Abstract
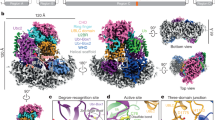
Homologous to E6-AP C terminus (HECT) E3 ligases recognize and directly catalyze ligation of ubiquitin (Ub) to their substrates. Molecular details of this process remain unknown. We report the first structure, to our knowledge, of a Ub-loaded E3, the human neural precursor cell–expressed developmentally downregulated protein 4 (Nedd4). The HECTNedd4~Ub transitory intermediate provides a structural basis for the proposed sequential addition mechanism. The donor Ub, transferred from the E2, is bound to the Nedd4 C lobe with its C-terminal tail locked in an extended conformation, primed for catalysis. We provide evidence that the Nedd4-family members are Lys63-specific enzymes whose catalysis is mediated by an essential C-terminal acidic residue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Scheffner, M., Nuber, U. & Huibregtse, J.M. Protein ubiquitination involving an E1-E2-E3 enzyme ubiquitin thioester cascade. Nature 373, 81–83 (1995).
Scheffner, M. & Staub, O. HECT E3s and human disease. BMC Biochem. 8 (suppl. 1), S6 (2007).
Marín, I. Animal HECT ubiquitin ligases: evolution and functional implications. BMC Evol. Biol. 10, 56 (2010).
Rotin, D. & Kumar, S. Physiological functions of the HECT family of ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 10, 398–409 (2009).
Verdecia, M.A. et al. Conformational flexibility underlies ubiquitin ligation mediated by the WWP1 HECT domain E3 ligase. Mol. Cell 11, 249–259 (2003).
Kamadurai, H.B. et al. Insights into ubiquitin transfer cascades from a structure of a UbcH5B~ubiquitin-HECT(NEDD4L) complex. Mol. Cell 36, 1095–1102 (2009).
Kim, H.C., Steffen, A.M., Oldham, M.L., Chen, J. & Huibregtse, J.M. Structure and function of a HECT domain ubiquitin-binding site. EMBO Rep. 12, 334–341 (2011).
Maspero, E. et al. Structure of the HECT:ubiquitin complex and its role in ubiquitin chain elongation. EMBO Rep. 12, 342–349 (2011).
Kim, H.C. & Huibregtse, J.M. Polyubiquitination by HECT E3s and the determinants of chain type specificity. Mol. Cell Biol. 29, 3307–3318 (2009).
Wang, M. & Pickart, C.M. Different HECT domain ubiquitin ligases employ distinct mechanisms of polyubiquitin chain synthesis. EMBO J. 24, 4324–4333 (2005).
Remaut, H. & Waksman, G. Protein-protein interaction through β-strand addition. Trends Biochem. Sci. 31, 436–444 (2006).
Wang, M., Cheng, D., Peng, J. & Pickart, C.M. Molecular determinants of polyubiquitin linkage selection by an HECT ubiquitin ligase. EMBO J. 25, 1710–1719 (2006).
Dou, H., Buetow, L., Sibbet, G.J., Cameron, K. & Huang, D.T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Plechanovová, A., Jaffray, E.G., Tatham, M.H., Naismith, J.H. & Hay, R.T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Reverter, D. & Lima, C.D. Insights into E3 ligase activity revealed by a SUMO–RanGAP1–Ubc9–Nup358 complex. Nature 435, 687–692 (2005).
Salvat, C., Wang, G., Dastur, A., Lyon, N. & Huibregtse, J.M. The -4 phenylalanine is required for substrate ubiquitination catalyzed by HECT ubiquitin ligases. J. Biol. Chem. 279, 18935–18943 (2004).
Ronchi, V.P., Klein, J.M. & Haas, A.L. E6AP/UBE3A ubiquitin ligase harbors two e2~ubiquitin binding sites. J. Biol. Chem. published online, doi:10.1074/jbc.M113.458059 (25 February 2013).
Hochstrasser, M. Lingering mysteries of ubiquitin-chain assembly. Cell 124, 27–34 (2006).
Woelk, T. et al. Molecular mechanisms of coupled monoubiquitination. Nat. Cell Biol. 8, 1246–1254 (2006).
Serniwka, S.A., & Shaw, G.S. The structure of the UbcH8-ubiquitin complex shows a unique ubiquitin interaction site. Biochemistry 48, 12169–12179 (2009).
Dimasi, N., Flot, D., Dupeux, F. & Marquez, J.A. Expression, crystallization and X-ray data collection from microcrystals of the extracellular domain of the human inhibitory receptor expressed on myeloid cells IREM-1. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 63, 204–208 (2007).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–189 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Acknowledgements
We thank P. Romano for critically reading the manuscript and for helpful discussions and advice and the staff at the European Synchrotron Radiation Facility and Swiss Light Source for assistance in data collection. This work was supported by grants from the Associazione Italiana per la Ricerca sul Cancro (AIRC, IG11627), the European Community (Network of Excellence FP6, 100601–201012) and the European Molecular Biology Organization Young Investigator Program to S. Polo. Access to the High Throughput Crystallization Laboratory of the European Molecular Biology Laboratory Grenoble outstation was funded by the European Community's Seventh Framework Programme (FP7/2007-2013) PCUBE (grant agreement 227764).
Author information
Authors and Affiliations
Contributions
E.M. conducted the experiments with the help of E.V. and S.M.; V.C. assisted in crystallization; P.S. carried out MS analysis; S. Pasqualato performed structure determination and description; S. Pasqualato and E.M. participated in experimental design and data analysis; S. Polo conceived of the project, interpreted the results and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 7879 kb)
Rights and permissions
About this article
Cite this article
Maspero, E., Valentini, E., Mari, S. et al. Structure of a ubiquitin-loaded HECT ligase reveals the molecular basis for catalytic priming. Nat Struct Mol Biol 20, 696–701 (2013). https://doi.org/10.1038/nsmb.2566
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2566