Abstract
Although telomeres are heterochromatic, they are transcribed into noncoding telomeric repeat–containing RNA (TERRA). Here we show that RNA-DNA hybrids form at telomeres and are removed by RNase H enzymes in the budding yeast, Saccharomyces cerevisiae. In recombination-competent telomerase mutants, telomeric RNA-DNA hybrids promote recombination-mediated elongation events that delay the onset of cellular senescence. Reduction of TERRA and telomeric RNA-DNA–hybrid levels diminishes rates of recombination-mediated telomere elongation in cis. Overexpression of RNase H decreases telomere recombination rates and accelerates senescence in recombination-competent but not recombination-deficient cells. In contrast, in the absence of both telomerase and homologous recombination, accumulation of telomeric RNA-DNA hybrids leads to telomere loss and accelerated rates of cellular senescence. Therefore, the regulation of TERRA transcription and telomeric RNA-DNA–hybrid formation are important determinants of both telomere-length dynamics and proliferative potential after the inactivation of telomerase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
de Lange, T. How telomeres solve the end-protection problem. Science 326, 948–952 (2009).
Watson, J.D. Origin of concatemeric T7 DNA. Nat. New Biol. 239, 197–201 (1972).
Lingner, J., Cooper, J.P. & Cech, T.R. Telomerase and DNA end replication: no longer a lagging strand problem? Science 269, 1533–1534 (1995).
Greider, C.W. & Blackburn, E.H. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43, 405–413 (1985).
Shay, J.W. & Wright, W.E. Senescence and immortalization: role of telomeres and telomerase. Carcinogenesis 26, 867–874 (2005).
Wellinger, R.J. & Zakian, V.A. Everything you ever wanted to know about Saccharomyces cerevisiae telomeres: beginning to end. Genetics 191, 1073–1105 (2012).
Lundblad, V. & Szostak, J.W. A mutant with a defect in telomere elongation leads to senescence in yeast. Cell 57, 633–643 (1989).
Lundblad, V. & Blackburn, E.H. An alternative pathway for yeast telomere maintenance rescues est1- senescence. Cell 73, 347–360 (1993).
Luke, B. & Lingner, J. TERRA: telomeric repeat-containing RNA. EMBO J. 28, 2503–2510 (2009).
Schoeftner, S. & Blasco, M.A. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat. Cell Biol. 10, 228–236 (2008).
Luke, B. et al. The Rat1p 5′ to 3′ exonuclease degrades telomeric repeat-containing RNA and promotes telomere elongation in Saccharomyces cerevisiae. Mol. Cell 32, 465–477 (2008).
Porro, A., Feuerhahn, S., Reichenbach, P. & Lingner, J. Molecular dissection of telomeric repeat-containing RNA biogenesis unveils the presence of distinct and multiple regulatory pathways. Mol. Cell Biol. 30, 4808–4817 (2010).
Yehezkel, S., Segev, Y., Viegas-Pequignot, E., Skorecki, K. & Selig, S. Hypomethylation of subtelomeric regions in ICF syndrome is associated with abnormally short telomeres and enhanced transcription from telomeric regions. Hum. Mol. Genet. 17, 2776–2789 (2008).
Azzalin, C.M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Maicher, A., Kastner, L., Dees, M. & Luke, B. Deregulated telomere transcription causes replication-dependent telomere shortening and promotes cellular senescence. Nucleic Acids Res. 40, 6649–6659 (2012).
Pfeiffer, V. & Lingner, J. TERRA promotes telomere shortening through exonuclease 1-mediated resection of chromosome ends. PLoS Genet. 8, e1002747 (2012).
Chawla, R. et al. Human UPF1 interacts with TPP1 and telomerase and sustains telomere leading-strand replication. EMBO J. 30, 4047–4058 (2011).
Iglesias, N. et al. Subtelomeric repetitive elements determine TERRA regulation by Rap1/Rif and Rap1/Sir complexes in yeast. EMBO Rep. 12, 587–593 (2011).
Chawla, R. & Azzalin, C.M. The telomeric transcriptome and SMG proteins at the crossroads. Cytogenet. Genome Res. 122, 194–201 (2008).
Gómez -Gonzalez, B. et al. Genome-wide function of THO/TREX in active genes prevents R-loop-dependent replication obstacles. EMBO J. 30, 3106–3119 (2011).
Arudchandran, A. et al. The absence of ribonuclease H1 or H2 alters the sensitivity of Saccharomyces cerevisiae to hydroxyurea, caffeine and ethyl methanesulphonate: implications for roles of RNases H in DNA replication and repair. Genes Cells 5, 789–802 (2000).
Jeong, H.S., Backlund, P.S., Chen, H.C., Karavanov, A.A. & Crouch, R.J. RNase H2 of Saccharomyces cerevisiae is a complex of three proteins. Nucleic Acids Res. 32, 407–414 (2004).
Luna, R., Rondon, A.G. & Aguilera, A. New clues to understand the role of THO and other functionally related factors in mRNP biogenesis. Biochim. Biophys. Acta 1819, 514–520 (2012).
Gaillard, H. & Aguilera, A. Transcription coupled repair at the interface between transcription elongation and mRNP biogenesis. Biochim. Biophys. Acta 1829, 141–150 (2013).
Gaillard, H., Herrera-Moyano, E. & Aguilera, A. Transcription-associated genome instability. Chem. Rev., http://dx.doi.org/10.1021/cr400017y (18 April 2013).
Prado, F. & Aguilera, A. Impairment of replication fork progression mediates RNA polII transcription-associated recombination. EMBO J. 24, 1267–1276 (2005).
Wellinger, R.E., Prado, F. & Aguilera, A. Replication fork progression is impaired by transcription in hyperrecombinant yeast cells lacking a functional THO complex. Mol. Cell Biol. 26, 3327–3334 (2006).
Boguslawski, S.J. et al. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 89, 123–130 (1986).
Hu, Z., Zhang, A., Storz, G., Gottesman, S. & Leppla, S.H. An antibody-based microarray assay for small RNA detection. Nucleic Acids Res. 34, e52 (2006).
El Hage, A., French, S.L., Beyer, A.L. & Tollervey, D. Loss of Topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010).
Poveda, A.M., Le Clech, M. & Pasero, P. Transcription and replication: breaking the rules of the road causes genomic instability. Transcription 1, 99–102 (2010).
Chang, M., Dittmar, J.C. & Rothstein, R. Long telomeres are preferentially extended during recombination-mediated telomere maintenance. Nat. Struct. Mol. Biol. 18, 451–456 (2011).
Teixeira, M.T., Arneric, M., Sperisen, P. & Lingner, J. Telomere length homeostasis is achieved via a switch between telomerase-extendible and -nonextendible states. Cell 117, 323–335 (2004).
Sandell, L.L., Gottschling, D.E. & Zakian, V.A. Transcription of a yeast telomere alleviates telomere position effect without affecting chromosome stability. Proc. Natl. Acad. Sci. USA 91, 12061–12065 (1994).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003).
Griffith, J.D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Fouché, N. et al. The basic domain of TRF2 directs binding to DNA junctions irrespective of the presence of TTAGGG repeats. J. Biol. Chem. 281, 37486–37495 (2006).
Atkinson, J. & McGlynn, P. Replication fork reversal and the maintenance of genome stability. Nucleic Acids Res. 37, 3475–3492 (2009).
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012).
Gottipati, P., Cassel, T.N., Savolainen, L. & Helleday, T. Transcription-associated recombination is dependent on replication in mammalian cells. Mol. Cell Biol. 28, 154–164 (2008).
Gottipati, P. & Helleday, T. Transcription-associated recombination in eukaryotes: link between transcription, replication and recombination. Mutagenesis 24, 203–210 (2009).
Lowell, J.E., Roughton, A.I., Lundblad, V. & Pillus, L. Telomerase-independent proliferation is influenced by cell type in Saccharomyces cerevisiae. Genetics 164, 909–921 (2003).
Arnoult, N., Van Beneden, A. & Decottignies, A. Telomere length regulates TERRA levels through increased trimethylation of telomeric H3K9 and HP1α. Nat. Struct. Mol. Biol. 19, 948–956 (2012).
Azzalin, C.M. & Lingner, J. Telomeres: the silence is broken. Cell Cycle 7, 1161–1165 (2008).
Nergadze, S.G. et al. CpG-island promoters drive transcription of human telomeres. RNA 15, 2186–2194 (2009).
Ng, L.J., Cropley, J.E., Pickett, H.A., Reddel, R.R. & Suter, C.M. Telomerase activity is associated with an increase in DNA methylation at the proximal subtelomere and a reduction in telomeric transcription. Nucleic Acids Res. 37, 1152–1159 (2009).
Guthrie, C. & Fink, G.R. Guide to Yeast Genetics and Molecular Biology Vol 194 (Academic Press, 1991).
Acknowledgements
We acknowledge M. Chang for help with alignments and T. Schmidt and M.G. Montana for strain generation. B.B. was supported by a Landesgraduiertenförderung fellowship. S.L.-G. is supported by a long-term European Molecular Biology Organization–Marie Curie cofund fellowship (ALTF 9-2010). This work was supported by the Deutsche Forschungsgemeinschaft (SFB 1036) to B.L. and the Netzwerk Alters-Forschung (Ministerium für Wissenschaft, Forschung und Kunst Baden-Württemberg) to B.L.
Author information
Authors and Affiliations
Contributions
B.B., A.M. and B.L. designed experiments. B.B., A.M., M.D., J.K., S.L.-G. and K.B. carried out experimental work and performed the data analysis. B.B., A.M., J.K., S.L.-G. and B.L. wrote the manuscript. B.L. guided the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Telomeric RNA-DNA hybrids are regulated by RNase H enzymes
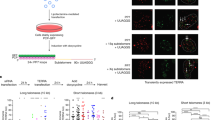
RNA-DNA hybrids exist at telomeres and accumulate in RNase H mutants. The identical ChIP as described in Fig. 1a, however quantified showing % input on the y axis. Values are represented as % input of telomeric DNA recovered and are the means of seven (+ab) and five (-ab) biological replicates, error bars depict ± s.e.m.P values were derived from two-tailed Student's t-tests (NS = not significant). (b) The RNA-DNA hybrid signal is RNase H sensitive. Following overnight immunoprecipitation with the S9.6 antibodies, one wild type sample was treated with recombinant RNase H before the washing steps (see online methods). (c) TERRA levels are not increased in rnh1 rnh201 mutants. qRT-PCR was performed for the 6Y', 1L and 15L telomeres using the indicated mutants (at approx. PD 15 following tetrad dissection). sir2 cells served as positive control where TERRA is upregulated. Average values were derived from 3 biological replicates where error bars depict ± s.e.m.
Supplementary Figure 2 Accumulation of RNA-DNA hybrids promotes homologous recombination
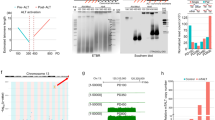
(a) Raw data from Fig. 1b. The individual pairs within a graph were derived from the same tetrad for direct comparisons. (b) The second and third biological replicates from Fig. 2a. (c) Premature survivor formation does not account for the increased rate of HR in rnh1 rnh201 cells. Genomic DNA derived from tetrad 2 of the curve shown in Fig. 1b was digested with Xho1. TRFs of Y' telomeres can be seen between 1164 and 992 bps. Recombination rates for Fig. 2a were determined at around PD 35, when type II survivors had clearly not yet formed.
Supplementary Figure 3 Rnh1 and Rnh201 act redundantly to remove telomeric RNA-DNA hybrids
(a) The increased rate of senescence shown in Fig. 2c only occurs in absence of telomerase. Senescence curves were performed as previously described (Fig. 1b). The loss of RNase H activity does not result in decreased cell viability in rad52 cells, but accelerates senescence only when both EST2 and RAD52 are co-deleted. Curve averages were derived from 6 biological replicates for each genotype depicted, error bars depict ± s.e.m. est2 rad52 and est2 rad52 rnh1 rnh201 curves are taken form Fig. 2c. (b, c) Senescence curves in an est2 rad52 background were performed with only one of the RNase H genes deleted (rnh1 or rnh201, respectively). Rates of senescence were not affected upon deletion of one RNase H gene only. Curves represent the means of 6 biological replicates ± s.e.m. (d) Raw data from Fig. 2c. Genotypes are indicated.
Supplementary Figure 4 The THO mutant hpr1 shows increased telomeric RNA-DNA hybrids and increased rates of telomeric recombination
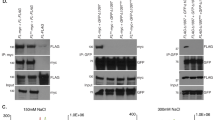
(a) hpr1 shows increased telomeric RNA-DNA hybrids. ChIP was carried out as described in Fig. 1a. The values are represented as % input of telomeric DNA recovered relative to wild type (set to 1) and are the means of 5 biological replicates, error bars depict ± s.e.m. P values were derived from a two-tailed one-sample Student's t-test. (b) Deletion of HPR1 accelerates senescence in HR deficient telomerase mutants. In the presence of RAD52 the viability loss was alleviated, suggesting HR-mediated compensation. Liquid senescence assays were performed on the indicated strains. Curves represent the average value for 6 biological replicates ± s.e.m. (c) hpr1 shows increased rates of telomeric recombination. 1L Telomere-PCR was performed on genomic DNA extracted from the two indicated clones, which were derived form the same tetrad. 1L Telomere-PCR products were cloned and sequenced as described in Fig. 2a and % of diverged telomeres was calculated (est2 n = 48, PD 30; est2 hpr1 n = 39; PD 26).
Supplementary Figure 5 Overexpression of RNH1 increases senescence in recombination-competent cells
(a) Raw data for Fig. 5a for the indicated genotypes. (b) Raw data for Fig. 5e for the indicated genotypes
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–3 (PDF 1978 kb)
Rights and permissions
About this article
Cite this article
Balk, B., Maicher, A., Dees, M. et al. Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat Struct Mol Biol 20, 1199–1205 (2013). https://doi.org/10.1038/nsmb.2662
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2662
This article is cited by
-
XPF activates break-induced telomere synthesis
Nature Communications (2022)
-
Telomerase subunit Est2 marks internal sites that are prone to accumulate DNA damage
BMC Biology (2021)
-
TERRA G-quadruplex RNA interaction with TRF2 GAR domain is required for telomere integrity
Scientific Reports (2021)
-
TERRA transcription destabilizes telomere integrity to initiate break-induced replication in human ALT cells
Nature Communications (2021)
-
R-loop proximity proteomics identifies a role of DDX41 in transcription-associated genomic instability
Nature Communications (2021)