Abstract
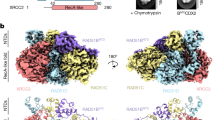
The Mre11–Rad50–Nbs1 (MRN) complex tethers, processes and signals DNA double-strand breaks, promoting genomic stability. To understand the functional architecture of MRN, we determined the crystal structures of the Schizosaccharomyces pombe Mre11 dimeric catalytic domain alone and in complex with a fragment of Nbs1. Two Nbs1 subunits stretch around the outside of the nuclease domains of Mre11, with one subunit additionally bridging and locking the Mre11 dimer via a highly conserved asymmetrical binding motif. Our results show that Mre11 forms a flexible dimer and suggest that Nbs1 not only is a checkpoint adaptor but also functionally influences Mre11-Rad50. Clinical mutations in Mre11 are located along the Nbs1-interaction sites and weaken the Mre11-Nbs1 interaction. However, they differentially affect DNA repair and telomere maintenance in Saccharomyces cerevisiae, potentially providing insight into their different human disease pathologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Mills, K.D., Ferguson, D.O. & Alt, F.W. The role of DNA breaks in genomic instability and tumorigenesis. Immunol. Rev. 194, 77–95 (2003).
Lee, K., Zhang, Y. & Lee, S.E. Saccharomyces cerevisiae ATM orthologue suppresses break-induced chromosome translocations. Nature 454, 543–546 (2008).
Hoeijmakers, J.H. Genome maintenance mechanisms for preventing cancer. Nature 411, 366–374 (2001).
Heyer, W.D., Ehmsen, K.T. & Liu, J. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 44, 113–139 (2010).
Mladenov, E. & Iliakis, G. Induction and repair of DNA double strand breaks: The increasing spectrum of non-homologous end joining pathways. Mutat. Res. 711, 61–72 (2011).
Harper, J.W. & Elledge, S.J. The DNA damage response: ten years after. Mol. Cell 28, 739–745 (2007).
San Filippo, J., Sung, P. & Klein, H. Mechanism of eukaryotic homologous recombination. Annu. Rev. Biochem. 77, 229–257 (2008).
Williams, G.J., Lees-Miller, S.P. & Tainer, J.A. Mre11-Rad50-Nbs1 conformations and the control of sensing, signaling, and effector responses at DNA double-strand breaks. DNA Repair (Amst.) 9, 1299–1306 (2010).
Lieber, M.R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Rahal, E.A. et al. ATM regulates Mre11-dependent DNA end-degradation and microhomology-mediated end joining. Cell Cycle 9, 2866–2877 (2010).
Stracker, T.H. & Petrini, J.H. The MRE11 complex: starting from the ends. Nat. Rev. Mol. Cell Biol. 12, 90–103 (2011).
Cejka, P. et al. DNA end resection by Dna2-Sgs1-RPA and its stimulation by Top3-Rmi1 and Mre11-Rad50-Xrs2. Nature 467, 112–116 (2010).
Faure, V., Coulon, S., Hardy, J. & Geli, V. Cdc13 and telomerase bind through different mechanisms at the lagging- and leading-strand telomeres. Mol. Cell 38, 842–852 (2010).
Rass, E. et al. Role of Mre11 in chromosomal nonhomologous end joining in mammalian cells. Nat. Struct. Mol. Biol. 16, 819–824 (2009).
Borde, V. The multiple roles of the Mre11 complex for meiotic recombination. Chromosome Res. 15, 551–563 (2007).
Hopfner, K.P. et al. The Rad50 zinc-hook is a structure joining Mre11 complexes in DNA recombination and repair. Nature 418, 562–566 (2002).
de Jager, M. et al. Human Rad50/Mre11 is a flexible complex that can tether DNA ends. Mol. Cell 8, 1129–1135 (2001).
Paull, T.T. & Gellert, M. Nbs1 potentiates ATP-driven DNA unwinding and endonuclease cleavage by the Mre11/Rad50 complex. Genes Dev. 13, 1276–1288 (1999).
Nicolette, M.L. et al. Mre11-Rad50-Xrs2 and Sae2 promote 5′ strand resection of DNA double-strand breaks. Nat. Struct. Mol. Biol. 17, 1478–1485 (2010).
Mimitou, E.P. & Symington, L.S. Sae2, Exo1 and Sgs1 collaborate in DNA double-strand break processing. Nature 455, 770–774 (2008).
Sartori, A.A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007).
Zhu, Z., Chung, W.H., Shim, E.Y., Lee, S.E. & Ira, G. Sgs1 helicase and two nucleases Dna2 and Exo1 resect DNA double-strand break ends. Cell 134, 981–994 (2008).
Neale, M.J., Pan, J. & Keeney, S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436, 1053–1057 (2005).
Lloyd, J. et al. A supramodular FHA/BRCT-repeat architecture mediates Nbs1 adaptor function in response to DNA damage. Cell 139, 100–111 (2009).
Williams, R.S. et al. Nbs1 flexibly tethers Ctp1 and Mre11-Rad50 to coordinate DNA double-strand break processing and repair. Cell 139, 87–99 (2009).
Stracker, T.H., Morales, M., Couto, S.S., Hussein, H. & Petrini, J.H. The carboxy terminus of NBS1 is required for induction of apoptosis by the MRE11 complex. Nature 447, 218–221 (2007).
Difilippantonio, S. et al. Distinct domains in Nbs1 regulate irradiation-induced checkpoints and apoptosis. J. Exp. Med. 204, 1003–1011 (2007).
Falck, J., Coates, J. & Jackson, S.P. Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature 434, 605–611 (2005).
Lee, J.H. & Paull, T.T. ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science 308, 551–554 (2005).
Lee, J.H. & Paull, T.T. Direct activation of the ATM protein kinase by the Mre11/Rad50/Nbs1 complex. Science 304, 93–96 (2004).
Costanzo, V., Paull, T., Gottesman, M. & Gautier, J. Mre11 assembles linear DNA fragments into DNA damage signaling complexes. PLoS Biol. 2, E110 (2004).
Dupré, A., Boyer-Chatenet, L. & Gautier, J. Two-step activation of ATM by DNA and the Mre11-Rad50-Nbs1 complex. Nat. Struct. Mol. Biol. 13, 451–457 (2006).
Yazdi, P.T. et al. SMC1 is a downstream effector in the ATM/NBS1 branch of the human S-phase checkpoint. Genes Dev. 16, 571–582 (2002).
Falck, J., Petrini, J.H., Williams, B.R., Lukas, J. & Bartek, J. The DNA damage-dependent intra-S phase checkpoint is regulated by parallel pathways. Nat. Genet. 30, 290–294 (2002).
Derheimer, F.A. & Kastan, M.B. Multiple roles of ATM in monitoring and maintaining DNA integrity. FEBS Lett. 584, 3675–3681 (2010).
Taylor, A.M., Groom, A. & Byrd, P.J. Ataxia-telangiectasia-like disorder (ATLD)-its clinical presentation and molecular basis. DNA Repair (Amst.) 3, 1219–1225 (2004).
Waltes, R. et al. Human RAD50 deficiency in a Nijmegen breakage syndrome-like disorder. Am. J. Hum. Genet. 84, 605–616 (2009).
Carney, J.P. et al. The hMre11/hRad50 protein complex and Nijmegen breakage syndrome: linkage of double-strand break repair to the cellular DNA damage response. Cell 93, 477–486 (1998).
Stewart, G.S. et al. The DNA double-strand break repair gene hMRE11 is mutated in individuals with an ataxia-telangiectasia-like disorder. Cell 99, 577–587 (1999).
Uchisaka, N. et al. Two brothers with ataxia-telangiectasia-like disorder with lung adenocarcinoma. J. Pediatr. 155, 435–438 (2009).
Shull, E.R. et al. Differential DNA damage signaling accounts for distinct neural apoptotic responses in ATLD and NBS. Genes Dev. 23, 171–180 (2009).
Matsumoto, Y. et al. Two unrelated patients with MRE11A mutations and Nijmegen breakage syndrome-like severe microcephaly. DNA Repair (Amst.) 10, 314–321 (2011).
Lammens, K. et al. The Mre11:Rad50 structure shows an ATP-dependent molecular clamp in DNA double-strand break repair. Cell 145, 54–66 (2011).
Ueno, M. et al. Molecular characterization of the Schizosaccharomyces pombe nbs1+ gene involved in DNA repair and telomere maintenance. Mol. Cell. Biol. 23, 6553–6563 (2003).
Fernet, M. et al. Identification and functional consequences of a novel MRE11 mutation affecting 10 Saudi Arabian patients with the ataxia telangiectasia-like disorder. Hum. Mol. Genet. 14, 307–318 (2005).
Chamankhah, M., Fontanie, T. & Xiao, W. The Saccharomyces cerevisiae mre11(ts) allele confers a separation of DNA repair and telomere maintenance functions. Genetics 155, 569–576 (2000).
Bressan, D.A., Olivares, H.A., Nelms, B.E. & Petrini, J.H. Alteration of N-terminal phosphoesterase signature motifs inactivates Saccharomyces cerevisiae Mre11. Genetics 150, 591–600 (1998).
Haber, J.E. Mating-type gene switching in Saccharomyces cerevisiae. Annu. Rev. Genet. 32, 561–599 (1998).
Tsukamoto, Y., Mitsuoka, C., Terasawa, M., Ogawa, H. & Ogawa, T. Xrs2p regulates Mre11p translocation to the nucleus and plays a role in telomere elongation and meiotic recombination. Mol. Biol. Cell 16, 597–608 (2005).
Boulton, S.J. & Jackson, S.P. Components of the Ku-dependent non-homologous end-joining pathway are involved in telomeric length maintenance and telomeric silencing. EMBO J. 17, 1819–1828 (1998).
van der Linden, E., Sanchez, H., Kinoshita, E., Kanaar, R. & Wyman, C. RAD50 and NBS1 form a stable complex functional in DNA binding and tethering. Nucleic Acids Res. 37, 1580–1588 (2009).
Park, Y.B., Chae, J., Kim, Y.C. & Cho, Y. Crystal structure of human Mre11: understanding tumorigenic mutations. Structure 19, 1591–1602 (2011).
Hopfner, K.P. et al. Structural biochemistry and interaction architecture of the DNA double-strand break repair Mre11 nuclease and Rad50-ATPase. Cell 105, 473–485 (2001).
Sabourin, M., Tuzon, C.T. & Zakian, V.A. Telomerase and Tel1p preferentially associate with short telomeres in S. cerevisiae. Mol. Cell 27, 550–561 (2007).
Lim, H.S., Kim, J.S., Park, Y.B., Gwon, G.H. & Cho, Y. Crystal structure of the Mre11-Rad50-ATPγS complex: understanding the interplay between Mre11 and Rad50. Genes Dev. 25, 1091–1104 (2011).
Möckel, C., Lammens, K., Schele, A. & Hopfner, K.P. ATP driven structural changes of the bacterial Mre11:Rad50 catalytic head complex. Nucleic Acids Res. 40, 914–927 (2012).
Williams, G.J. et al. ABC ATPase signature helices in Rad50 link nucleotide state to Mre11 interface for DNA repair. Nat. Struct. Mol. Biol. 18, 423–431 (2011).
Williams, R.S. et al. Mre11 dimers coordinate DNA end bridging and nuclease processing in double-strand-break repair. Cell 135, 97–109 (2008).
Shima, H., Suzuki, M. & Shinohara, M. Isolation and characterization of novel xrs2 mutations in Saccharomyces cerevisiae. Genetics 170, 71–85 (2005).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
D'Amours, D. & Jackson, S.P. The yeast Xrs2 complex functions in S phase checkpoint regulation. Genes Dev. 15, 2238–2249 (2001).
Strahl-Bolsinger, S., Hecht, A., Luo, K. & Grunstein, M. SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes Dev. 11, 83–93 (1997).
Janke, R. et al. A truncated DNA-damage-signaling response is activated after DSB formation in the G1 phase of Saccharomyces cerevisiae. Nucleic Acids Res. 38, 2302–2313 (2010).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Ye, Y. & Godzik, A. FATCAT: a web server for flexible structure comparison and structure similarity searching. Nucleic Acids Res. 32, W582–W585 (2004).
Acknowledgements
We are grateful to J. Petrini (Memorial Sloan-Kettering Cancer Center, New York) for his gift of antibodies to Mre11, Rad50 and Xrs2; members of the Hopfner lab for technical support and discussions; and M. Bennett, A. Rojowska, A. Kopetzki and C. Jung for help with experimentation. We thank the Max Planck Institute crystallization facility for crystallization trials, the staffs of the synchrotron beamlines for help with data collection and processing and SLS and ESRF for beamtime allowance. Research in the K.-P.H. lab was funded by grants from the German Research Council (SFBs 684, 646 and TR5), the German Excellence Initiative, European Commission (IP DNA repair), and US National Institutes of Health (U19AI83025). Research in the K.S. lab was funded by grants from the German Research Council (SFB 646) and the European Research Council (ERC; ERC Starting Grant, project 204522). Research in the S.P.J. lab is supported by grants from Cancer Research UK (C6/A11226), the European Research Council, the European Community's Seventh Framework Program (FP7/2007-2013) under grant agreement HEALTH-F2-2010-259893 and by core infrastructure funding from Cancer Research UK and the Wellcome Trust. S.P.J. receives his salary from the University of Cambridge, supplemented by Cancer Research UK.
Author information
Authors and Affiliations
Contributions
C.B.S., I.G., B.C., H.F. and C.M. designed experiments; C.B.S., F.S. and A.S. cloned constructs and purified proteins; C.B.S. and F.S. crystallized proteins; C.B.S. and K.L. determined crystal structures; I.G., C.B.S., B.C. and H.F. carried out S. cerevisiae assays; C.B.S. did analytical size exclusion experiments; C.M. carried out nuclease activity assays; K.-P.H. and C.B.S. wrote the manuscript; K.L., I.G. and B.C. contributed to the writing and S.P.J., H.F. and C.M. revised the manuscript; K.-P.H., S.P.J. and K.S. supervised the research; K.-P.H. initiated the project and designed the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3 and Supplementary Methods (PDF 6645 kb)
Rights and permissions
About this article
Cite this article
Schiller, C., Lammens, K., Guerini, I. et al. Structure of Mre11–Nbs1 complex yields insights into ataxia-telangiectasia–like disease mutations and DNA damage signaling. Nat Struct Mol Biol 19, 693–700 (2012). https://doi.org/10.1038/nsmb.2323
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2323