein. Proc. Natl. Acad. Sci. USA 108, 10466–10471 (2011).
Eulalio, A., Huntzinger, E. & Izaurralde, E. GW182 interaction with Argonaute is essential for miRNA-mediated translational repression and mRNA decay. Nat. Struct. Mol. Biol. 15, 346–353 (2008).
Bähler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951 (1998).
Leeds, P., Peltz, S.W., Jacobson, A. & Culbertson, M.R. The product of the yeast UPF1 gene is required for rapid turnover of mRNAs containing a premature translational termination codon. Genes Dev. 5, 2303–2314 (1991).
Bühler, M., Verdel, A. & Moazed, D. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125, 873–886 (2006).
Huang, J. & Moazed, D. Association of the RENT complex with nontranscribed and coding regions of rDNA and a regional requirement for the replication fork block protein Fob1 in rDNA silencing. Genes Dev. 17, 2162–2176 (2003).
Acknowledgements
We thank members of the Moazed laboratory for helpful discussions; E. Gerace, M. Halic, N. Iglesias and J. Xiol for advice on protocols; M. Halic and R. Yu for scripts and advice on bioinformatic analysis; C. Centrella for technical assistance; and E. Egan, N. Iglesias, R. Jain, J. Xiol and R. Yu for critical reading of the manuscript. This work was supported by the US National Science Foundation Graduate Research Fellowship Program (D.H.) and the US National Institutes of Health grant R01 GM072805 (D.M.). D.M. is supported as a Howard Hughes Medical Institute Investigator. D.H. dedicates this paper to the memory of his beloved father, George Holoch, who died on 9 April 2013.
Author information
Authors and Affiliations
Contributions
D.H. and D.M. designed the study. D.H. performed all experiments and bioinformatic analysis. D.H. and D.M. analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Ago1 associates with Arb1 and Arb2 independently of Tas3.
Mixture mass spectrometry analysis of the indicated FLAG purifications. Uniquely detected peptides, including overlapping peptides, are reported for each protein in each purification. Percent coverages shown are calculated by dividing the total number of amino acid residues represented in any peptide for a given protein in a given purification by the protein’s total length.
Supplementary Figure 2 Ago1 D580A, Ago1 F276A R773E, Ago1 F276A and Ago1 R773E are null-mutant proteins whose small RNA–binding activity correlates with their assembly into RITS.
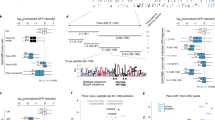
(a) Schematic illustrating the ectopic insertion of 3xFLAG-tagged alleles of ago1 with endogenous promoter and terminator sequences near the trp1+ locus on chromosome 2. The native ago1+ locus harbors either a wild-type untagged allele or a deletion. (b) Tenfold serial dilutions of otr1R::ura4+ pericentromeric silencing reporter cells of the indicated genotypes, plated on non-selective medium or medium containing 5-FOA. (c) Northern blot analysis of dg siRNAs in total small RNA fractions and RNA immunoprecipitated with wild-type and mutant 3xFLAG-Ago1 proteins, in cells also expressing untagged wild-type Ago1, and Western blot analysis of input extracts and FLAG-immunoprecipitated material. (d) Western blot analysis of co-immunoprecipitation experiment to assay association of Tas3 with the indicated Ago1 proteins. (e) Western blot analysis of co-immunoprecipitation experiment to assay association of Arb1 with the indicated Ago1 proteins.
Supplementary Figure 3 RITS and ARC bind populations of pericentromeric small RNAs with similar sequences.
Tracks showing the normalized numbers of reads mapping to the dg and dh repeats flanking the centromere of chromosome 1, for small RNAs co-purifying with Tas3-TAP (RITS) or TAP-Arb1 (ARC), excluding tRNAs.
Supplementary Figure 4 Arb2 is not required for in vitro loading of duplex small RNAs onto immunopurified Ago1.
(a) Western blot analysis of input fractions and IgG magnetic beads after one-step purification of RITS and ARC from dcr1Δ cells expressing TAP-tagged subunits Tas3 or Arb1 or no tagged protein. (b) Phosphorimager scan of a non-denaturing polyacrylamide gel showing the 5’-end-labeled single-stranded and annealed duplex small RNAs used in in vitro binding assays. (c) Western blot analysis of the beads from FLAG purifications from the indicated cells, in aliquots equal to one-eighth of those used in the binding assay. (d) Phosphorimager scan of eluted RNAs after in vitro binding to immobilized 3xFLAG-Ago1 purified from the indicated wild-type and mutant cells. (e) Quantification by densitometry of the results shown in (d).
Supplementary Figure 5 Overexpressed Ago1 suppresses arb1Δ and arb2Δ not simply by rescuing protein stability and does not suppress other pericentromeric silencing mutants.
(a) Western blot analysis of total protein prepared from arb1Δ and arb2Δ cells expressing 3xFLAG-ago1 either from the endogenous locus or from an overexpression plasmid. Relative quantity of total protein loaded is indicated for each lane. Red pixels indicate saturated signal. (b) Western blot analysis of total protein prepared from cells of the indicated genotypes. Relative quantity of total protein loaded is indicated for each lane. (c) Tenfold serial dilutions of otr1R::ura4+ pericentromeric silencing reporter cells of the indicated genotypes, transformed with an empty vector (denoted with a “–”) or a 3xFLAG-ago1 overexpression plasmid (denoted with a red “+”), plated on non-selective medium or medium containing 5-FOA. Similarly, tenfold serial dilutions of otr1R::ade6+ cells of the indicated genotypes plated on medium containing a limiting concentration of adenine.
Supplementary Figure 6 Ago1 L317A protein does not complement ago1Δ but does not show a significant reduction in loading of pericentromeric siRNAs.
(a) Tenfold serial dilutions of otr1R::ura4+ or imr1R::ura4+ pericentromeric silencing reporter cells of the indicated genotypes plated on non-selective medium or medium containing 5-FOA. (b) Western blot analysis of whole cell extracts and FLAG immunoprecipitates prepared from wild-type cells transformed with the indicated 3xFLAG-ago1 overexpression plasmids, and Northern blot analysis of RNA extracted from each immunoprecipitated sample.
Supplementary Figure 7 ago1 D580A and ago1 F276A R773E are dominant-negative alleles when overexpressed.
Tenfold serial dilutions of otr1R::ura4+ pericentromeric silencing reporter cells of the indicated genotypes, transformed with an empty vector or a 3xFLAG-ago1 overexpression plasmid (wild-type or mutant as noted), plated on non-selective medium or medium containing 5-FOA.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–4 and Supplementary Note (PDF 738 kb)
Supplementary Data Set 1
Uncropped gel images (PDF 14944 kb)
Rights and permissions
About this article
Cite this article
Holoch, D., Moazed, D. Small-RNA loading licenses Argonaute for assembly into a transcriptional silencing complex. Nat Struct Mol Biol 22, 328–335 (2015). https://doi.org/10.1038/nsmb.2979
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2979
This article is cited by
-
Argonaute bypasses cellular obstacles without hindrance during target search
Nature Communications (2019)
-
Chromatin-associated RNAs as facilitators of functional genomic interactions
Nature Reviews Genetics (2019)
-
RNAi-dependent heterochromatin assembly in fission yeast Schizosaccharomyces pombe requires heat-shock molecular chaperones Hsp90 and Mas5
Epigenetics & Chromatin (2018)
-
Unique roles for histone H3K9me states in RNAi and heritable silencing of transcription
Nature (2017)
-
New methods in the diagnosis of cancer and gene therapy of cancer based on nanoparticles
Cancer Gene Therapy (2017)