Abstract
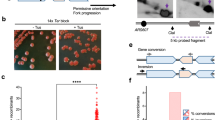
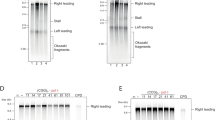
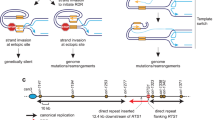
Expansion of GAA/TTC repeats is the causative event in Friedreich's ataxia. GAA repeats have been shown to hinder replication in model systems, but the mechanisms of replication interference and expansion in human cells remained elusive. To study in vivo replication structures at GAA repeats, we designed a new plasmid-based system that permits the analysis of human replication intermediates by two-dimensional gel electrophoresis and EM. We found that replication forks transiently pause and reverse at long GAA/TTC tracts in both orientations. Furthermore, we identified replication-associated intramolecular junctions, located between GAA/TTC repeats and other homopurine-homopyrimidine tracts, that were associated with breakage of the plasmid fork not traversing the repeats. Finally, we detected postreplicative, sister-chromatid hemicatenanes on control plasmids, which were replaced by persistent homology-driven junctions at GAA/TTC repeats. These data prove that GAA/TTC tracts interfere with replication in humans and implicate postreplicative mechanisms in trinucleotide repeat expansion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
McMurray, C.T. Mechanisms of trinucleotide repeat instability during human development. Nat. Rev. Genet. 11, 786–799 (2010).
Mirkin, S.M. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007).
Schmucker, S. & Puccio, H. Understanding the molecular mechanisms of Friedreich's ataxia to develop therapeutic approaches. Hum. Mol. Genet. 19, R103–R110 (2010).
Gacy, A.M. et al. GAA instability in Friedreich's Ataxia shares a common, DNA-directed and intraallelic mechanism with other trinucleotide diseases. Mol. Cell 1, 583–593 (1998).
Sakamoto, N. et al. Sticky DNA: self-association properties of long GAA.TTC repeats in R.R.Y triplex structures from Friedreich's ataxia. Mol. Cell 3, 465–475 (1999).
Vetcher, A.A., Napierala, M. & Wells, R.D. Sticky DNA: effect of the polypurine.polypyrimidine sequence. J. Biol. Chem. 277, 39228–39234 (2002).
Ditch, S., Sammarco, M.C., Banerjee, A. & Grabczyk, E. Progressive GAA.TTC repeat expansion in human cell lines. PLoS Genet. 5, e1000704 (2009).
Rindler, P.M. & Bidichandani, S.I. Role of transcript and interplay between transcription and replication in triplet-repeat instability in mammalian cells. Nucleic Acids Res. 39, 526–535 (2011).
Mirkin, S.M. & Smirnova, E.V. Positioned to expand. Nat. Genet. 31, 5–6 (2002).
Samadashwily, G.M., Raca, G. & Mirkin, S.M. Trinucleotide repeats affect DNA replication in vivo. Nat. Genet. 17, 298–304 (1997).
Voineagu, I., Surka, C.F., Shishkin, A.A., Krasilnikova, M.M. & Mirkin, S.M. Replisome stalling and stabilization at CGG repeats, which are responsible for chromosomal fragility. Nat. Struct. Mol. Biol. 16, 226–228 (2009).
Freudenreich, C.H., Kantrow, S.M. & Zakian, V.A. Expansion and length-dependent fragility of CTG repeats in yeast. Science 279, 853–856 (1998).
Krasilnikova, M.M. & Mirkin, S.M. Replication stalling at Friedreich's ataxia (GAA)n repeats in vivo. Mol. Cell Biol. 24, 2286–2295 (2004).
Miret, J.J., Pessoa-Brandao, L. & Lahue, R.S. Orientation-dependent and sequence-specific expansions of CTG/CAG trinucleotide repeats in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 95, 12438–12443 (1998).
Fouché, N., Ozgur, S., Roy, D. & Griffith, J.D. Replication fork regression in repetitive DNAs. Nucleic Acids Res. 34, 6044–6050 (2006).
Mirkin, S.M. DNA structures, repeat expansions and human hereditary disorders. Curr. Opin. Struct. Biol. 16, 351–358 (2006).
Shishkin, A.A. et al. Large-scale expansions of Friedreich's ataxia GAA repeats in yeast. Mol. Cell 35, 82–92 (2009).
Daee, D.L., Mertz, T. & Lahue, R.S. Postreplication repair inhibits CAG.CTG repeat expansions in Saccharomyces cerevisiae. Mol. Cell Biol. 27, 102–110 (2007).
Kerrest, A. et al. SRS2 and SGS1 prevent chromosomal breaks and stabilize triplet repeats by restraining recombination. Nat. Struct. Mol. Biol. 16, 159–167 (2009).
Chandok, G.S., Patel, M.P., Mirkin, S.M. & Krasilnikova, M.M. Effects of Friedreich's ataxia GAA repeats on DNA replication in mammalian cells. Nucleic Acids Res. 40, 3964–3974 (2012).
Cleary, J.D., Nichol, K., Wang, Y.H. & Pearson, C.E. Evidence of cis-acting factors in replication-mediated trinucleotide repeat instability in primate cells. Nat. Genet. 31, 37–46 (2002).
M. Rindler, P., Clark, R.M., Pollard, L.M., De Biase, I. & Bidichandani, S.I. Replication in mammalian cells recapitulates the locus-specific differences in somatic instability of genomic GAA triplet-repeats. Nucleic Acids Res. 34, 6352–6361 (2006).
Sogo, J.M., Stahl, H., Koller, T. & Knippers, R. Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J. Mol. Biol. 189, 189–204 (1986).
Follonier, C. & Lopes, M. Combined bi-dimensional electrophoresis and electron microscopy to study specific plasmid DNA replication intermediates in human cells. Methods Mol. Biol. (in the press).
Bénard, M., Maric, C. & Pierron, G. DNA replication-dependent formation of joint DNA molecules in Physarum polycephalum. Mol. Cell 7, 971–980 (2001).
Lopes, M., Cotta-Ramusino, C., Liberi, G. & Foiani, M. Branch migrating sister chromatid junctions form at replication origins through Rad51/Rad52-independent mechanisms. Mol. Cell 12, 1499–1510 (2003).
Lucas, I. & Hyrien, O. Hemicatenanes form upon inhibition of DNA replication. Nucleic Acids Res. 28, 2187–2193 (2000).
Segurado, M., Gomez, M. & Antequera, F. Increased recombination intermediates and homologous integration hot spots at DNA replication origins. Mol. Cell 10, 907–916 (2002).
Zou, H. & Rothstein, R. Holliday junctions accumulate in replication mutants via a RecA homolog-independent mechanism. Cell 90, 87–96 (1997).
Liberi, G. et al. Methods to study replication fork collapse in budding yeast. Methods Enzymol. 409, 442–462 (2006).
Neelsen, K.J., Ray Chaudhuri, A., Follonier, C., Herrador, R. & Lopes, M. Visualization and interpretation of eukaryotic DNA replication intermediates by electron microscopy in vivo. Methods Mol. Biol. (in the press).
Kalejta, R.F. & Hamlin, J.L. Composite patterns in neutral/neutral two-dimensional gels demonstrate inefficient replication origin usage. Mol. Cell Biol. 16, 4915–4922 (1996).
Duckett, D.R. et al. The structure of the Holliday junction, and its resolution. Cell 55, 79–89 (1988).
Malkov, V.A., Voloshin, O.N., Soyfer, V.N. & Frank-Kamenetskii, M.D. Cation and sequence effects on stability of intermolecular pyrimidine-purine-purine triplex. Nucleic Acids Res. 21, 585–591 (1993).
Son, L.S., Bacolla, A. & Wells, R.D. Sticky DNA: in vivo formation in E. coli and in vitro association of long GAA*TTC tracts to generate two independent supercoiled domains. J. Mol. Biol. 360, 267–284 (2006).
Lopes, M. Electron microscopy methods for studying in vivo DNA replication intermediates. Methods Mol. Biol. 521, 605–631 (2009).
Beal, P.A. & Dervan, P.B. The influence of single base triplet changes on the stability of a pur.pur.pyr triple helix determined by affinity cleaving. Nucleic Acids Res. 20, 2773–2776 (1992).
Potaman, V.N. et al. Length-dependent structure formation in Friedreich ataxia (GAA)n*(TTC)n repeats at neutral pH. Nucleic Acids Res. 32, 1224–1231 (2004).
Pandolfo, M. Friedreich ataxia: the clinical picture. J. Neurol. 256 (suppl. 1), 3–8 (2009).
Frank-Kamenetskii, M.D. & Mirkin, S.M. Triplex DNA structures. Annu. Rev. Biochem. 64, 65–95 (1995).
Kovtun, I.V. et al. OGG1 initiates age-dependent CAG trinucleotide expansion in somatic cells. Nature 447, 447–452 (2007).
Kim, H.M. et al. Chromosome fragility at GAA tracts in yeast depends on repeat orientation and requires mismatch repair. EMBO J. 27, 2896–2906 (2008).
Tang, W. et al. Friedreich's ataxia (GAA)n*(TTC)n repeats strongly stimulate mitotic crossovers in Saccharomyces cerevisae. PLoS Genet. 7, e1001270 (2011).
Raghavan, S.C. & Lieber, M.R. DNA structures at chromosomal translocation sites. Bioessays 28, 480–494 (2006).
Glazkov, M.V. Loop organization of eukaryotic chromosomes and triple-stranded DNA structures. Mol. Biol. (Mosk.) 45, 294–306 (2011).
Vetcher, A.A. et al. Sticky DNA, a long GAA.GAA.TTC triplex that is formed intramolecularly, in the sequence of intron 1 of the frataxin gene. J. Biol. Chem. 277, 39217–39227 (2002).
Sinha, N.K., Morris, C.F. & Alberts, B.M. Efficient in vitro replication of double-stranded DNA templates by a purified T4 bacteriophage replication system. J. Biol. Chem. 255, 4290–4293 (1980).
Ohshima, K., Montermini, L., Wells, R.D. & Pandolfo, M. Inhibitory effects of expanded GAA.TTC triplet repeats from intron I of the Friedreich ataxia gene on transcription and replication in vivo. J. Biol. Chem. 273, 14588–14595 (1998).
Ziegler, K., Bui, T., Frisque, R.J., Grandinetti, A. & Nerurkar, V.R. A rapid in vitro polyomavirus DNA replication assay. J. Virol. Methods 122, 123–127 (2004).
Brewer, B.J. & Fangman, W.L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell 51, 463–471 (1987).
Acknowledgements
We thank M. Pandolfo (Campus ERASME, Université Libre de Bruxelles, Brussels, Belgium) for providing plasmids and L. Pelkmans (Institute of Molecular Life Science, University of Zürich, Zürich, Switzerland) for the SV40 strain. We thank also the Center for Microscopy and Image Analysis of the University of Zürich for technical assistance with EM. We thank J. Jiricny for critical reading of the manuscript. We are grateful to S. Mirkin, V. Zakian and all members of the Lopes lab for helpful discussions and valuable input on this project. This work was supported by the Swiss National Science Foundation grants PP0033-114922 and PP00P3-135292 to M.L.
Author information
Authors and Affiliations
Contributions
C.F. established and optimized the plasmid-based system used in this study and performed the 2D-gel and EM experiments. J.O. performed the 2D gel in Figure 1b. R.H. contributed to the RAD51-depletion experiment and performed the western blot. M.L. planned and supervised the project and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 2034 kb)
Rights and permissions
About this article
Cite this article
Follonier, C., Oehler, J., Herrador, R. et al. Friedreich's ataxia–associated GAA repeats induce replication-fork reversal and unusual molecular junctions. Nat Struct Mol Biol 20, 486–494 (2013). https://doi.org/10.1038/nsmb.2520
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2520
This article is cited by
-
The mechanism of replication stalling and recovery within repetitive DNA
Nature Communications (2022)
-
Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites
Nature Communications (2021)
-
Sequential role of RAD51 paralog complexes in replication fork remodeling and restart
Nature Communications (2020)
-
Mechanisms of genetic instability caused by (CGG)n repeats in an experimental mammalian system
Nature Structural & Molecular Biology (2018)
-
RNA biology of disease-associated microsatellite repeat expansions
Acta Neuropathologica Communications (2017)