Abstract
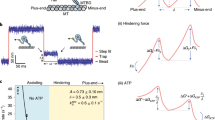
Processivity, the ability of single molecules to move continuously along a track, is a fundamental requirement of cargo-transporting molecular motors. Here, we investigate how cytoplasmic dynein, a homodimeric, microtubule-based motor, achieves processive motion. To do this, we developed a versatile method for assembling Saccharomyces cerevisiae dynein heterodimers, using complementary DNA oligonucleotides covalently linked to dynein monomers labeled with different organic fluorophores. Using two-color, single-molecule microscopy and high-precision, two-dimensional tracking, we find that dynein has a highly variable stepping pattern that is distinct from all other processive cytoskeletal motors, which use 'hand-over-hand' mechanisms. Uniquely, dynein stepping is stochastic when its two motor domains are close together. However, coordination emerges as the distance between motor domains increases, implying that a tension-based mechanism governs these steps. This plasticity may allow tuning of dynein for its diverse cellular functions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Eschbach, J. & Dupuis, L. Cytoplasmic dynein in neurodegeneration. Pharmacol. Ther. 130, 348–363 (2011).
Wynshaw-Boris, A. Lissencephaly and LIS1: insights into the molecular mechanisms of neuronal migration and development. Clin. Genet. 72, 296–304 (2007).
King, S.J. & Schroer, T.A. Dynactin increases the processivity of the cytoplasmic dynein motor. Nat. Cell Biol. 2, 20–24 (2000).
Mallik, R., Carter, B.C., Lex, S.A., King, S.J. & Gross, S.P. Cytoplasmic dynein functions as a gear in response to load. Nature 427, 649–652 (2004).
Reck-Peterson, S.L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Ross, J.L., Wallace, K., Shuman, H., Goldman, Y.E. & Holzbaur, E.L. Processive bidirectional motion of dynein-dynactin complexes in vitro. Nat. Cell Biol. 8, 562–570 (2006).
Toba, S., Watanabe, T.M., Yamaguchi-Okimoto, L., Toyoshima, Y.Y. & Higuchi, H. Overlapping hand-over-hand mechanism of single molecular motility of cytoplasmic dynein. Proc. Natl. Acad. Sci. USA 103, 5741–5745 (2006).
Wang, Z., Khan, S. & Sheetz, M.P. Single cytoplasmic dynein molecule movements: characterization and comparison with kinesin. Biophys. J. 69, 2011–2023 (1995).
Kardon, J.R. & Vale, R.D. Regulators of the cytoplasmic dynein motor. Nat. Rev. Mol. Cell Biol. 10, 854–865 (2009).
Burgess, S.A., Walker, M.L., Sakakibara, H., Knight, P.J. & Oiwa, K. Dynein structure and power stroke. Nature 421, 715–718 (2003).
Kon, T. et al. Helix sliding in the stalk coiled coil of dynein couples ATPase and microtubule binding. Nat. Struct. Mol. Biol. 16, 325–333 (2009).
Roberts, A.J. et al. AAA+ Ring and linker swing mechanism in the dynein motor. Cell 136, 485–495 (2009).
Vale, R.D. Switches, latches, and amplifiers: common themes of G proteins and molecular motors. J. Cell Biol. 135, 291–302 (1996).
Gibbons, I.R. et al. Photosensitized cleavage of dynein heavy chains. Cleavage at the 'V1 site' by irradiation at 365 nm in the presence of ATP and vanadate. J. Biol. Chem. 262, 2780–2786 (1987).
Cho, C., Reck-Peterson, S.L. & Vale, R.D. Regulatory ATPase sites of cytoplasmic dynein affect processivity and force generation. J. Biol. Chem. 283, 25839–25845 (2008).
Kon, T., Nishiura, M., Ohkura, R., Toyoshima, Y.Y. & Sutoh, K. Distinct functions of nucleotide-binding/hydrolysis sites in the four AAA modules of cytoplasmic dynein. Biochemistry 43, 11266–11274 (2004).
Silvanovich, A., Li, M.G., Serr, M., Mische, S. & Hays, T.S. The third P-loop domain in cytoplasmic dynein heavy chain is essential for dynein motor function and ATP-sensitive microtubule binding. Mol. Biol. Cell 14, 1355–1365 (2003).
Carter, A.P., Cho, C., Jin, L. & Vale, R.D. Crystal structure of the dynein motor domain. Science 331, 1159–1165 (2011).
Carter, A.P. et al. Structure and functional role of dynein's microtubule-binding domain. Science 322, 1691–1695 (2008).
Kon, T., Sutoh, K. & Kurisu, G. X-ray structure of a functional full-length dynein motor domain. Nat. Struct. Mol. Biol. 18, 638–642 (2011).
Shima, T., Imamula, K., Kon, T., Ohkura, R. & Sutoh, K. Head-head coordination is required for the processive motion of cytoplasmic dynein, an AAA+ molecular motor. J. Struct. Biol. 156, 182–189 (2006).
Gennerich, A., Carter, A.P., Reck-Peterson, S.L. & Vale, R.D. Force-induced bidirectional stepping of cytoplasmic dynein. Cell 131, 952–965 (2007).
Gennerich, A. & Vale, R.D. Walking the walk: how kinesin and dynein coordinate their steps. Curr. Opin. Cell Biol. 21, 59–67 (2009).
Sellers, J.R. & Veigel, C. Walking with myosin V. Curr. Opin. Cell Biol. 18, 68–73 (2006).
Sweeney, H.L. & Houdusse, A. Myosin VI rewrites the rules for myosin motors. Cell 141, 573–582 (2010).
Yildiz, A. et al. Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. Science 300, 2061–2065 (2003).
Yildiz, A., Tomishige, M., Vale, R.D. & Selvin, P.R. Kinesin walks hand-over-hand. Science 303, 676–678 (2004).
Samsó, M. & Koonce, M.P. 25 Angstrom resolution structure of a cytoplasmic dynein motor reveals a seven-member planar ring. J. Mol. Biol. 340, 1059–1072 (2004).
Su, X. et al. Mechanisms underlying the dual-mode regulation of microtubule dynamics by Kip3/kinesin8. Mol. Cell 43, 751–763 (2011).
Ray, S., Meyhofer, E., Milligan, R.A. & Howard, J. Kinesin follows the microtubule's protofilament axis. J. Cell Biol. 121, 1083–1093 (1993).
Ray, S., Wolf, S.G., Howard, J. & Downing, K.H. Kinesin does not support the motility of zinc-macrotubes. Cell Motil. Cytoskeleton 30, 146–152 (1995).
Banaszynski, L.A., Liu, C.W. & Wandless, T.J. Characterization of the FKBP.rapamycin.FRB ternary complex. J. Am. Chem. Soc. 127, 4715–4721 (2005).
Markham, N.R. & Zuker, M. DINAMelt web server for nucleic acid melting prediction. Nucleic Acids Res. 33, W577–W581 (2005).
Miyazono, Y., Hayashi, M., Karagiannis, P., Harada, Y. & Tadakuma, H. Strain through the neck linker ensures processive runs: a DNA-kinesin hybrid nanomachine study. EMBO J. 29, 93–106 (2010).
Essevaz-Roulet, B., Bockelmann, U. & Heslot, F. Mechanical separation of the complementary strands of DNA. Proc. Natl. Acad. Sci. USA 94, 11935–11940 (1997).
Thompson, R.E., Larson, D.R. & Webb, W.W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Churchman, L.S., Okten, Z., Rock, R.S., Dawson, J.F. & Spudich, J.A. Single molecule high-resolution colocalization of Cy3 and Cy5 attached to macromolecules measures intramolecular distances through time. Proc. Natl. Acad. Sci. USA 102, 1419–1423 (2005).
Yildiz, A., Tomishige, M., Gennerich, A. & Vale, R.D. Intramolecular strain coordinates kinesin stepping behavior along microtubules. Cell 134, 1030–1041 (2008).
Rosenfeld, S.S. & Sweeney, H.L. A model of myosin V processivity. J. Biol. Chem. 279, 40100–40111 (2004).
Purcell, T.J., Sweeney, H.L. & Spudich, J.A. A force-dependent state controls the coordination of processive myosin V. Proc. Natl. Acad. Sci. USA 102, 13873–13878 (2005).
Veigel, C., Schmitz, S., Wang, F. & Sellers, J.R. Load-dependent kinetics of myosin-V can explain its high processivity. Nat. Cell Biol. 7, 861–869 (2005).
Sweeney, H.L. et al. How myosin VI coordinates its heads during processive movement. EMBO J. 26, 2682–2692 (2007).
Dunn, A.R., Chuan, P., Bryant, Z. & Spudich, J.A. Contribution of the myosin VI tail domain to processive stepping and intramolecular tension sensing. Proc. Natl. Acad. Sci. USA 107, 7746–7750 (2010).
Baboolal, T.G. et al. The SAH domain extends the functional length of the myosin lever. Proc. Natl. Acad. Sci. USA 106, 22193–22198 (2009).
Elting, M.W., Bryant, Z., Liao, J.C. & Spudich, J.A. Detailed tuning of structure and intramolecular communication are dispensable for processive motion of myosin VI. Biophys. J. 100, 430–439 (2011).
Numata, N., Shima, T., Ohkura, R., Kon, T. & Sutoh, K. C-sequence of the Dictyostelium cytoplasmic dynein participates in processivity modulation. FEBS Lett. 585, 1185–1190 (2011).
Dixit, R., Ross, J.L., Goldman, Y.E. & Holzbaur, E.L. Differential regulation of dynein and kinesin motor proteins by tau. Science 319, 1086–1089 (2008).
Ross, J.L., Shuman, H., Holzbaur, E.L. & Goldman, Y.E. Kinesin and dynein-dynactin at intersecting microtubules: motor density affects dynein function. Biophys. J. 94, 3115–3125 (2008).
Lawrence, C.J., Morris, N.R., Meagher, R.B. & Dawe, R.K. Dyneins have run their course in plant lineage. Traffic 2, 362–363 (2001).
Wickstead, B. & Gull, K. Dyneins across eukaryotes: a comparative genomic analysis. Traffic 8, 1708–1721 (2007).
Ishikawa, T., Sakakibara, H. & Oiwa, K. The architecture of outer dynein arms in situ. J. Mol. Biol. 368, 1249–1258 (2007).
Mizuno, N., Narita, A., Kon, T., Sutoh, K. & Kikkawa, M. Three-dimensional structure of cytoplasmic dynein bound to microtubules. Proc. Natl. Acad. Sci. USA 104, 20832–20837 (2007).
Nicastro, D. et al. The molecular architecture of axonemes revealed by cryoelectron tomography. Science 313, 944–948 (2006).
Ueno, H., Yasunaga, T., Shingyoji, C. & Hirose, K. Dynein pulls microtubules without rotating its stalk. Proc. Natl. Acad. Sci. USA 105, 19702–19707 (2008).
Acknowledgements
We thank X. Su and D. Pellman (Harvard Medical School) for providing purified kinesin-8; S. Zou for technical assistance; A. Carter, S. Churchman, A. Gennerich, Y. Goldman, A. Hendricks, J. Huang, A. Leschziner and A. Roberts for critical comments on the manuscript; F. Aguet, A. Besser, M. Vilela and G. Danuser for discussions of data analysis; M. Bagonis for early work on oligomer-SNAP linking; A. Leschziner for help with figure design; and A. Carter for providing MATLAB code. W.Q. is supported by a postdoctoral fellowship from the American Heart Association. S.L.R.-P. is funded by the Rita Allen Foundation, the Harvard Armenise Foundation and a US National Institutes of Health New Innovator award (1 DP2 OD004268-01).
Author information
Authors and Affiliations
Contributions
W.Q. and N.D.D. contributed equally. W.Q., N.D.D., W.S. and S.L.R.-P. designed the experiments. W.Q., N.D.D. and B.S.G. conducted the experiments and analyzed the data. W.Q., N.D.D., B.S.G. and S.L.R.-P. wrote the paper. E.V. and D.W. wrote the two-dimensional particle tracking code.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 5618 kb)
Rights and permissions
About this article
Cite this article
Qiu, W., Derr, N., Goodman, B. et al. Dynein achieves processive motion using both stochastic and coordinated stepping. Nat Struct Mol Biol 19, 193–200 (2012). https://doi.org/10.1038/nsmb.2205
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2205
This article is cited by
-
Lis1 slows force-induced detachment of cytoplasmic dynein from microtubules
Nature Chemical Biology (2024)
-
Regulatory mechanisms of the dynein-2 motility by post-translational modification revealed by MD simulation
Scientific Reports (2023)
-
Effect of detachment of motor protein from track on its transport
Journal of Biological Physics (2022)
-
A synthetic tubular molecular transport system
Nature Communications (2021)
-
One Dimensional Exclusion Process with Dynein Inspired Hops: Simulation and Mean Field Analysis
Journal of Statistical Physics (2021)