Abstract
Eukaryotic translation initiation requires cooperative assembly of a large protein complex at the 40S ribosomal subunit. We have resolved a budding yeast initiation complex by cryo-EM, allowing placement of prior structures of eIF1, eIF1A, eIF3a, eIF3b and eIF3c. Our structure highlights differences in initiation-complex binding to the ribosome compared to that of mammalian eIF3, demonstrates a direct contact between eIF3j and eIF1A and reveals the network of interactions between eIF3 subunits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hinnebusch, A.G. & Lorsch, J.R. Cold Spring Harb. Perspect. Biol. 4, a011544 (2012).
Jivotovskaya, A.V., Valasek, L., Hinnebusch, A.G. & Nielsen, K.H. Mol. Cell. Biol. 26, 1355–1372 (2006).
Sun, C. et al. Proc. Natl. Acad. Sci. USA 108, 20473–20478 (2011).
Zhou, M. et al. Proc. Natl. Acad. Sci. USA 105, 18139–18144 (2008).
Hashem, Y. et al. Cell 153, 1108–1119 (2013).
Hussain, T. et al. Cell 159, 597–607 (2014).
Erzberger, J.P. et al. Cell 158, 1123–1135 (2014).
Sokabe, M., Fraser, C.S. & Hershey, J.W. Nucleic Acids Res. 40, 905–913 (2012).
Sokabe, M. & Fraser, C.S. J. Biol. Chem. 289, 31827–31836 (2014).
Weisser, M., Voigts-Hoffmann, F., Rabl, J., Leibundgut, M. & Ban, N. Nat. Struct. Mol. Biol. 20, 1015–1017 (2013).
Lomakin, I.B. & Steitz, T.A. Nature 500, 307–311 (2013).
Querol-Audi, J. et al. Structure 21, 920–928 (2013).
Pisarev, A.V. et al. EMBO J. 27, 1609–1621 (2008).
Kouba, T. et al. PLoS ONE 7, e40464 (2012).
Munzarová, V. et al. PLoS Genet. 7, e1002137 (2011).
Szamecz, B. et al. Genes Dev. 22, 2414–2425 (2008).
Roy, B. et al. RNA 16, 748–761 (2010).
Chiu, W.L. et al. Mol. Cell. Biol. 30, 4415–4434 (2010).
Phan, L., Schoenfeld, L.W., Valásek, L., Nielsen, K.H. & Hinnebusch, A.G. EMBO J. 20, 2954–2965 (2001).
Dong, Z., Qi, J., Peng, H., Liu, J. & Zhang, J.T. J. Biol. Chem. 288, 27951–27959 (2013).
Liu, Y. et al. Structure 22, 923–930 (2014).
Cuchalová, L. et al. Mol. Cell. Biol. 30, 4671–4686 (2010).
Herrmannová, A. et al. Nucleic Acids Res. 40, 2294–2311 (2012).
Khoshnevis, S., Neumann, P. & Ficner, R. PLoS ONE 5, e12784 (2010).
Elantak, L. et al. J. Mol. Biol. 396, 1097–1116 (2010).
Fraser, C.S., Berry, K.E., Hershey, J.W. & Doudna, J.A. Mol. Cell 26, 811–819 (2007).
Nielsen, K.H. et al. Mol. Cell. Biol. 26, 2984–2998 (2006).
Valásek, L. et al. EMBO J. 20, 891–904 (2001).
Valásek, L., Hasek, J., Trachsel, H., Imre, E.M. & Ruis, H. J. Biol. Chem. 274, 27567–27572 (1999).
Valásek, L., Nielsen, K.H., Zhang, F., Fekete, C.A. & Hinnebusch, A.G. Mol. Cell. Biol. 24, 9437–9455 (2004).
Pettersen, E.F. et al. J. Comput. Chem. 25, 1605–1612 (2004).
Rabl, J., Leibundgut, M., Ataide, S.F., Haag, A. & Ban, N. Science 331, 730–736 (2011).
Cabrita, L.D., Dai, W. & Bottomley, S.P. BMC Biotechnol. 6, 12 (2006).
Leitner, A. et al. Mol. Cell. Proteomics 9, 1634–1649 (2010).
Li, X. et al. Nat. Methods 10, 584–590 (2013).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. J. Struct. Biol. 128, 82–97 (1999).
Scheres, S.H.W. J. Struct. Biol. 180, 519–530 (2012).
Kucukelbir, A., Sigworth, F.J. & Tagare, H.D. Nat. Methods 11, 63–65 (2014).
Ben-Shem, A. et al. Science 334, 1524–1529 (2011).
Kelley, L.A. & Sternberg, M.J.E. Nat. Protoc. 4, 363–371 (2009).
Acknowledgements
We would like to thank the ETH Zürich Scientific Center for Optical and Electron Microscopy for access to EM equipment and are indebted to P. Tittmann for technical support. C.H.S.A. was supported by an ETH Zürich postdoctoral fellowship and an European Molecular Biology Organization (EMBO) long-term fellowship and J.P.E. by an EMBO long-term fellowship. This work was supported by the Swiss National Science Foundation (SNSF), the National Center of Excellence in Research (of Switzerland), the Structural Biology, and RNA and Disease programs of the SNSF and European Research Council grant 250071 (to N.B.) under the European Community's Seventh Framework Programme.
Author information
Authors and Affiliations
Contributions
N.B. and C.H.S.A. designed the study. C.H.S.A., J.P.E. and T.S. prepared samples and performed biochemical experiments. C.H.S.A. and D.B. collected electron micrographs and calculated reconstructions. C.H.S.A., D.B., J.P.E. and N.B. interpreted the structures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Local and global resolution statistics for the 40S–eIF1–eIF1A–eIF3–eIF3j complex.
Fourier shell correlation for 40S–eIF1–eIF1A–eIF3–eIF3j calculated between two independently refined half sets; resolution cut-off indicated by the red line (left panel). The “gold-standard” Fourier shell correlation was determined, and resolution estimated, according to the method of S. Scheres37. The surface of the 40S–eIF1–eIF1A–eIF3–eIF3j reconstruction low-pass filtered at global resolution (6.47 Å) is shown coloured according to the local resolution (centre and right panels – 5 Å shown in blue through to 15 Å shown in red). The mean local resolution was 12.9 Å for eIF3ac, 9.5 Å for eIF3a CTD, 10.3 Å for eIF3b WD40, 14.7 Å for eIF3gi, 14.0 Å for eIF3b RRM and 10.8 Å for eIF1–1a–3j. Local resolution was determined using the implementation of Kucukelbir and colleagues: ResMap38.
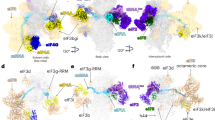
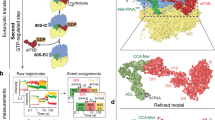
Supplementary Figure 2 Overview comparing the rabbit 43S preinitiation complex, budding-yeast eIF3–eIF1–eIF1A complex and partial yeast 48S initiation complex.
Cryo-EM structures of (left) the rabbit 43S5, (centre) the budding yeast eIF3–eIF1–eIF1A initiation complex and (right) the partial yeast 48S6 are shown with accompanying fitted or directly interpreted structures. A composite between the modelled PCI-MPN core7, the deposited structure of the partial yeast 48S and our eIF3b fit is shown docked into the 43S (for which only a fit of the PCI-MPN core is currently available) to facilitate comparison. Each structure is shown from the interface (top) and solvent (bottom) sides; RNA is shown in grey, ribosomal proteins in yellow, the ternary complex (TC) in black, eIF1 in red, eIF1A in orange, eIF3a in blue, 3b in green, 3c in purple, 3e in dark green, 3f in turquoise, 3h in deep orange, 3k in spring green, 3l in light yellow and 3m in pink. The trace of the C-terminal tail of eIF3a is shown by a blue dotted line in, while difference density assigned to eIF3j is shown in magenta.
Supplementary Figure 3 Cryo-EM classification schema for the 40S–eIF1–eIF1A–eIF3–eIF3j complex.
The three-dimensional reconstructions from each round of classification are shown from the solvent exposed surface of the 40S subunit. The number of particles involved in each classification, and the flow of particles between stages of classification are shown. The last steps of classification involved the occupied particles from both datasets and several different parameter sets were trialed in order to minimize the presence of unoccupied, or lower resolution, particles. Magenta boxes indicate classes from which particles were retained earlier in classification.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Note (PDF 4289 kb)
Rights and permissions
About this article
Cite this article
Aylett, C., Boehringer, D., Erzberger, J. et al. Structure of a Yeast 40S–eIF1–eIF1A–eIF3–eIF3j initiation complex. Nat Struct Mol Biol 22, 269–271 (2015). https://doi.org/10.1038/nsmb.2963
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2963
This article is cited by
-
The molecular basis of translation initiation and its regulation in eukaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Molecular basis for recognition and deubiquitination of 40S ribosomes by Otu2
Nature Communications (2023)
-
Eukaryotic translation initiation factor 3 subunit B promotes head and neck cancer via CEBPB translation
Cancer Cell International (2022)
-
Knockdown of eIF3a attenuated cell growth in K1 human thyroid cancer cells
Genes & Genomics (2021)
-
The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs
Nature (2018)