Abstract
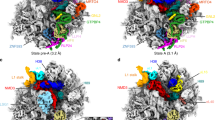
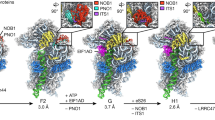
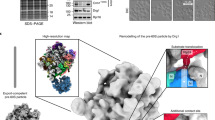
Preribosomal particles evolve in the nucleus through transient interaction with biogenesis factors before export to the cytoplasm. Here, we report the architecture of the late pre-60S particle, purified from Saccharomyces cerevisiae, through Arx1, a nuclear export factor with structural homology to methionine aminopeptidases, or its binding partner Alb1. Cryo-EM reconstruction of the Arx1 particle at 11.9-Å resolution reveals regions of extra density on the pre-60S particle attributed to associated biogenesis factors, confirming the immature state of the nascent subunit. One of these densities could be unambiguously assigned to Arx1. Immunoelectron microscopy and UV cross-linking localize Arx1 close to the ribosomal exit tunnel, in direct contact with ES27, a highly dynamic eukaryotic rRNA expansion segment. The binding of Arx1 at the exit tunnel may position this export factor to prevent premature recruitment of ribosome-associated factors active during translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kressler, D., Hurt, E. & Bassler, J. Driving ribosome assembly. Biochim. Biophys. Acta 1803, 673–683 (2010).
Karbstein, K. Inside the 40S ribosome assembly machinery. Curr. Opin. Chem. Biol. 15, 657–663 (2011).
Görlich, D. & Kutay, U. Transport between the cell nucleus and the cytoplasm. Annu. Rev. Cell Dev. Biol. 15, 607–660 (1999).
Zemp, I. & Kutay, U. Nuclear export and cytoplasmic maturation of ribosomal subunits. FEBS Lett. 581, 2783–2793 (2007).
Spahn, C.M. et al. Structure of the 80S ribosome from Saccharomyces cerevisiae–tRNA-ribosome and subunit-subunit interactions. Cell 107, 373–386 (2001).
Armache, J.P. et al. Cryo-EM structure and rRNA model of a translating eukaryotic 80S ribosome at 5.5-A resolution. Proc. Natl. Acad. Sci. USA 107, 19748–19753 (2010).
Ben-Shem, A., Jenner, L., Yusupova, G. & Yusupov, M. Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209 (2010).
Rabl, J., Leibundgut, M., Ataide, S.F., Haag, A. & Ban, N. Crystal structure of the eukaryotic 40S ribosomal subunit in complex with initiation factor 1. Science 331, 730–736 (2011).
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011).
Tschochner, H. & Hurt, E. Pre-ribosomes on the road from the nucleolus to the cytoplasm. Trends Cell Biol. 13, 255–263 (2003).
Schäfer, T. et al. Hrr25-dependent phosphorylation state regulates organization of the pre-40S subunit. Nature 441, 651–655 (2006).
Strunk, B.S. et al. Ribosome assembly factors prevent premature translation initiation by 40S assembly intermediates. Science 333, 1449–1453 (2011).
Nissan, T.A. et al. A pre-ribosome with a tadpole-like structure functions in ATP-dependent maturation of 60S subunits. Mol. Cell 15, 295–301 (2004).
Ulbrich, C. et al. Mechanochemical removal of ribosome biogenesis factors from nascent 60S ribosomal subunits. Cell 138, 911–922 (2009).
Ho, J.H., Kallstrom, G. & Johnson, A.W. Nmd3p is a Crm1p-dependent adapter protein for nuclear export of the large ribosomal subunit. J. Cell Biol. 151, 1057–1066 (2000).
Gadal, O. et al. Nuclear export of 60s ribosomal subunits depends on Xpo1p and requires a nuclear export sequence-containing factor, Nmd3p, that associates with the large subunit protein Rpl10p. Mol. Cell. Biol. 21, 3405–3415 (2001).
Sengupta, J. et al. Characterization of the nuclear export adaptor protein Nmd3 in association with the 60S ribosomal subunit. J. Cell Biol. 189, 1079–1086 (2010).
Yao, W. et al. Nuclear export of ribosomal 60S subunits by the general mRNA export receptor Mex67-Mtr2. Mol. Cell 26, 51–62 (2007).
Yao, W., Lutzmann, M. & Hurt, E. A versatile interaction platform on the Mex67-Mtr2 receptor creates an overlap between mRNA and ribosome export. EMBO J. 27, 6–16 (2008).
Bradatsch, B. et al. Arx1 functions as an unorthodox nuclear export receptor for the 60S preribosomal subunit. Mol. Cell 27, 767–779 (2007).
Hung, N.J., Lo, K.Y., Patel, S.S., Helmke, K. & Johnson, A.W. Arx1 is a nuclear export receptor for the 60S ribosomal subunit in yeast. Mol. Biol. Cell 19, 735–744 (2008).
Yao, Y. et al. Ecm1 is a new pre-ribosomal factor involved in pre-60S particle export. RNA 16, 1007–1017 (2010).
Oeffinger, M., Dlakic, M. & Tollervey, D. A pre-ribosome-associated HEAT-repeat protein is required for export of both ribosomal subunits. Genes Dev. 18, 196–209 (2004).
Hackmann, A., Gross, T., Baierlein, C. & Krebber, H. The mRNA export factor Npl3 mediates the nuclear export of large ribosomal subunits. EMBO Rep. 12, 1024–1031 (2011).
Lebreton, A. et al. A functional network involved in the recycling of nucleocytoplasmic pre-60S factors. J. Cell Biol. 173, 349–360 (2006).
Nissan, T.A., Bassler, J., Petfalski, E., Tollervey, D. & Hurt, E. 60S pre-ribosome formation viewed from assembly in the nucleolus until export to the cytoplasm. EMBO J. 21, 5539–5547 (2002).
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 A resolution. Science 334, 1524–1529 (2011).
Kowalinski, E. et al. The crystal structure of Ebp1 reveals a methionine aminopeptidase fold as binding platform for multiple interactions. FEBS Lett. 581, 4450–4454 (2007).
Monie, T.P. et al. Structural insights into the transcriptional and translational roles of Ebp1. EMBO J. 26, 3936–3944 (2007).
Gartmann, M. et al. Mechanism of eIF6-mediated inhibition of ribosomal subunit joining. J. Biol. Chem. 285, 14848–14851 (2010).
Groft, C.M., Beckmann, R., Sali, A. & Burley, S.K. Crystal structures of ribosome anti-association factor IF6. Nat. Struct. Biol. 7, 1156–1164 (2000).
Lebreton, A., Saveanu, C., Decourty, L., Jacquier, A. & Fromont-Racine, M. Nsa2 is an unstable, conserved factor required for the maturation of 27 SB pre-rRNAs. J. Biol. Chem. 281, 27099–27108 (2006).
Granneman, S., Petfalski, E. & Tollervey, D. A cluster of ribosome synthesis factors regulate pre-rRNA folding and 5.8S rRNA maturation by the Rat1 exonuclease. EMBO J. 30, 4006–4019 (2011).
Granneman, S., Petfalski, E., Swiatkowska, A. & Tollervey, D. Cracking pre-40S ribosomal subunit structure by systematic analyses of RNA-protein cross-linking. EMBO J. 29, 2026–2036 (2010).
Granneman, S., Kudla, G., Petfalski, E. & Tollervey, D. Identification of protein binding sites on U3 snoRNA and pre-rRNA by UV cross-linking and high-throughput analysis of cDNAs. Proc. Natl. Acad. Sci. USA 106, 9613–9618 (2009).
Beckmann, R. et al. Architecture of the protein-conducting channel associated with the translating 80S ribosome. Cell 107, 361–372 (2001).
Sweeney, R., Chen, L. & Yao, M.C. An rRNA variable region has an evolutionarily conserved essential role despite sequence divergence. Mol. Cell. Biol. 14, 4203–4215 (1994).
Wilson, D.N. & Nierhaus, K.H. Ribosomal proteins in the spotlight. Crit. Rev. Biochem. Mol. Biol. 40, 243–267 (2005).
Kemmler, S., Occhipinti, L., Veisu, M. & Panse, V.G. Yvh1 is required for a late maturation step in the 60S biogenesis pathway. J. Cell Biol. 186, 863–880 (2009).
Lo, K.Y., Li, Z., Wang, F., Marcotte, E.M. & Johnson, A.W. Ribosome stalk assembly requires the dual-specificity phosphatase Yvh1 for the exchange of Mrt4 with P0. J. Cell Biol. 186, 849–862 (2009).
Kruiswijk, T., Planta, R.J. & Krop, J.M. The course of the assembly of ribosomal subunits in yeast. Biochim. Biophys. Acta 517, 378–389 (1978).
Bassler, J., Kallas, M. & Hurt, E. The NUG1 GTPase reveals an N-terminal RNA-binding domain that is essential for association with 60 S pre-ribosomal particles. J. Biol. Chem. 281, 24737–24744 (2006).
Hung, N.J. & Johnson, A.W. Nuclear recycling of the pre-60S ribosomal subunit-associated factor Arx1 depends on Rei1 in Saccharomyces cerevisiae. Mol. Cell. Biol. 26, 3718–3727 (2006).
Hedges, J., West, M. & Johnson, A.W. Release of the export adapter, Nmd3p, from the 60S ribosomal subunit requires Rpl10p and the cytoplasmic GTPase Lsg1p. EMBO J. 24, 567–579 (2005).
Strunk, B.S., Novak, M.N., Young, C.L. & Karbstein, K. A translation-like cycle is a quality control checkpoint for maturing 40S ribosome subunits. Cell 150, 111–121 (2012).
Thomson, E. & Tollervey, D. The final step in 5.8S rRNA processing is cytoplasmic in Saccharomyces cerevisiae. Mol. Cell. Biol. 30, 976–984 (2010).
Becker, T. et al. Structure of monomeric yeast and mammalian Sec61 complexes interacting with the translating ribosome. Science 326, 1369–1373 (2009).
DeLano, W.L. The PyMOL Molecular Graphics System. http://www.pymol.org/ (2002).
Puig, O. et al. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24, 218–229 (2001).
Longtine, M.S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli. J. Microsc. 146, 113–136 (1987).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Wagenknecht, T., Grassucci, R. & Frank, J. Electron microscopy and computer image averaging of ice-embedded large ribosomal subunits from Escherichia coli. J. Mol. Biol. 199, 137–147 (1988).
Penczek, P.A., Frank, J. & Spahn, C.M. A method of focused classification, based on the bootstrap 3D variance analysis, and its application to EF-G-dependent translocation. J. Struct. Biol. 154, 184–194 (2006).
Ho, J.H., Kallstrom, G. & Johnson, A.W. Nascent 60S ribosomal subunits enter the free pool bound by Nmd3p. RNA 6, 1625–1634 (2000).
Santos-Rosa, H. et al. Nuclear mRNA export requires complex formation between Mex67p and Mtr2p at the nuclear pores. Mol. Cell. Biol. 18, 6826–6838 (1998).
Vilardell, J. & Warner, J.R. Ribosomal protein L32 of Saccharomyces cerevisiae influences both the splicing of its own transcript and the processing of rRNA. Mol. Cell. Biol. 17, 1959–1965 (1997).
Acknowledgements
We are very thankful to the European Molecular Biology Laboratory–Heidelberg for providing the electron microscopy facility and computational infrastructure, without which this work would not have been possible, and especially M. Diepholz, C. Blachiere-Batisse, J. Briggs, F. Thommen and M. Wahlers for advice and technical support. The plasmid pFA6a-HTpA-HIS3MX4 was a kind gift of D. Kressler (Unit of Biochemistry, Department of Biology, University of Fribourg, Fribourg, Switzerland). We thank C. Dargemont (Institut Jacques Monod, Universités Paris VI and VII, Centre National de la Recherche Scientifique, Paris, France), A.W. Johnson (Molecular Genetics & Microbiology, University of Texas at Austin, Austin, Texas, USA), A. Lebreton (Unité des Interactions Bactéries-Cellules, Institut Pasteur, Paris, France) and J.R. Warner (Department of Cell Biology, Albert Einstein College of Medicine of Yeshiva University, Bronx, New York, USA) for providing antibodies. This work was supported by grants from the Wellcome Trust (to D.T.) and the Deutsche Forschungsgemeinschaft (DFG Hu363/10-4 to E.H.).
Author information
Authors and Affiliations
Contributions
B. Bradatsch designed and performed the experiments and wrote the manuscript; C.L. performed the cryo-EM analysis, supervised by R.B.; S.G. performed the CRAC experiments and the computational analyses in the laboratory of D.T.; M.G. performed some of the biochemical purifications and growth analysis; B. Böttcher provided know-how and supplied the electron microscopy facility for the negative-stain electron microscopy; E.H. directed the project, designed experiments and wrote the manuscript; all authors contributed to the interpretation of the results and helped write the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 2514 kb)
Rights and permissions
About this article
Cite this article
Bradatsch, B., Leidig, C., Granneman, S. et al. Structure of the pre-60S ribosomal subunit with nuclear export factor Arx1 bound at the exit tunnel. Nat Struct Mol Biol 19, 1234–1241 (2012). https://doi.org/10.1038/nsmb.2438
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2438
This article is cited by
-
Structural basis of Naa20 activity towards a canonical NatB substrate
Communications Biology (2021)
-
In situ cryo-electron tomography reveals gradient organization of ribosome biogenesis in intact nucleoli
Nature Communications (2021)
-
Nuclear export of the pre-60S ribosomal subunit through single nuclear pores observed in real time
Nature Communications (2021)
-
Inhibiting eukaryotic ribosome biogenesis
BMC Biology (2019)
-
Ribosome assembly coming into focus
Nature Reviews Molecular Cell Biology (2019)