Abstract
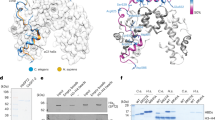
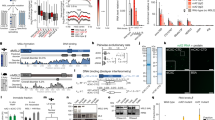
X-chromosome dosage compensation by the MSL (male-specific lethal) complex is required in Drosophila melanogaster to increase gene expression from the single male X to equal that of both female X chromosomes. Instead of focusing solely on protein complexes released from DNA, here we used chromatin-interacting protein MS (ChIP-MS) to identify MSL interactions on cross-linked chromatin. We identified MSL-enriched histone modifications, including histone H4 Lys16 acetylation and histone H3 Lys36 methylation, and CG4747, a putative Lys36-trimethylated histone H3 (H3K36me3)-binding protein. CG4747 is associated with the bodies of active genes, coincident with H3K36me3, and is mislocalized in the Set2 mutant lacking H3K36me3. CG4747 loss of function in vivo results in partial mislocalization of the MSL complex to autosomes, and RNA interference experiments confirm that CG4747 and Set2 function together to facilitate targeting of the MSL complex to active genes, validating the ChIP-MS approach.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Lucchesi, J.C., Kelly, W.G. & Panning, B. Chromatin remodeling in dosage compensation. Annu. Rev. Genet. 39, 615–651 (2005).
Conrad, T. & Akhtar, A. Dosage compensation in Drosophila melanogaster: epigenetic fine-tuning of chromosome-wide transcription. Nat. Rev. Genet. 13, 123–134 (2011).
Alekseyenko, A.A., Larschan, E., Lai, W.R., Park, P.J. & Kuroda, M.I. High-resolution ChIP-chip analysis reveals that the Drosophila MSL complex selectively identifies active genes on the male X chromosome. Genes Dev. 20, 848–857 (2006).
Gilfillan, G.D. et al. Chromosome-wide gene-specific targeting of the Drosophila dosage compensation complex. Genes Dev. 20, 858–870 (2006).
Gelbart, M.E. & Kuroda, M.I. Drosophila dosage compensation: a complex voyage to the X chromosome. Development 136, 1399–1410 (2009).
Hilfiker, A., Hilfiker-Kleiner, D., Pannuti, A. & Lucchesi, J.C. mof, a putative acetyl transferase gene related to the Tip60 and MOZ human genes and to the SAS genes of yeast, is required for dosage compensation in Drosophila. EMBO J. 16, 2054–2060 (1997).
Jin, Y., Wang, Y., Johansen, J. & Johansen, K.M. JIL-1, a chromosomal kinase implicated in regulation of chromatin structure, associates with the male specific lethal (MSL) dosage compensation complex. J. Cell Biol. 149, 1005–1010 (2000).
Mendjan, S. et al. Nuclear pore components are involved in the transcriptional regulation of dosage compensation in Drosophila. Mol. Cell 21, 811–823 (2006).
Morales, V. et al. Functional integration of the histone acetyltransferase MOF into the dosage compensation complex. EMBO J. 23, 2258–2268 (2004).
Smith, E.R. et al. The Drosophila MSL complex acetylates histone H4 at lysine 16, a chromatin modification linked to dosage compensation. Mol. Cell. Biol. 20, 312–318 (2000).
Lambert, J.-P., Mitchell, L., Rudner, A., Baetz, K. & Figeys, D. A novel proteomics approach for the discovery of chromatin-associated protein networks. Mol. Cell. Proteomics 8, 870–882 (2009).
Lambert, J.-P. et al. Defining the budding yeast chromatin-associated interactome. Mol. Syst. Biol. 6, 448 (2010).
Smart, S.K., Mackintosh, S.G., Edmondson, R.D., Taverna, S.D. & Tackett, A.J. Mapping the local protein interactome of the NuA3 histone acetyltransferase. Protein Sci. 18, 1987–1997 (2009).
Guerrero, C., Tagwerker, C., Kaiser, P. & Huang, L. An integrated mass spectrometry-based proteomic approach: quantitative analysis of tandem affinity-purified in vivo cross-linked protein complexes (QTAX) to decipher the 26 S proteasome-interacting network. Mol. Cell. Proteomics 5, 366–378 (2006).
Tagwerker, C. et al. A tandem affinity tag for two-step purification under fully denaturing conditions: application in ubiquitin profiling and protein complex identification combined with in vivocross-linking. Mol. Cell. Proteomics 5, 737–748 (2006).
Tardiff, D.F., Abruzzi, K.C. & Rosbash, M. Protein characterization of Saccharomyces cerevisiae RNA polymerase II after in vivo cross-linking. Proc. Natl. Acad. Sci. USA 104, 19948–19953 (2007).
Déjardin, J. & Kingston, R.E. Purification of proteins associated with specific genomic loci. Cell 136, 175–186 (2009).
Larschan, E. et al. Identification of chromatin-associated regulators of MSL complex targeting in Drosophila dosage compensation. PLoS Genet. 8, e1002830 (2012).
Cronan, J.E. Biotination of proteins in vivo. A post-translational modification to label, purify, and study proteins. J. Biol. Chem. 265, 10327–10333 (1990).
Copps, K. et al. Complex formation by the Drosophila MSL proteins: role of the MSL2 RING finger in protein complex assembly. EMBO J. 17, 5409–5417 (1998).
Hamada, F.N., Park, P.J., Gordadze, P.R. & Kuroda, M.I. Global regulation of X chromosomal genes by the MSL complex in Drosophila melanogaster. Genes Dev. 19, 2289–2294 (2005).
Straub, T., Gilfillan, G.D., Maier, V.K. & Becker, P.B. The Drosophila MSL complex activates the transcription of target genes. Genes Dev. 19, 2284–2288 (2005).
Kharchenko, P.V. et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 471, 480–485 (2011).
Kho, Y. et al. A tagging-via-substrate technology for detection and proteomics of farnesylated proteins. Proc. Natl. Acad. Sci. USA 101, 12479–12484 (2004).
Turner, B.M., Birley, A.J. & Lavender, J. Histone H4 isoforms acetylated at specific lysine residues define individual chromosomes and chromatin domains in Drosophila polytene nuclei. Cell 69, 375–384 (1992).
Larschan, E. et al. MSL complex is attracted to genes marked by H3K36 trimethylation using a sequence-independent mechanism. Mol. Cell 28, 121–133 (2007).
Sural, T.H. et al. The MSL3 chromodomain directs a key targeting step for dosage compensation of the Drosophila melanogaster X chromosome. Nat. Struct. Mol. Biol. 15, 1318–1325 (2008).
Bell, O. et al. Transcription-coupled methylation of histone H3 at lysine 36 regulates dosage compensation by enhancing recruitment of the MSL complex in Drosophila melanogaster. Mol. Cell Biol. 28, 3401–3409 (2008).
Plazas-Mayorca, M.D. et al. One-pot shotgun quantitative mass spectrometry characterization of histones. J. Proteome Res. 8, 5367–5374 (2009).
Akhtar, A. & Becker, P.B. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol. Cell 5, 367–375 (2000).
Smith, E.R., Allis, C.D. & Lucchesi, J.C. Linking global histone acetylation to the transcription enhancement of X-chromosomal genes in Drosophila males. J. Biol. Chem. 276, 31483–31486 (2001).
Kind, J. et al. Genome-wide analysis reveals MOF as a key regulator of dosage compensation and gene expression in Drosophila. Cell 133, 813–828 (2008).
Gelbart, M.E., Larschan, E., Peng, S., Park, P.J. & Kuroda, M.I. Drosophila MSL complex globally acetylates H4K16 on the male X chromosome for dosage compensation. Nat. Struct. Mol. Biol. 16, 825–832 (2009).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Nguyen, A.T. & Zhang, Y. The diverse functions of Dot1 and H3K79 methylation. Genes Dev. 25, 1345–1358 (2011).
Vermeulen, M. et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142, 967–980 (2010).
Carrozza, M.J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Huh, J.-W. et al. Multivalent di-nucleosome recognition enables the Rpd3S histone deacetylase complex to tolerate decreased H3K36 methylation levels. EMBO J. 31, 3564–3574 (2012).
Ge, H., Si, Y. & Wolffe, A.P. A novel transcriptional coactivator, p52, functionally interacts with the essential splicing factor ASF/SF2. Mol. Cell 2, 751–759 (1998).
Pradeepa, M.M., Sutherland, H.G., Ule, J., Grimes, G.R. & Bickmore, W.A. Psip1/Ledgf p52 binds methylated histone H3K36 and splicing factors and contributes to the regulation of alternative splicing. PLoS Genet. 8, e1002717 (2012).
Maurer-Stroh, S. et al. The Tudor domain 'Royal Family': Tudor, plant Agenet, Chromo, PWWP and MBT domains. Trends Biochem. Sci. 28, 69–74 (2003).
Vezzoli, A. et al. Molecular basis of histone H3K36me3 recognition by the PWWP domain of Brpf1. Nat. Struct. Mol. Biol. 17, 617–619 (2010).
Dhayalan, A. et al. The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J. Biol. Chem. 285, 26114–26120 (2010).
Eissenberg, J.C. & Shilatifard, A. Histone H3 lysine 4 (H3K4) methylation in development and differentiation. Dev. Biol. 339, 240–249 (2010).
Mohan, M. et al. The COMPASS family of H3K4 methylases in Drosophila. Mol. Cell. Biol. 31, 4310–4318 (2011).
Cheng, H., He, X. & Moore, C. The essential WD repeat protein Swd2 has dual functions in RNA polymerase II transcription termination and lysine 4 methylation of histone H3. Mol. Cell. Biol. 24, 2932–2943 (2004).
Regnard, C. et al. Global analysis of the relationship between JIL-1 kinase and transcription. PLoS Genet. 7, e1001327 (2011).
Qiu, C., Sawada, K., Zhang, X. & Cheng, X. The PWWP domain of mammalian DNA methyltransferase Dnmt3b defines a new family of DNA-binding folds. Nat. Struct. Biol. 9, 217–224 (2002).
Handler, D. et al. A systematic analysis of Drosophila TUDOR domain-containing proteins identifies Vreteno and the Tdrd12 family as essential primary piRNA pathway factors. EMBO J. 30, 3977–3993 (2011).
Ni, J.-Q. et al. A genome-scale shRNA resource for transgenic RNAi in Drosophila. Nat. Methods 8, 405–407 (2011).
Mito, Y., Henikoff, J.G. & Henikoff, S. Genome-scale profiling of histone H3.3 replacement patterns. Nat. Genet. 37, 1090–1097 (2005).
Moore, S.A., Ferhatoglu, Y., Jia, Y., Al-Jiab, R.A. & Scott, M.J. Structural and biochemical studies on the chromo-barrel domain of male specific lethal 3 (MSL3) reveal a binding preference for mono or dimethyl lysine 20 on histone H4. J. Biol. Chem. 285, 40879–40890 (2010).
Kim, D. et al. Corecognition of DNA and a methylated histone tail by the MSL3 chromodomain. Nat. Struct. Mol. Biol. 17, 1027–1029 (2010).
Alekseyenko, A.A. et al. A sequence motif within chromatin entry sites directs MSL establishment on the Drosophila X chromosome. Cell 134, 599–609 (2008).
Schotta, G. & Reuter, G. Controlled expression of tagged proteins in Drosophila using a new modular P-element vector system. Mol. Gen. Genet. 262, 916–920 (2000).
Puig, O. et al. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24, 218–229 (2001).
Kelley, R.L. et al. Epigenetic spreading of the Drosophila dosage compensation complex from roX RNA genes into flanking chromatin. Cell 98, 513–522 (1999).
Saumweber, H., Symmons, P., Kabisch, R., Will, H. & Bonhoeffer, F. Monoclonal antibodies against chromosomal proteins of Drosophila melanogaster: establishment of antibody producing cell lines and partial characterization of corresponding antigens. Chromosoma 80, 253–275 (1980).
Acknowledgements
We thank P. Kaiser (University of California, Irvine, Irvine, California, USA) for the HTB tag and insightful discussions. We thank G. Schotta (Ludwig-Maximilians Universität, Munich, Germany) for the pGS-mw[+] vector. The anti-Z4 antibody was a generous gift from H. Saumweber (Humboldt University, Berlin, Germany). We thank E. Gerace from the Moazed lab (Harvard Medical School, Boston, Massachusetts, USA) for providing the protocol for coupling IgG to magnetic beads. We thank the Bloomington Stock Center and TRiP at Harvard Medical School (NIH/NIGMS R01-GM084947) for providing fly stocks used in this study. We are grateful to M. Gelbart for support and expertise, E. Smith for technical assistance and A. Ciccia, B. Adamson and A. Plachetka for critical reading of the manuscript. This work was supported by grants from the US National Institutes of Health (NIH) to M.I.K. (GM45744 and GM101958), P.V.K. (K25AG037596) and S.J.E. (GM44664). A.E.H.E. is supported by fellowships from The Jane Coffin Childs Foundation and The American Society for Radiation Oncology. B.A.G. is supported by a US National Science Foundation Early Faculty CAREER award and NIH award number DP2OD007447 from the Office of the Director.
Author information
Authors and Affiliations
Contributions
C.I.W., A.A.A. and A.A.G. performed ChIP-MS experiments; A.E.H.E. performed LC-MS/MS to identify MSL-interacting proteins; G.L. and L.-M.P.B. performed quantitative MS of histone PTM; C.I.W. performed all other experiments; P.V.K. performed all bioinformatics analyses; S.J.E., B.A.G. and M.I.K. supervised analyses; C.I.W. and M.I.K. prepared the manuscript in consultation with all coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Tables 1–3 and Supplementary Note (PDF 347 kb)
Rights and permissions
About this article
Cite this article
Wang, C., Alekseyenko, A., LeRoy, G. et al. Chromatin proteins captured by ChIP–mass spectrometry are linked to dosage compensation in Drosophila. Nat Struct Mol Biol 20, 202–209 (2013). https://doi.org/10.1038/nsmb.2477
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2477
This article is cited by
-
A bioinformatics screen reveals hox and chromatin remodeling factors at the Drosophila histone locus
BMC Genomic Data (2023)
-
VULCAN integrates ChIP-seq with patient-derived co-expression networks to identify GRHL2 as a key co-regulator of ERa at enhancers in breast cancer
Genome Biology (2019)
-
JASPer controls interphase histone H3S10 phosphorylation by chromosomal kinase JIL-1 in Drosophila
Nature Communications (2019)
-
Versatile and efficient chromatin pull-down methodology based on DNA triple helix formation
Scientific Reports (2018)
-
ADP-ribose-specific chromatin-affinity purification for investigating genome-wide or locus-specific chromatin ADP-ribosylation
Nature Protocols (2017)