Abstract
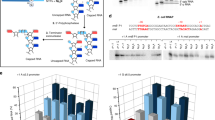
Recent studies showed that Rai1 is a crucial component of the mRNA 5′-end-capping quality-control mechanism in yeast. The yeast genome encodes a weak homolog of Rai1, Ydr370C, but little is known about this protein. Here we report the crystal structures of Ydr370C from Kluyveromyces lactis and the first biochemical and functional studies on this protein. The overall structure of Ydr370C is similar to Rai1. Ydr370C has robust decapping activity on RNAs with unmethylated caps, but it has no detectable pyrophosphohydrolase activity. Unexpectedly, Ydr370C also possesses distributive, 5′-3′ exoRNase activity, and we propose the name Dxo1 for this new eukaryotic enzyme with both decapping and exonuclease activities. Studies of yeast in which both Dxo1 and Rai1 are disrupted reveal that mRNAs with incomplete caps are produced even under normal growth conditions, in sharp contrast to current understanding of the capping process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Shatkin, A.J. & Manley, J.L. The ends of the affair: capping and polyadenylation. Nat. Struct. Biol. 7, 838–842 (2000).
Shuman, S. The mRNA capping apparatus as drug target and guide to eukaryotic phylogeny. Cold Spring Harb. Symp. Quant. Biol. 66, 301–312 (2001).
Hocine, S., Singer, R.H. & Grunwald, D. RNA processing and export. Cold Spring Harb. Perspect. Biol. 2, a000752 (2010).
Ghosh, A. & Lima, C.D. Enzymology of RNA cap synthesis. Wiley Interdiscip. Rev. RNA 1, 152–172 (2010).
Coller, J. & Parker, R. Eukaryotic mRNA decapping. Annu. Rev. Biochem. 73, 861–890 (2004).
Cougot, N., van Dijk, E., Babajko, S. & Seraphin, B. 'Cap-tabolism'. Trends Biochem. Sci. 29, 436–444 (2004).
Franks, T.M. & Lykke-Andersen, S. The control of mRNA decapping and P-body formation. Mol. Cell 32, 605–615 (2008).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Li, Y. & Kiledjian, M. Regulation of mRNA decapping. Wiley Interdiscip. Rev. RNA 1, 253–265 (2010).
Stevens, A. An exoribonuclease from Saccharomyces cerevisiae: effect of modifications of 5′ end groups on the hydrolysis of substrates to 5′ mononucleotides. Biochem. Biophys. Res. Commun. 81, 656–661 (1978).
Stevens, A. Purification and characterization of a Saccharomyces cerevisiae exoribonuclease which yields 5′-mononucleotides by a 5′→3′ mode of hydrolysis. J. Biol. Chem. 255, 3080–3085 (1980).
Chang, J.H., Xiang, S. & Tong, L. 5′-3′ exoribonucleases. in Ribonucleases, Nucleic Acids and Molecular Biology Vol. 26 (ed. Nicholson, A.W.) Ch. 7, 167–192 (Springer, 2011).
Song, M.-G., Li, Y. & Kiledjian, M. Multiple mRNA decapping enzymes in mammalian cells. Mol. Cell 40, 423–432 (2010).
Xiang, S. et al. Structure and function of the 5′→3′ exoribonuclease Rat1 and its activating partner Rai1. Nature 458, 784–788 (2009).
Jiao, X. et al. Identification of a quality-control mechanism for mRNA 5′-end capping. Nature 467, 608–611 (2010).
Xue, Y. et al. Saccharomyces cerevisiae RAI1 (YGL246c) is homologous to human DOM3Z and encodes a protein that binds the nuclear exoribonuclease Rat1p. Mol. Cell. Biol. 20, 4006–4015 (2000).
Huh, W.K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Aravind, L., Makarova, K.S. & Koonin, E.V. Holliday junction resolvases and related nucleases: identification of new families, phyletic distribution and evolutionary trajectories. Nucleic Acids Res. 28, 3417–3432 (2000).
Buisson, M. et al. A bridge crosses the active-site canyon of the Epstein-Barr virus nuclease with DNase and RNase activities. J. Mol. Biol. 391, 717–728 (2009).
Dahlroth, S.-L., Gurmu, D., Haas, J., Erlandsen, H. & Nordlund, P. Crystal structure of the shutoff and exonuclease protein from the oncogenic Kaposi's sarcoma-associated herpesvirus. FEBS J. 276, 6636–6645 (2009).
Holm, L., Kaariainen, S., Rosenstrom, P. & Schenkel, A. Searching protein structure databases with DaliLite v.3. Bioinformatics 24, 2780–2781 (2008).
Zhang, J., Xing, X., Herr, A.B. & Bell, C.E. Crystal structure of E. coli RecE protein reveals a toroidal tetramer for processing double-stranded DNA breaks. Structure 17, 690–702 (2009).
Kovall, R. & Matthews, B.W. Toroidal structure of lambda-exonuclease. Science 277, 1824–1827 (1997).
Kovall, R.A. & Matthews, B.W. Type II restriction endonucleases: structural, functional and evolutionary relationships. Curr. Opin. Chem. Biol. 3, 578–583 (1999).
Joshi, H.K., Etzkorn, C., Chatwell, L., Bitinaite, J. & Horton, N.C. Alteration of sequence specificity of the type II restriction endonuclease HincII through an indirect readout mechanism. J. Biol. Chem. 281, 23852–23869 (2006).
Sinturel, F. et al. Real-time fluorescence detection of exoribonucleases. RNA 15, 2057–2062 (2009).
Chang, J.H., Xiang, S., Xiang, K., Manley, J.L. & Tong, L. Structural and biochemical studies of the 5′→3′ exoribonuclease Xrn1. Nat. Struct. Mol. Biol. 18, 270–276 (2011).
Buratowski, S. Connections between mRNA 3′ end processing and transcription termination. Curr. Opin. Cell Biol. 17, 257–261 (2005).
Conti, E. & Izaurralde, E. Nonsense-mediated mRNA decay: molecular insights and mechanistic variations across species. Curr. Opin. Cell Biol. 17, 316–325 (2005).
Isken, O. & Maquat, L.E. Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function. Genes Dev. 21, 1833–1856 (2007).
van Dijk, E.L. et al. XUTs are a class of Xrn1-sensitive antisense regulatory non-coding RNA in yeast. Nature 475, 114–117 (2011).
Thompson, D.M. & Parker, R. Cytoplasmic decay of intergenic transcripts in Saccharomyces cerevisiae. Mol. Cell. Biol. 27, 92–101 (2007).
Doublié, S. et al. Crystallization and preliminary X-ray analysis of the 9 kDa protein of the mouse signal recognition particle and the selenomethionyl-SRP9. FEBS Lett. 384, 219–221 (1996).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Weeks, C.M. & Miller, R. The design and implementation of SnB v2.0. J. Appl. Crystallogr. 32, 120–124 (1999).
Terwilliger, T.C. SOLVE and RESOLVE: automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K.D. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brünger, A.T. et al. Crystallography & NMR System: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Schwer, B., Saha, N., Mao, X., Chen, H.W. & Shuman, S. Structure-function analysis of yeast mRNA cap methyltransferase and high-copy suppression of conditional mutants by AdoMet synthase and the ubiquitin conjugating enzyme Cdc34p. Genetics 155, 1561–1576 (2000).
Köhrer, K. & Domdey, H. Preparation of high molecular weight RNA. Methods Enzymol. 194, 398–405 (1991).
Wang, Z., Day, N., Trifillis, P. & Kiledjian, M. An mRNA stability complex functions with poly(A)-binding protein to stabilize mRNA in vitro. Mol. Cell. Biol. 19, 4552–4560 (1999).
Wang, Z., Jiao, X., Carr-Schmid, A. & Kiledjian, M. The hDcp2 protein is a mammalian mRNA decapping enzyme. Proc. Natl. Acad. Sci. USA 99, 12663–12668 (2002).
Acknowledgements
We thank T.G. Kinzy (University of Medicine and Dentistry of New Jersey, Piscataway , New Jersey, USA) for the TAP-tagged Ydr370C strain; N. Whalen, S. Myers and H. Robinson for setting up the X29A beamline at the National Synchrotron Light Source . This research was supported by grants from the US National Institutes of Health to L.T. (GM090059) and M.K. (GM67005).
Author information
Authors and Affiliations
Contributions
J.H.C. and K.C. performed protein expression, purification and crystallization experiments. J.H.C. carried out crystallographic data collection, structure determination and refinement, as well as mutagenesis and exonuclease assays. X.J. carried out decapping assays and all the studies in yeast cells. C.O. and C.E.M. generated the Rai1 and Dxo1 deletion strains. All authors commented on the manuscript. L.T. and M.K. designed experiments, analyzed data, supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–3 (PDF 2029 kb)
Rights and permissions
About this article
Cite this article
Chang, J., Jiao, X., Chiba, K. et al. Dxo1 is a new type of eukaryotic enzyme with both decapping and 5′-3′ exoribonuclease activity. Nat Struct Mol Biol 19, 1011–1017 (2012). https://doi.org/10.1038/nsmb.2381
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2381
This article is cited by
-
Extensive 5′-surveillance guards against non-canonical NAD-caps of nuclear mRNAs in yeast
Nature Communications (2020)
-
No-Go Decay mRNA cleavage in the ribosome exit tunnel produces 5′-OH ends phosphorylated by Trl1
Nature Communications (2020)
-
Mechanism and structural diversity of exoribonuclease-resistant RNA structures in flaviviral RNAs
Nature Communications (2018)
-
Interconnections between mRNA degradation and RDR-dependent siRNA production in mRNA turnover in plants
Journal of Plant Research (2017)
-
eIF3d is an mRNA cap-binding protein that is required for specialized translation initiation
Nature (2016)