Abstract
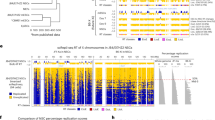
Heterochromatin assembly at Schizosaccharomyces pombe centromeres involves a self-reinforcing loop mechanism wherein chromatin-bound RNAi factors facilitate targeting of Clr4–Rik1 methyltransferase. However, the initial nucleation of heterochromatin has remained elusive. We show that cells lacking Mlo3, a protein involved in mRNP biogenesis and RNA quality control, assemble functional heterochromatin in RNAi-deficient cells. Heterochromatin restoration is linked to RNA surveillance because loss of Mlo3-associated TRAMP also rescues heterochromatin defects of RNAi mutants. mlo3Δ, which causes accumulation of bidirectional repeat-transcripts, restores Rik1 enrichment at repeats and triggers de novo heterochromatin formation in the absence of RNAi. RNAi-independent heterochromatin nucleation occurs at selected euchromatic loci that show upregulation of antisense RNAs in mlo3Δ cells. We find that the exosome RNA degradation machinery acts parallel to RNAi to promote heterochromatin formation at centromeres. These results suggest that RNAi-independent mechanisms exploit transcription and non-coding RNAs to nucleate heterochromatin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Grewal, S.I. & Elgin, S.C. Transcription and RNA interference in the formation of heterochromatin. Nature 447, 399–406 (2007).
Ekwall, K. Epigenetic control of centromere behavior. Annu. Rev. Genet. 41, 63–81 (2007).
Cam, H.P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Noma, K. et al. RITS acts in cis to promote RNA interference–mediated transcriptional and post-transcriptional silencing. Nat. Genet. 36, 1174–1180 (2004).
Schalch, T. et al. High-affinity binding of Chp1 chromodomain to K9 methylated histone H3 is required to establish centromeric heterochromatin. Mol. Cell 34, 36–46 (2009).
Bayne, E.H. et al. Stc1: a critical link between RNAi and chromatin modification required for heterochromatin integrity. Cell 140, 666–677 (2010).
Zhang, K., Mosch, K., Fischle, W. & Grewal, S.I. Roles of the Clr4 methyltransferase complex in nucleation, spreading and maintenance of heterochromatin. Nat. Struct. Mol. Biol. 15, 381–388 (2008).
Sugiyama, T., Cam, H., Verdel, A., Moazed, D. & Grewal, S.I. RNA-dependent RNA polymerase is an essential component of a self-enforcing loop coupling heterochromatin assembly to siRNA production. Proc. Natl. Acad. Sci. USA 102, 152–157 (2005).
Fischer, T. et al. Diverse roles of HP1 proteins in heterochromatin assembly and functions in fission yeast. Proc. Natl. Acad. Sci. USA 106, 8998–9003 (2009).
Sugiyama, T. et al. SHREC, an effector complex for heterochromatic transcriptional silencing. Cell 128, 491–504 (2007).
Motamedi, M.R. et al. HP1 proteins form distinct complexes and mediate heterochromatic gene silencing by nonoverlapping mechanisms. Mol. Cell 32, 778–790 (2008).
Yamane, K. et al. Asf1/HIRA facilitate global histone deacetylation and associate with HP1 to promote nucleosome occupancy at heterochromatic loci. Mol. Cell 41, 56–66 (2011).
Chen, E.S. et al. Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451, 734–737 (2008).
Kloc, A., Zaratiegui, M., Nora, E. & Martienssen, R. RNA interference guides histone modification during the S phase of chromosomal replication. Curr. Biol. 18, 490–495 (2008).
Djupedal, I. et al. RNA Pol II subunit Rpb7 promotes centromeric transcription and RNAi-directed chromatin silencing. Genes Dev. 19, 2301–2306 (2005).
Kato, H. et al. RNA polymerase II is required for RNAi-dependent heterochromatin assembly. Science 309, 467–469 (2005).
Bayne, E.H. et al. Splicing factors facilitate RNAi-directed silencing in fission yeast. Science 322, 602–606 (2008).
Smith, E. & Shilatifard, A. The chromatin signaling pathway: diverse mechanisms of recruitment of histone-modifying enzymes and varied biological outcomes. Mol. Cell 40, 689–701 (2010).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003).
Strässer, K. et al. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417, 304–308 (2002).
Houseley, J., LaCava, J. & Tollervey, D. RNA-quality control by the exosome. Nat. Rev. Mol. Cell Biol. 7, 529–539 (2006).
Wang, S.W., Stevenson, A.L., Kearsey, S.E., Watt, S. & Bahler, J. Global role for polyadenylation-assisted nuclear RNA degradation in posttranscriptional gene silencing. Mol. Cell. Biol. 28, 656–665 (2008).
Bühler, M., Haas, W., Gygi, S.P. & Moazed, D. RNAi-dependent and -independent RNA turnover mechanisms contribute to heterochromatic gene silencing. Cell 129, 707–721 (2007).
Halic, M. & Moazed, D. Dicer-independent primal RNAs trigger RNAi and heterochromatin formation. Cell 140, 504–516 (2010).
Thakurta, A.G., Gopal, G., Yoon, J.H., Kozak, L. & Dhar, R. Homolog of BRCA2-interacting Dss1p and Uap56p link Mlo3p and Rae1p for mRNA export in fission yeast. EMBO J. 24, 2512–2523 (2005).
Bernard, P. & Allshire, R. Centromeres become unstuck without heterochromatin. Trends Cell Biol. 12, 419–424 (2002).
Hall, I.M., Noma, K. & Grewal, S.I. RNA interference machinery regulates chromosome dynamics during mitosis and meiosis in fission yeast. Proc. Natl. Acad. Sci. USA 100, 193–198 (2003).
Provost, P. et al. Dicer is required for chromosome segregation and gene silencing in fission yeast cells. Proc. Natl. Acad. Sci. USA 99, 16648–16653 (2002).
Zhang, K. et al. Clr4/Suv39 and RNA quality control factors cooperate to trigger RNAi and suppress antisense RNA. Science 331, 1624–1627 (2011).
Yamada, T., Fischle, W., Sugiyama, T., Allis, C.D. & Grewal, S.I. The nucleation and maintenance of heterochromatin by a histone deacetylase in fission yeast. Mol. Cell 20, 173–185 (2005).
Kulish, D. & Struhl, K. TFIIS enhances transcriptional elongation through an artificial arrest site in vivo. Mol. Cell. Biol. 21, 4162–4168 (2001).
Williams, L.A. & Kane, C.M. Isolation and characterization of the Schizosaccharomyces pombe gene encoding transcript elongation factor TFIIS. Yeast 12, 227–236 (1996).
Kvint, K. et al. Reversal of RNA polymerase II ubiquitylation by the ubiquitin protease Ubp3. Mol. Cell 30, 498–506 (2008).
Iida, T., Nakayama, J. & Moazed, D. siRNA-mediated heterochromatin establishment requires HP1 and is associated with antisense transcription. Mol. Cell 31, 178–189 (2008).
Simmer, F. et al. Hairpin RNA induces secondary small interfering RNA synthesis and silencing in trans in fission yeast. EMBO Rep. 11, 112–118 (2010).
Hilleren, P., McCarthy, T., Rosbash, M., Parker, R. & Jensen, T.H. Quality control of mRNA 3′-end processing is linked to the nuclear exosome. Nature 413, 538–542 (2001).
Libri, D. et al. Interactions between mRNA export commitment, 3′-end quality control, and nuclear degradation. Mol. Cell. Biol. 22, 8254–8266 (2002).
Grewal, S.I. & Klar, A.J. A recombinationally repressed region between mat2 and mat3 loci shares homology to centromeric repeats and regulates directionality of mating-type switching in fission yeast. Genetics 146, 1221–1238 (1997).
Hall, I.M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Ponting, C.P., Oliver, P.L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Bonasio, R., Tu, S. & Reinberg, D. Molecular signals of epigenetic states. Science 330, 612–616 (2010).
Sadaie, M., Iida, T., Urano, T. & Nakayama, J. A chromodomain protein, Chp1, is required for the establishment of heterochromatin in fission yeast. EMBO J. 23, 3825–3835 (2004).
Shanker, S. et al. Continuous requirement for the Clr4 complex but not RNAi for centromeric heterochromatin assembly in fission yeast harboring a disrupted RITS complex. PLoS Genet. 6, e1001174 (2010).
Moshkovich, N. & Lei, E.P. HP1 recruitment in the absence of argonaute proteins in Drosophila. PLoS Genet. 6, e1000880 (2010).
Freitag, M. et al. DNA methylation is independent of RNA interference in Neurospora. Science 304, 1939 (2004).
Henderson, I.R. & Jacobsen, S.E. Epigenetic inheritance in plants. Nature 447, 418–424 (2007).
Kavi, H.H. & Birchler, J.A. Interaction of RNA polymerase II and the small RNA machinery affects heterochromatic silencing in Drosophila. Epigenetics Chromatin 2, 15 (2009).
Guang, S. et al. Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription. Nature 465, 1097–1101 (2010).
Orphanides, G. & Reinberg, D. RNA polymerase II elongation through chromatin. Nature 407, 471–475 (2000).
Li, X. & Manley, J.L. Cotranscriptional processes and their influence on genome stability. Genes Dev. 20, 1838–1847 (2006).
Pal-Bhadra, M., Bhadra, U. & Birchler, J.A. RNAi related mechanisms affect both transcriptional and posttranscriptional transgene silencing in Drosophila. Mol. Cell 9, 315–327 (2002).
Bühler, M., Verdel, A. & Moazed, D. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125, 873–886 (2006).
Lee, J.T. Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome. Genes Dev. 23, 1831–1842 (2009).
Sharp, P.A. The centrality of RNA. Cell 136, 577–580 (2009).
Matzke, M.A. & Birchler, J.A. RNAi-mediated pathways in the nucleus. Nat. Rev. Genet. 6, 24–35 (2005).
Maison, C. et al. SUMOylation promotes de novo targeting of HP1α to pericentric heterochromatin. Nat. Genet. 43, 220–227 (2011).
Chapman, R.D. et al. Transcribing RNA polymerase II is phosphorylated at CTD residue serine-7. Science 318, 1780–1782 (2007).
Zofall, M. et al. Histone H2A.Z cooperates with RNAi and heterochromatin factors to suppress antisense RNAs. Nature 461, 419–422 (2009).
Gilbert, C. & Svejstrup, J.Q. RNA immunoprecipitation for determining RNA-protein associations in vivo. Curr. Protoc. Mol. Biol. 27, 27.4 (2006).
Acknowledgements
We are thankful to D. Eick (Helmholtz Center Munich) for the gift of phospho (Ser2) RNAPII antibody, R. Dhar and N. Krogan (University of California, San Francisco) for strains, J. Dhakshnamoorthy, N. Komissarova and S. Mehta for helpful contributions, and members of the Grewal laboratory for discussions. This research was supported by the Intramural Research Program of the US National Institutes of Health, National Cancer Institute, Center for Cancer Research.
Author information
Authors and Affiliations
Contributions
F.E.R.-T., K.Z. and S.I.S.G. designed the research. K.Z., F.E.R.-T. and M.Z. conducted the experiments. E.C. contributed the reagents. F.E.R.-T., K.Z. and S.I.S.G. analyzed the data. F.E.R.-T. and S.I.S.G. wrote the paper. F.E.R.-T., K.Z. and S.I.S.G. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Methods (PDF 9646 kb)
Rights and permissions
About this article
Cite this article
Reyes-Turcu, F., Zhang, K., Zofall, M. et al. Defects in RNA quality control factors reveal RNAi-independent nucleation of heterochromatin. Nat Struct Mol Biol 18, 1132–1138 (2011). https://doi.org/10.1038/nsmb.2122
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2122
This article is cited by
-
Dicer promotes genome stability via the bromodomain transcriptional co-activator BRD4
Nature Communications (2022)
-
Regulation of long non-coding RNAs and genome dynamics by the RNA surveillance machinery
Nature Reviews Molecular Cell Biology (2020)
-
The nascent RNA binding complex SFiNX licenses piRNA-guided heterochromatin formation
Nature Structural & Molecular Biology (2019)
-
Epigenetica: passepartout della doppia elica?
La Rivista Italiana della Medicina di Laboratorio - Italian Journal of Laboratory Medicine (2018)
-
The 19S proteasome regulates subtelomere silencing and facultative heterochromatin formation in fission yeast
Current Genetics (2018)