Abstract
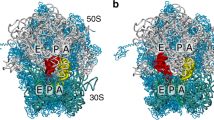
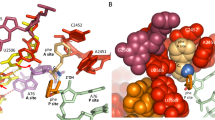
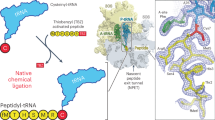
During peptide-bond formation on the ribosome, the α-amine of an aminoacyl-tRNA attacks the ester carbonyl carbon of a peptidyl-tRNA to yield a peptide lengthened by one amino acid. Although the ribosome's contribution to catalysis is predominantly entropic, the lack of high-resolution structural data for the complete active site in complex with full-length ligands has made it difficult to assess how the ribosome might influence the pathway of the reaction. Here, we present crystal structures of preattack and postcatalysis complexes of the Thermus thermophilus 70S ribosome at ~2.6-Å resolution. These structures reveal a network of hydrogen bonds along which proton transfer could take place to ensure the concerted, rate-limiting formation of a tetrahedral intermediate. We propose that, unlike earlier models, the ribosome and the A-site tRNA facilitate the deprotonation of the nucleophile through the activation of a water molecule.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Voorhees, R.M., Weixlbaumer, A., Loakes, D., Kelley, A.C. & Ramakrishnan, V. Insights into substrate stabilization from snapshots of the peptidyl transferase center of the intact 70S ribosome. Nat. Struct. Mol. Biol. 16, 528–533 (2009).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Schmeing, T.M., Huang, K.S., Kitchen, D.E., Strobel, S.A. & Steitz, T.A. Structural insights into the roles of water and the 2′ hydroxyl of the P site tRNA in the peptidyl transferase reaction. Mol. Cell 20, 437–448 (2005).
Schmeing, T.M., Huang, K.S., Strobel, S.A. & Steitz, T.A. An induced-fit mechanism to promote peptide bond formation and exclude hydrolysis of peptidyl-tRNA. Nature 438, 520–524 (2005).
Schmeing, T.M. et al. A pre-translocational intermediate in protein synthesis observed in crystals of enzymatically active 50S subunits. Nat. Struct. Biol. 9, 225–230 (2002).
Nagle, J.F. & Morowitz, H.J. Molecular mechanisms for proton transport in membranes. Proc. Natl. Acad. Sci. USA 75, 298–302 (1978).
Trobro, S. & Aqvist, J. Role of ribosomal protein L27 in peptidyl transfer. Biochemistry 47, 4898–4906 (2008).
Katunin, V.I., Muth, G.W., Strobel, S.A., Wintermeyer, W. & Rodnina, M.V. Important contribution to catalysis of peptide bond formation by a single ionizing group within the ribosome. Mol. Cell 10, 339–346 (2002).
Kuhlenkoetter, S., Wintermeyer, W. & Rodnina, M.V. Different substrate-dependent transition states in the active site of the ribosome. Nature 476, 351–354 (2011).
Wallin, G. & Aqvist, J. The transition state for peptide bond formation reveals the ribosome as a water trap. Proc. Natl. Acad. Sci. USA 107, 1888–1893 (2010).
Hiller, D.A., Singh, V., Zhong, M. & Strobel, S.A. A two-step chemical mechanism for ribosome-catalysed peptide bond formation. Nature 476, 236–239 (2011).
Carrasco, N., Hiller, D.A. & Strobel, S.A. Minimal transition state charge stabilization of the oxyanion during peptide bond formation by the ribosome. Biochemistry 50, 10491–10498 (2011).
Maguire, B.A., Beniaminov, A.D., Ramu, H., Mankin, A.S. & Zimmermann, R.A. A protein component at the heart of an RNA machine: the importance of protein l27 for the function of the bacterial ribosome. Mol. Cell 20, 427–435 (2005).
Hammes-Schiffer, S. & Stuchebrukhov, A.A. Theory of coupled electron and proton transfer reactions. Chem. Rev. 110, 6939–6960 (2010).
Kingery, D.A. et al. An uncharged amine in the transition state of the ribosomal peptidyl transfer reaction. Chem. Biol. 15, 493–500 (2008).
Zaher, H.S., Shaw, J.J., Strobel, S.A. & Green, R. The 2′-OH group of the peptidyl-tRNA stabilizes an active conformation of the ribosomal PTC. EMBO J. 30, 2445–2453 (2011).
Huang, Y. & Sprinzl, M. Peptide bond formation on the ribosome: the role of the 2′-OH group on the terminal adenosine of peptidyl-tRNA and of the length of nascent peptide chain. Angew. Chem. Int. Ed. Engl. 50, 7287–7289 (2011).
Koch, M., Huang, Y. & Sprinzl, M. Peptide-bond synthesis on the ribosome: no free vicinal hydroxy group required on the terminal ribose residue of peptidyl-tRNA. Angew. Chem. Int. Edn Engl. 47, 7242–7245 (2008).
Erlacher, M.D. et al. Efficient ribosomal peptidyl transfer critically relies on the presence of the ribose 2′-OH at A2451 of 23S rRNA. J. Am. Chem. Soc. 128, 4453–4459 (2006).
Lang, K., Erlacher, M., Wilson, D.N., Micura, R. & Polacek, N. The role of 23S ribosomal RNA residue A2451 in peptide bond synthesis revealed by atomic mutagenesis. Chem. Biol. 15, 485–492 (2008).
Weinger, J.S., Parnell, K.M., Dorner, S., Green, R. & Strobel, S.A. Substrate-assisted catalysis of peptide bond formation by the ribosome. Nat. Struct. Mol. Biol. 11, 1101–1106 (2004).
Youngman, E.M., Brunelle, J.L., Kochaniak, A.B. & Green, R. The active site of the ribosome is composed of two layers of conserved nucleotides with distinct roles in peptide bond formation and peptide release. Cell 117, 589–599 (2004).
Bieling, P., Beringer, M., Adio, S. & Rodnina, M.V. Peptide bond formation does not involve acid-base catalysis by ribosomal residues. Nat. Struct. Mol. Biol. 13, 423–428 (2006).
Polacek, N. et al. The critical role of the universally conserved A2602 of 23S ribosomal RNA in the release of the nascent peptide during translation termination. Mol. Cell 11, 103–112 (2003).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science 289, 905–920 (2000).
Armache, J.P. et al. Localization of eukaryote-specific ribosomal proteins in a 5.5-A cryo-EM map of the 80S eukaryotic ribosome. Proc. Natl. Acad. Sci. USA 107, 19754–19759 (2010).
Hofer, A., Bussiere, C. & Johnson, A.W. Mutational analysis of the ribosomal protein Rpl10 from yeast. J. Biol. Chem. 282, 32630–32639 (2007).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006).
Schmitt, E., Blanquet, S. & Mechulam, Y. Crystallization and preliminary X-ray analysis of Escherichia coli methionyl-tRNAMet(f) formyltransferase complexed with formyl-methionyl-tRNAMet(f). Acta Crystallogr. D Biol. Crystallogr. 55, 332–334 (1999).
Jünemann, R. et al. In vivo deuteration of transfer RNAs: overexpression and large-scale purification of deuterated specific tRNAs. Nucleic Acids Res. 24, 907–913 (1996).
Fraser, T.H. & Rich, A. Synthesis and aminoacylation of 3′-amino-3′-deoxy transfer RNA and its activity in ribosomal protein synthesis. Proc. Natl. Acad. Sci. USA 70, 2671–2675 (1973).
Korostelev, A., Trakhanov, S., Laurberg, M. & Noller, H.F. Crystal structure of a 70S ribosome-tRNA complex reveals functional interactions and rearrangements. Cell 126, 1065–1077 (2006).
Polikanov, Y.S., Blaha, G.M. & Steitz, T.A. How hibernation factors RMF, HPF, and YfiA turn off protein synthesis. Science 336, 915–918 (2012).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Byrne, R.T., Konevega, A.L., Rodnina, M.V. & Antson, A.A. The crystal structure of unmodified tRNAPhe from Escherichia coli. Nucleic Acids Res. 38, 4154–4162 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
We thank the staff at the Advanced Photon Source (beamline 24ID) and at the National Synchrotron Light Source (beamline X25) for help during data collection and the staff at the Richards Center at Yale University for computational support. We also thank P.B. Moore, S.A. Strobel and D.A. Hiller for critical reading of the manuscript, R.L. Grodzicki for preparation of the unmodified tRNAs, J. Lin, C. Mackereth and D. Dupuy for discussions and advice and members of the T.A.S. and C.A.I. laboratories for discussions. This work was supported by US National Institutes of Health grant GM022778 (T.A.S.) and by starting funds from the Fondation pour la Recherche Médicale, the Conseil Régional d'Aquitaine, the Institut National de la Santé et de la Recherche Médicale and the Centre National de la Recherche Scientifique (C.A.I.).
Author information
Authors and Affiliations
Contributions
C.A.I. and Y.S.P. devised experiments and prepared modified-tRNA substrates, Y.S.P. crystallized ribosomal complexes and performed crystallographic data collection and processing, C.A.I. and Y.S.P. analyzed the data, and C.A.I., Y.S.P. and T.A.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Experimental approach for enhancement of unbiased Fo – Fc maps.
a, c, Unbiased Fo-Fc maps of the Tth 70S post-catalysis complex at 2.55 Å resolution with (a) or without (c) bulk solvent correction applied to the active site of the ribosome. For clarity, maps are shown only for the terminal A76 residues of the tRNAs in both A- and P-sites. In all three panels, fitted models of a 3’-terminal A76 residue with an attached di-peptide in the A-site or a deacylated A76 residue in the P-site are shown only for reference, as they were not used for refinement and phase calculation. To prevent a specific region of the asymmetric unit from being included into the solvent mask during bulk solvent correction, a sphere of evenly spaced dummy atoms with zero occupancy (b) was used to cover the active site region. Any dummy atom located <2 Å away from any atom of the physical model was excluded. The resulting unbiased Fo–Fc map shown in (c) was calculated using phases derived from exactly the same model as in (a), but without bulk solvent correction applied to the active site.
Supplementary Figure 2 Electron density maps for the CPmn and CCPmn preattack complexes.
a, c, The chemical structures of CPmn and fMet-NH-tRNAiMet (a), and CCPmn and fMet-NH-tRNAiMet (c). b, d, Unbiased Fo-Fc electron density maps from co-crystallization experiments with the respective substrates: CPmn and fMet-NH-tRNAiMet at 2.9 Å resolution (contoured at +2.0σ) (b), and CCPmn and fMet-NH-tRNAiMet at 2.8 Å resolution (+2.0σ) (d). Bulk solvent correction was not applied to the active site region, as explained in Fig. S1, S2. The color scheme for the tRNA-ligands is the same as in Fig. 1. A-site bound tRNA analogs, CPmn and CCPmn, are colored in red.
Supplementary Figure 3 Comparison of the Tth 70S and Hma 50S preattack and postcatalysis structures and of the Tth 70S and Hma 50S structures in complex with short tRNA analogs.
a, Superimposed models of the Tth 70S and Hma 50S pre-attack complexes. In the Tth structure, Phe-NH-tRNAPhe is in the A-site and fMet-NH-tRNAiMet is in the P-site, whereas the Hma structure features CChPmn in the A-site and CCApcb in the P-site (PDB code 1VQN). Structural differences between the two models are within experimental error. Hma numbering is shown in brackets in this and subsequent panels. b, Superimposed models of the Tth 70S and Hma 50S post-catalysis complexes. The Tth structure contains fMet-Phe-NH-tRNAPhe in the A-site and deacylated tRNAiMet in the P-site, while the Hma structure has CCPmn-pcb and CCA in the A- and P-sites, respectively (PDB code 1KQS). In the latter structure, several ribosomal residues fail to undergo the movements induced by proper A-site substrate accommodation and residue A2602(A2637) is not held in place by the 3’-termini of the tRNAs, resulting in a conformation that would clash with the phosphate group of the P-site residue C74. The color guide is shown at the bottom of each panel. c, d, Superimposed models of our Tth 70S pre-attack (c) and post-catalysis (d) complexes and lower resolution Tth 70S complexes from an earlier study (PDB codes 2WDK/2WDL and 2WDG/2WDI for the pre- and post-catalysis structures, respectively). Differences between the new and old models are within experimental error. However, the amount of detail in the structures presented here is greater owing to a nearly 1-Å increase in resolution. e, f, Superimposed models of Tth 70S featuring full-length Phe-NH-tRNAPhe in the A-site with Tth 70S (e) or Hma 50S (f) in complex with the short aminoacyl-tRNA analogs CPmn and CCPmn. Structural differences between the active sites of the three Tth complexes shown in (e) are within experimental error, while residues U2506, G2583, U2584 and U2585 of the active site of Hma 50S featuring the CPmn analog in the A-site (PDB code 1VQ6) fail to undergo the movements elicited by proper A-site binding (f). Thus, all of the models shown in (e) correspond to the induced state, with a fully accommodated A-site substrate, whereas CPmn and CCPmn complexes of Hma 50S shown in (f) are in uninduced and induced states, respectively. The observed conformation of the active site for each given structure and the color guide are shown at the bottom of each panel. tRNA backbone and rRNA base movements are shown by black arrows. Hydrogen bonds are shown using black dotted lines.
Supplementary Figure 4 Schematic diagrams of the hydrogen-bond networks and reaction schemes.
Schematic diagrams of the hydrogen-bond networks and reaction schemes are shown for the proton wire (a) and the eight-membered (b) and six-membered (c) proton shuttles. Hydrogen bond distances and the ω angles between three successive non-hydrogen atoms are indicated. The reaction scheme shown in panel (a) is only for the first half of the peptidyl transfer reaction. Nucleophilic attack is shown by red arrows.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 4354 kb)
Proton-wire model for peptide-bond formation.
High-resolution structures of the 70S ribosome from Thermus thermophiles reveal a network of ordered waters in the peptidyl transferase center that suggests a mechanism for proton movement and formation and breakdown of the tetrahedral intermediate. (MP4 28018 kb)
Rights and permissions
About this article
Cite this article
Polikanov, Y., Steitz, T. & Innis, C. A proton wire to couple aminoacyl-tRNA accommodation and peptide-bond formation on the ribosome. Nat Struct Mol Biol 21, 787–793 (2014). https://doi.org/10.1038/nsmb.2871
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2871
This article is cited by
-
The SecM arrest peptide traps a pre-peptide bond formation state of the ribosome
Nature Communications (2024)
-
RAPP-containing arrest peptides induce translational stalling by short circuiting the ribosomal peptidyltransferase activity
Nature Communications (2024)
-
Establishing the fundamental rules for genetic code expansion
Nature Chemistry (2023)
-
A molecular network of conserved factors keeps ribosomes dormant in the egg
Nature (2023)
-
Atomistic simulations of the Escherichia coli ribosome provide selection criteria for translationally active substrates
Nature Chemistry (2023)