Abstract
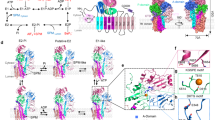
DNA ligases finalize DNA replication and repair through DNA nick-sealing reactions that can abort to generate cytotoxic 5′-adenylation DNA damage. Aprataxin (Aptx) catalyzes direct reversal of 5′-adenylate adducts to protect genome integrity. Here the structure of a Schizosaccharomyces pombe Aptx–DNA–AMP–Zn2+ complex reveals active site and DNA interaction clefts formed by fusing a histidine triad (HIT) nucleotide hydrolase with a DNA minor groove–binding C2HE zinc finger (Znf). An Aptx helical 'wedge' interrogates the base stack for sensing DNA ends or DNA nicks. The HIT-Znf, the wedge and an '[F/Y]PK' pivot motif cooperate to distort terminal DNA base-pairing and direct 5′-adenylate into the active site pocket. Structural and mutational data support a wedge-pivot-cut HIT-Znf catalytic mechanism for 5′-adenylate adduct recognition and removal and suggest that mutations affecting protein folding, the active site pocket and the pivot motif underlie Aptx dysfunction in the neurodegenerative disorder ataxia with oculomotor apraxia 1 (AOA1).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
16 October 2011
In the HTML version of this article initially published online, the corresponding author was given as Jessica S. Williams, instead of R. Scott Willliams. The error has been corrected in the HTML version of this article.
References
Pascal, J.M., O'Brien, P.J., Tomkinson, A.E. & Ellenberger, T. Human DNA ligase I completely encircles and partially unwinds nicked DNA. Nature 432, 473–478 (2004).
Ellenberger, T. & Tomkinson, A.E. Eukaryotic DNA ligases: structural and functional insights. Annu. Rev. Biochem. 77, 313–338 (2008).
Ahel, I. et al. The neurodegenerative disease protein aprataxin resolves abortive DNA ligation intermediates. Nature 443, 713–716 (2006).
Reynolds, J.J. et al. Defective DNA ligation during short-patch single-strand break repair in ataxia oculomotor apraxia 1. Mol. Cell. Biol. 29, 1354–1362 (2009).
Harris, J.L. et al. Aprataxin, poly-ADP ribose polymerase 1 (PARP-1) and apurinic endonuclease 1 (APE1) function together to protect the genome against oxidative damage. Hum. Mol. Genet. 18, 4102–4117 (2009).
Date, H. et al. Early-onset ataxia with ocular motor apraxia and hypoalbuminemia is caused by mutations in a new HIT superfamily gene. Nat. Genet. 29, 184–188 (2001).
Moreira, M.C. et al. The gene mutated in ataxia-ocular apraxia 1 encodes the new HIT/Zn-finger protein aprataxin. Nat. Genet. 29, 189–193 (2001).
Quinzii, C.M. et al. Coenzyme Q deficiency and cerebellar ataxia associated with an aprataxin mutation. Neurology 64, 539–541 (2005).
Baba, Y. et al. Aprataxin (APTX) gene mutations resembling multiple system atrophy. Parkinsonism Relat. Disord. 13, 139–142 (2007).
Hirano, M. et al. DNA single-strand break repair is impaired in aprataxin-related ataxia. Ann. Neurol. 61, 162–174 (2007).
Mosesso, P. et al. The novel human gene aprataxin is directly involved in DNA single-strand-break repair. Cell. Mol. Life Sci. 62, 485–491 (2005).
Daley, J.M., Wilson, T.E. & Ramotar, D. Genetic interactions between HNT3/Aprataxin and RAD27/FEN1 suggest parallel pathways for 5′ end processing during base excision repair. DNA Repair (Amst.) 9, 690–699 (2010).
Deshpande, G.P. et al. Screening a genome-wide S. pombe deletion library identifies novel genes and pathways involved in genome stability maintenance. DNA Repair (Amst.) 8, 672–679 (2009).
Caldecott, K.W. Single-strand break repair and genetic disease. Nat. Rev. Genet. 9, 619–631 (2008).
Rass, U., Ahel, I. & West, S.C. Actions of aprataxin in multiple DNA repair pathways. J. Biol. Chem. 282, 9469–9474 (2007).
Lima, C.D., Klein, M.G. & Hendrickson, W.A. Structure-based analysis of catalysis and substrate definition in the HIT protein family. Science 278, 286–290 (1997).
Brenner, C. Hint, Fhit, and GalT: function, structure, evolution, and mechanism of three branches of the histidine triad superfamily of nucleotide hydrolases and transferases. Biochemistry 41, 9003–9014 (2002).
Rass, U., Ahel, I. & West, S.C. Molecular mechanism of DNA deadenylation by the neurological disease protein aprataxin. J. Biol. Chem. 283, 33994–34001 (2008).
Clements, P.M. et al. The ataxia-oculomotor apraxia 1 gene product has a role distinct from ATM and interacts with the DNA strand break repair proteins XRCC1 and XRCC4. DNA Repair (Amst.) 3, 1493–1502 (2004).
Becherel, O.J. et al. CK2 phosphorylation-dependent interaction between aprataxin and MDC1 in the DNA damage response. Nucleic Acids Res. 38, 1489–1503 (2010).
Lima, C.D. et al. MAD analysis of FHIT, a putative human tumor suppressor from the HIT protein family. Structure 5, 763–774 (1997).
Pavletich, N.P. & Pabo, C.O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Klug, A. The discovery of zinc fingers and their applications in gene regulation and genome manipulation. Annu. Rev. Biochem. 79, 213–231 (2010).
Takahashi, T. et al. Aprataxin, causative gene product for EAOH/AOA1, repairs DNA single-strand breaks with damaged 3′-phosphate and 3′-phosphoglycolate ends. Nucleic Acids Res. 35, 3797–3809 (2007).
Kijas, A.W., Harris, J.L., Harris, J.M. & Lavin, M.F. Aprataxin forms a discrete branch in the HIT (histidine triad) superfamily of proteins with both DNA/RNA binding and nucleotide hydrolase activities. J. Biol. Chem. 281, 13939–13948 (2006).
Tranchant, C., Fleury, M., Moreira, M.C., Koenig, M. & Warter, J.M. Phenotypic variability of aprataxin gene mutations. Neurology 60, 868–870 (2003).
Huffman, J.L., Sundheim, O. & Tainer, J.A. DNA base damage recognition and removal: new twists and grooves. Mutat. Res. 577, 55–76 (2005).
Wilson, S.H. & Kunkel, T.A. Passing the baton in base excision repair. Nat. Struct. Biol. 7, 176–178 (2000).
Bernstein, N.K. et al. The molecular architecture of the mammalian DNA repair enzyme, polynucleotide kinase. Mol. Cell 17, 657–670 (2005).
Klug, A. The discovery of zinc fingers and their development for practical applications in gene regulation and genome manipulation. Q. Rev. Biophys. 43, 1–21 (2010).
Dopeso, H. et al. Aprataxin tumor levels predict response of colorectal cancer patients to irinotecan-based treatment. Clin. Cancer Res. 16, 2375–2382 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Cowtan, K. dm: an automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newslett. on Protein Crystallogr. 31, 34–38 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Acknowledgements
This research was supported by the intramural research program of the US National Institutes of Health (NIH), National Institute of Environmental Health Sciences (NIEHS) (1Z01ES102765-01 to R.S.W.). We thank L. Pedersen, T. Kunkel and S. Wilson for discussions and critical reading of the manuscript, the Advanced Photon Source (APS) Southeast Regional Collaborative Access Team (SER-CAT) staff for assistance with crystallographic data collection, and we thank J. Williams of the NIEHS Protein Microcharacterization Core Facility for mass spectrometry analysis.
Author information
Authors and Affiliations
Contributions
P.T. conducted and analyzed experiments and helped write the manuscript. C.D.A., R.K. and J.S.W. conducted experiments. P.D.R. and J.K. analyzed results. I.A. designed experiments and analyzed results. R.S.W. designed research, did experiments, analyzed results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 3010 kb)
Supplementary Movie 1
Aptx engagement of a nicked or gapped DNA-adenylate. Aptx employs the helical wedge with Phe34 (Blue) to displace a stacked 5′-adenylate (orange). The DNA (green) morphs between a model B-DNA conformation and the backbone conformation observed in the Aptx–DNA–AMP–Zn complex structure. A slight under-winding of the duplex is observed upon binding. Grey DNA is a modeled conformation of the predicted positioning of the upstream region of nick or gapped duplex bearing a 5′-AMP. Intercalation of the wedge helix into the base stack necessitates displacement of the upstream DNA. HIT domain is shown in purple and Znf in gold/brown. (MOV 21034 kb)
Rights and permissions
About this article
Cite this article
Tumbale, P., Appel, C., Kraehenbuehl, R. et al. Structure of an aprataxin–DNA complex with insights into AOA1 neurodegenerative disease. Nat Struct Mol Biol 18, 1189–1195 (2011). https://doi.org/10.1038/nsmb.2146
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2146
This article is cited by
-
Molecular basis for DarT ADP-ribosylation of a DNA base
Nature (2021)
-
Two-tiered enforcement of high-fidelity DNA ligation
Nature Communications (2019)
-
Aprataxin resolves adenylated RNA–DNA junctions to maintain genome integrity
Nature (2014)
-
Mechanism of repair of 5′-topoisomerase II–DNA adducts by mammalian tyrosyl-DNA phosphodiesterase 2
Nature Structural & Molecular Biology (2012)