Abstract
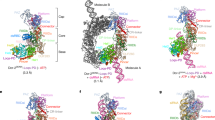
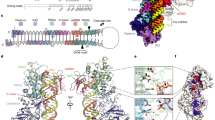
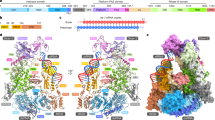
Dicer is a multidomain enzyme that generates small RNAs for gene silencing in eukaryotes. Current understanding of Dicer structure is restricted to simple forms of the enzyme, whereas that of the large and complex Dicer in metazoans is unknown. Here we describe a new domain localization strategy developed to determine the structure of human Dicer by EM. A rearrangement of the nuclease core, compared to the archetypal Giardia lamblia Dicer, explains how metazoan Dicers generate products that are 21–23 nucleotides in length. The helicase domains form a clamp-like structure adjacent to the RNase III active site, facilitating recognition of pre-miRNA loops or translocation on long dsRNAs. Drosophila melanogaster Dicer-2 shows similar features, revealing that the three-dimensional architecture is conserved. These results illuminate the structural basis for small RNA production in eukaryotes and provide a versatile new tool for determining structures of large molecular machines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Carthew, R.W. & Sontheimer, E.J. Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hutvágner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Hammond, S.M., Bernstein, E., Beach, D. & Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–296 (2000).
Pillai, R.S. et al. Inhibition of translational initiation by Let-7 microRNA in human cells. Science 309, 1573–1576 (2005).
Volpe, T.A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Lee, S.R. & Collins, K. Physical and functional coupling of RNA-dependent RNA polymerase and Dicer in the biogenesis of endogenous siRNAs. Nat. Struct. Mol. Biol. 14, 604–610 (2007).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Duchaine, T.F. et al. Functional proteomics reveals the biochemical niche of C. elegans DCR-1 in multiple small-RNA-mediated pathways. Cell 124, 343–354 (2006).
Thivierge, C. et al. Tudor domain ERI-5 tethers an RNA-dependent RNA polymerase to DCR-1 to potentiate endo-RNAi. Nat. Struct. Mol. Biol. 19, 90–97 (2011).
MacRae, I.J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006).
Du, Z., Lee, J.K., Tjhen, R., Stroud, R.M. & James, T.L. Structural and biochemical insights into the dicing mechanism of mouse Dicer: a conserved lysine is critical for dsRNA cleavage. Proc. Natl. Acad. Sci. USA 105, 2391–2396 (2008).
Takeshita, D. et al. Homodimeric structure and double-stranded RNA cleavage activity of the C-terminal RNase III domain of human dicer. J. Mol. Biol. 374, 106–120 (2007).
Qin, H. et al. Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein-protein interaction. RNA 16, 474–481 (2010).
Weinberg, D.E., Nakanishi, K., Patel, D.J. & Bartel, D.P. The inside-out mechanism of dicers from budding yeasts. Cell 146, 262–276 (2011).
Lau, P.W., Potter, C.S., Carragher, B. & MacRae, I.J. Structure of the human Dicer-TRBP complex by electron microscopy. Structure 17, 1326–1332 (2009).
Wang, H.W. et al. Structural insights into RNA processing by the human RISC-loading complex. Nat. Struct. Mol. Biol. 16, 1148–1153 (2009).
Noland, C.L., Ma, E. & Doudna, J.A. siRNA repositioning for guide strand selection by human Dicer complexes. Mol. Cell 43, 110–121 (2011).
Sashital, D.G. & Doudna, J.A. Structural insights into RNA interference. Curr. Opin. Struct. Biol. 20, 90–97 (2010).
Chakravarthy, S., Sternberg, S.H., Kellenberger, C.A. & Doudna, J.A. Substrate-specific kinetics of Dicer-catalyzed RNA processing. J. Mol. Biol. 404, 392–402 (2010).
Hartig, J.V. & Forstemann, K. Loqs-PD and R2D2 define independent pathways for RISC generation in Drosophila. Nucleic Acids Res. 39, 3836–3851 (2011).
Snead, N.M. & Rossi, J.J. Biogenesis and function of endogenous and exogenous siRNAs. Wiley Interdiscip Rev RNA 1, 117–131 (2010).
Cenik, E.S. et al. Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease. Mol. Cell 42, 172–184 (2011).
Tsutsumi, A., Kawamata, T., Izumi, N., Seitz, H. & Tomari, Y. Recognition of the pre-miRNA structure by Drosophila Dicer-1. Nat. Struct. Mol. Biol. 18, 1153–1158 (2011).
Welker, N.C. et al. Dicer's helicase domain discriminates dsRNA termini to promote an altered reaction mode. Mol. Cell 41, 589–599 (2011).
Zhang, H., Kolb, F.A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Allen, E., Xie, Z., Gustafson, A.M. & Carrington, J.C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005).
Ye, X., Paroo, Z. & Liu, Q. Functional anatomy of the Drosophila microRNA-generating enzyme. J. Biol. Chem. 282, 28373–28378 (2007).
Ma, J.B., Ye, K. & Patel, D.J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004).
Schatz, P.J. Use of peptide libraries to map the substrate specificity of a peptide-modifying enzyme: a 13 residue consensus peptide specifies biotinylation in Escherichia coli. Bio/Technology 11, 1138–1143 (1993).
Howarth, M. et al. A monovalent streptavidin with a single femtomolar biotin binding site. Nat. Methods 3, 267–273 (2006).
MacRae, I.J., Zhou, K. & Doudna, J.A. Structural determinants of RNA recognition and cleavage by Dicer. Nat. Struct. Mol. Biol. 14, 934–940 (2007).
Zou, J., Chang, M., Nie, P. & Secombes, C.J. Origin and evolution of the RIG-I like RNA helicase gene family. BMC Evol. Biol. 9, 85 (2009).
Kowalinski, E. et al. Structural basis for the activation of innate immune pattern-recognition receptor RIG-I by viral RNA. Cell 147, 423–435 (2011).
Soifer, H.S. et al. A role for the Dicer helicase domain in the processing of thermodynamically unstable hairpin RNAs. Nucleic Acids Res. 36, 6511–6522 (2008).
Ma, E., MacRae, I.J., Kirsch, J.F. & Doudna, J.A. Autoinhibition of human dicer by its internal helicase domain. J. Mol. Biol. 380, 237–243 (2008).
Zhang, H., Kolb, F.A., Brondani, V., Billy, E. & Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 21, 5875–5885 (2002).
Kowalinski, E. et al. Structural basis for the activation of innate immune pattern-recognition receptor RIG-I by viral RNA. Cell 147, 423–435 (2011).
Qin, H. et al. Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein–protein interaction. RNA 16, 474–481 (2010).
MacRae, I.J. et al. Structural basis for double-stranded RNA processing by Dicer. Science 311, 195–198 (2006).
Du, Z. et al. Structural and biochemical insights into the dicing mechanism of mouse Dicer: a conserved lysine is critical for dsRNA cleavage. Proc. Natl. Acad. Sci. 105, 2391–2396 (2002).
MacRae, I.J., Ma, E., Zhou, M., Robinson, C.V. & Doudna, J.A. In vitro reconstitution of the human RISC-loading complex. Proc. Natl. Acad. Sci. USA 105, 512–517 (2008).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Yoshioka, C. et al. Automation of random conical tilt and orthogonal tilt data collection using feature-based correlation. J. Struct. Biol. 159, 335–346 (2007).
Lander, G.C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Voss, N.R., Yoshioka, C.K., Radermacher, M., Potter, C.S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009).
Scheres, S.H. et al. Maximum-likelihood multi-reference refinement for electron microscopy images. J. Mol. Biol. 348, 139–149 (2005).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Pintilie, G.D., Zhang, J., Goddard, T.D., Chiu, W. & Gossard, D.C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010).
Acknowledgements
We are grateful to D.K. Simanshu, Y. Tian and D.J. Patel (Memorial Sloan-Kettering Cancer Center) for sharing the human Dicer PAZ-platform structure before publication. EM imaging and reconstruction was conducted at the National Resource for Automated Molecular Microscopy, which is supported by the US National Institutes of Health (NIH) through the P41 program of the National Center for Research Resources (RR017573) and the National Institute of General Medical Sciences (GM103310). This work was also supported by NIH grant R01 GM086701 to I.J.M. P.-W.L. is a predoctoral fellow of the American Heart Association. I.J.M. is a Pew Scholar in the Biomedical Sciences.
Author information
Authors and Affiliations
Contributions
C.S.P., B.C. and I.J.M. conceived of the project. P.-W.L., K.Z.G. and N.D. prepared the samples. P.-W.L. carried out the data collection. P.-W.L., C.S.P., B.C. and I.J.M. analyzed the data. P.-W.L. and I.J.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Methods (PDF 3448 kb)
Rights and permissions
About this article
Cite this article
Lau, PW., Guiley, K., De, N. et al. The molecular architecture of human Dicer. Nat Struct Mol Biol 19, 436–440 (2012). https://doi.org/10.1038/nsmb.2268
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2268
This article is cited by
-
Substrate promiscuity of Dicer toward precursors of the let-7 family and their 3′-end modifications
Cellular and Molecular Life Sciences (2024)
-
Secondary structure RNA elements control the cleavage activity of DICER
Nature Communications (2022)
-
A novel plant-fungal association reveals fundamental sRNA and gene expression reprogramming at the onset of symbiosis
BMC Biology (2021)
-
RNA interference of an orthologue of Dicer of Meloidogyne incognita alludes to the gene’s importance in nematode development
Scientific Reports (2021)
-
RNA and DNA G-quadruplexes bind to human dicer and inhibit its activity
Cellular and Molecular Life Sciences (2021)