Abstract
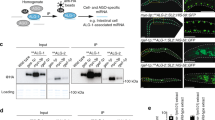
Homeostatic mechanisms regulate the abundance of several components in small-RNA pathways. We used Drosophila and mammalian systems to demonstrate a conserved homeostatic system in which the status of miRNA biogenesis controls Argonaute protein stability. Clonal analyses of multiple mutants of core Drosophila miRNA factors revealed that stability of the miRNA effector AGO1 is dependent on miRNA biogenesis. Reciprocally, ectopic transcription of miRNAs within in vivo clones induced accumulation of AGO1, as did genetic interference with the ubiquitin-proteasome system. In mouse cells, we found that the stability of Ago2 declined in Dicer-knockout cells and was rescued by proteasome blockade or introduction of either Dicer plasmid or Dicer-independent miRNA constructs. Notably, Dicer-dependent miRNA constructs generated pre-miRNAs that bound Ago2 but did not rescue Ago2 stability. We conclude that Argonaute levels are finely tuned by cellular availability of mature miRNAs and the ubiquitin-proteasome system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Czech, B. & Hannon, G.J. Small RNA sorting: matchmaking for Argonautes. Nat. Rev. Genet. 12, 19–31 (2011).
Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Yang, J.S. & Lai, E.C. Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol. Cell 43, 892–903 (2011).
Fabian, M.R. & Sonenberg, N. The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC. Nat. Struct. Mol. Biol. 19, 586–593 (2012).
Heo, I. & Kim, V.N. Regulating the regulators: posttranslational modifications of RNA silencing factors. Cell 139, 28–31 (2009).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Kadener, S. et al. Genome-wide identification of targets of the drosha-pasha/DGCR8 complex. RNA 15, 537–545 (2009).
Smibert, P. et al. A Drosophila genetic screen yields allelic series of core microRNA biogenesis factors and reveals post-developmental roles for microRNAs. RNA 17, 1997–2010 (2011).
Iki, T., Yoshikawa, M., Meshi, T. & Ishikawa, M. Cyclophilin 40 facilitates HSP90-mediated RISC assembly in plants. EMBO J. 31, 267–278 (2012).
Iki, T. et al. In vitro assembly of plant RNA-induced silencing complexes facilitated by molecular chaperone HSP90. Mol. Cell 39, 282–291 (2010).
Miyoshi, T., Takeuchi, A., Siomi, H. & Siomi, M.C. A direct role for Hsp90 in pre-RISC formation in Drosophila. Nat. Struct. Mol. Biol. 17, 1024–1026 (2010).
Iwasaki, S. et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 39, 292–299 (2010).
Johnston, M., Geoffroy, M.C., Sobala, A., Hay, R. & Hutvagner, G. HSP90 protein stabilizes unloaded argonaute complexes and microscopic P-bodies in human cells. Mol. Biol. Cell 21, 1462–1469 (2010).
Ishizu, H., Siomi, H. & Siomi, M.C. Biology of PIWI-interacting RNAs: new insights into biogenesis and function inside and outside of germlines. Genes Dev. 26, 2361–2373 (2012).
Handler, D. et al. A systematic analysis of Drosophila TUDOR domain-containing proteins identifies Vreteno and the Tdrd12 family as essential primary piRNA pathway factors. EMBO J. 30, 3977–3993 (2011).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Jiang, F. et al. Dicer-1 and R3D1-L catalyze microRNA maturation in Drosophila. Genes Dev. 19, 1674–1679 (2005).
Förstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Miyoshi, K., Okada, T.N., Siomi, H. & Siomi, M.C. Characterization of the miRNA-RISC loading complex and miRNA-RISC formed in the Drosophila miRNA pathway. RNA 15, 1282–1291 (2009).
Azzam, G., Smibert, P., Lai, E.C. & Liu, J.L. Drosophila Argonaute 1 and its miRNA biogenesis partners are required for oocyte formation and germline cell division. Dev. Biol. 365, 384–394 (2012).
Herranz, H. et al. The miRNA machinery targets Mei-P26 and regulates Myc protein levels in the Drosophila wing. EMBO J. 29, 1688–1698 (2010).
Yao, B., La, L.B., Chen, Y.C., Chang, L.J. & Chan, E.K. Defining a new role of GW182 in maintaining miRNA stability. EMBO Rep. 13, 1102–1108 (2012).
Neumüller, R.A. et al. Mei-P26 regulates microRNAs and cell growth in the Drosophila ovarian stem cell lineage. Nature 454, 241–245 (2008).
Okamura, K. et al. The Drosophila hairpin RNA pathway generates endogenous short interfering RNAs. Nature 453, 803–806 (2008).
Czech, B. et al. Hierarchical rules for Argonaute loading in Drosophila. Mol. Cell 36, 445–456 (2009).
Czech, B. et al. An endogenous siRNA pathway in Drosophila. Nature 453, 798–802 (2008).
Okamura, K., Robine, N., Liu, Y., Liu, Q. & Lai, E.C. R2D2 organizes small regulatory RNA pathways in Drosophila. Mol. Cell Biol. 31, 884–896 (2011).
Lee, Y.S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Yang, J.S. et al. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 107, 15163–15168 (2010).
Castilla-Llorente, V. et al. Mammalian GW220/TNGW1 is essential for the formation of GW/P bodies containing miRISC. J. Cell Biol. 198, 529–544 (2012).
Gibbings, D. et al. Selective autophagy degrades DICER and AGO2 and regulates miRNA activity. Nat. Cell Biol. 14, 1314–1321 (2012).
Yang, J.S. & Lai, E.C. Dicer-independent, Ago2-mediated microRNA biogenesis in vertebrates. Cell Cycle 9, 4455–4460 (2010).
Maurin, T., Cazalla, D., Yang, J.S., Bortolamiol-Becet, D. & Lai, E.C. RNase III-independent microRNA biogenesis in mammalian cells. RNA 18, 2166–2173 (2012).
Okamura, K., Liu, N. & Lai, E.C. Distinct mechanisms for microRNA strand selection by Drosophila Argonautes. Mol. Cell 36, 431–444 (2009).
Okamura, K. et al. The regulatory activity of microRNA* species has substantial influence on microRNA and 3′ UTR evolution. Nat. Struct. Mol. Biol. 15, 354–363 (2008).
Elkayam, E. et al. The structure of human Argonaute-2 in complex with miR-20a. Cell 150, 100–110 (2012).
Schirle, N.T. & MacRae, I.J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Martinez, N.J. & Gregory, R.I. Argonaute2 expression is post-transcriptionally coupled to microRNA abundance. RNA 19, 605–612 (2013).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Diederichs, S. & Haber, D.A. Dual role for Argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 131, 1097–1108 (2007).
Yang, J.S., Maurin, T. & Lai, E.C. Functional parameters of Dicer-independent microRNA biogenesis. RNA 18, 945–957 (2012).
Janas, M.M. et al. Alternative RISC assembly: binding and repression of microRNA-mRNA duplexes by human Ago proteins. RNA 18, 2041–2055 (2012).
Wang, D. et al. Quantitative functions of Argonaute proteins in mammalian development. Genes Dev. 26, 693–704 (2012).
Ameres, S.L., Hung, J.H., Xu, J., Weng, Z. & Zamore, P.D. Target RNA-directed tailing and trimming purifies the sorting of endo-siRNAs between the two Drosophila Argonaute proteins. RNA 17, 54–63 (2011).
Zhao, S. et al. piRNA-triggered MIWI ubiquitination and removal by APC/C in late spermatogenesis. Dev. Cell 24, 13–25 (2013).
Bronevetsky, Y. et al. T cell activation induces proteasomal degradation of Argonaute and rapid remodeling of the microRNA repertoire. J. Exp. Med. 210, 417–432 (2013).
Tan, G.S. et al. Expanded RNA-binding activities of mammalian Argonaute 2. Nucleic Acids Res. 37, 7533–7545 (2009).
Kataoka, Y., Takeichi, M. & Uemura, T. Developmental roles and molecular characterization of a Drosophila homologue of Arabidopsis Argonaute1, the founder of a novel gene superfamily. Genes Cells 6, 313–325 (2001).
Yang, L. et al. Argonaute 1 regulates the fate of germline stem cells in Drosophila. Development 134, 4265–4272 (2007).
Martin, R. et al. A Drosophila pasha mutant distinguishes the canonical miRNA and mirtron pathways. Mol. Cell Biol. 29, 861–870 (2009).
Schweisguth, F. Dominant-negative mutation in the b2 and b6 proteasome-subunit genes affect alternative cell fate decisions in the Drosophila sense organ lineage. Proc. Natl. Acad. Sci. USA 96, 11382–11386 (1999).
Xu, X.L., Li, Y., Wang, F. & Gao, F.B. The steady-state level of the nervous-system-specific microRNA-124a is regulated by dFMR1 in Drosophila. J. Neurosci. 28, 11883–11889 (2008).
Bejarano, F. et al. A genome-wide transgenic resource for conditional expression of Drosophila microRNAs. Development 139, 2821–2831 (2012).
Lai, E.C. & Rubin, G.M. neuralized functions cell-autonomously to regulate a subset of Notch-dependent processes during adult Drosophila development. Dev. Biol. 231, 217–233 (2001).
Liu, N., Han, H. & Lasko, P. Vasa promotes Drosophila germline stem cell differentiation by activating mei-P26 translation by directly interacting with a (U)-rich motif in its 3′ UTR. Genes Dev. 23, 2742–2752 (2009).
Okamura, K., Hagen, J.W., Duan, H., Tyler, D.M. & Lai, E.C. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell 130, 89–100 (2007).
Acknowledgements
We thank G. Meister (Universität Regensburg, Regensburg, Germany), M. Overholtzer (Sloan-Kettering Institute, New York, New York, USA), J. Liu (Sloan-Kettering Institute, New York, New York, USA), X. Jiang (Sloan-Kettering Institute, New York, New York, USA), K. Okamura (Temasek Institute, Singapore), V. Kim (Seoul National University, Seoul, Korea), H. Siomi (Keio University, Tokyo, Japan), A. Tarakhovsky (Rockefeller University, New York, New York, USA), D. O'Carroll (European Molecular Biology Laboratory, Monterotondo Scalo, Italy), R. Carthew (Northwestern University, Chicago, Illinois, USA), F. Schweisguth (Institut Pasteur, Paris, France), F. Gao (University of Massachusetts Medical School, Worcester, Massachusetts, USA), Paul Lasko (McGill University, Montreal, Canada), the Vienna Drosophila RNAi Center (Vienna, Austria), the Transgenic RNAi Project at Harvard Medical School (Boston, Massachusetts, USA) and the Bloomington Drosophila Stock Center (Bloomington, Indiana, USA) for reagents and discussions. Work in J.-L.L.'s group was supported by the UK Medical Research Council. G.A. was supported by the Malaysian Ministry of Higher Education. Work in E.C.L.'s group was supported by the US National Institutes of Health R01-GM083300.
Author information
Authors and Affiliations
Contributions
P.S., G.A. and J.-L.L. made the initial observations that demonstrated AGO1 homeostasis in vivo. Mechanistic experiments were carried out in flies by P.S., in S2 cells by P.S. and J.-S.Y. and in mammalian cells by J.-S.Y. P.S., J.-S.Y. and E.C.L. wrote the text with input from G.A. and J.-L.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 3131 kb)
Rights and permissions
About this article
Cite this article
Smibert, P., Yang, JS., Azzam, G. et al. Homeostatic control of Argonaute stability by microRNA availability. Nat Struct Mol Biol 20, 789–795 (2013). https://doi.org/10.1038/nsmb.2606
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2606
This article is cited by
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)
-
Argonaute 2 is lost from neuromuscular junctions affected with amyotrophic lateral sclerosis in SOD1G93A mice
Scientific Reports (2022)
-
Cell-type-specific profiling of loaded miRNAs from Caenorhabditis elegans reveals spatial and temporal flexibility in Argonaute loading
Nature Communications (2021)
-
Impaired AGO2/miR-185-3p/NRP1 axis promotes colorectal cancer metastasis
Cell Death & Disease (2021)
-
XPO5 promotes primary miRNA processing independently of RanGTP
Nature Communications (2020)