Abstract
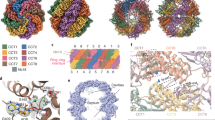
Protein folding is assisted by molecular chaperones. CCT (chaperonin containing TCP-1, or TRiC) is a 1-MDa oligomer that is built by two rings comprising eight different 60-kDa subunits. This chaperonin regulates the folding of important proteins including actin, α-tubulin and β-tubulin. We used an electron density map at 5.5 Å resolution to reconstruct CCT, which showed a substrate in the inner cavities of both rings. Here we present the crystal structure of the open conformation of this nanomachine in complex with tubulin, providing information about the mechanism by which it aids tubulin folding. The structure showed that the substrate interacts with loops in the apical and equatorial domains of CCT. The organization of the ATP-binding pockets suggests that the substrate is stretched inside the cavity. Our data provide the basis for understanding the function of this chaperonin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mogk, A., Bukau, B. & Deuerling, E. in Cellular Functions of Cytosolic E. coli Chaperones (ed. Lund, P.) 1–34 (Oxford Univ. Press, Oxford, 2001).
Braig, K. et al. The crystal structure of the bacterial chaperonin GroEL at 2.8 Å. Nature 371, 578–586 (1994).
Ditzel, L. et al. Crystal structure of the thermosome, the archaeal chaperonin and homolog of CCT. Cell 93, 125–138 (1998).
Valpuesta, J.M., Carrascosa, J.L. & Willison, K.R. in Structure and Function of the Cytosolic Chaperonin CCT (eds. Buchner, J. & Kiefhaber, T.) 725–755 (Wiley-VCH, Weinheim, 2005).
Llorca, O. et al. Eukaryotic chaperonin CCT stabilizes actin and tubulin folding intermediates in open quasi-native conformations. EMBO J. 19, 5971–5979 (2000).
Llorca, O. et al. Eukaryotic type II chaperonin CCT interacts with actin through specific subunits. Nature 402, 693–696 (1999).
Lin, Y.F., Tsai, W.P., Liu, H.G. & Liang, P.H. Intracellular beta-tubulin/chaperonin containing TCP1-beta complex serves as a novel chemotherapeutic target against drug-resistant tumors. Cancer Res. 69, 6879–6888 (2009).
Zhang, J. et al. Mechanism of folding chamber closure in a group II chaperonin. Nature 463, 379–383 (2010).
Cong, Y. et al. 4.0-A resolution cryo-EM structure of the mammalian chaperonin TRiC/CCT reveals its unique subunit arrangement. Proc. Natl. Acad. Sci. USA 107, 4967–4972 (2010).
Kubota, S., Kubota, H. & Nagata, K. Cytosolic chaperonin protects folding intermediates of Gbeta from aggregation by recognizing hydrophobic beta-strands. Proc. Natl. Acad. Sci. USA 103, 8360–8365 (2006).
Yam, A.Y. et al. Defining the TRiC/CCT interactome links chaperonin function to stabilization of newly made proteins with complex topologies. Nat. Struct. Mol. Biol. 15, 1255–1262 (2008).
Ban, N. et al. Placement of protein and RNA structures into a 5 Å-resolution map of the 50S ribosomal subunit. Nature 400, 841–847 (1999).
Liou, A.K. & Willison, K.R. Elucidation of the subunit orientation in CCT (chaperonin containing TCP1) from the subunit composition of CCT micro-complexes. EMBO J. 16, 4311–4316 (1997).
Martín-Benito, J. et al. The inter-ring arrangement of the cytosolic chaperonin CCT. EMBO Rep. 8, 252–257 (2007).
Meyer, A.S. et al. Closing the folding chamber of the eukaryotic chaperonin requires the transition state of ATP hydrolysis. Cell 113, 369–381 (2003).
Spiess, C., Meyer, A.S., Reissmann, S. & Frydman, J. Mechanism of the eukaryotic chaperonin: protein folding in the chamber of secrets. Trends Cell Biol. 14, 598–604 (2004).
Rivenzon-Segal, D., Wolf, S.G., Shimon, L., Willison, K.R. & Horovitz, A. Sequential ATP-induced allosteric transitions of the cytoplasmic chaperonin containing TCP-1 revealed by EM analysis. Nat. Struct. Mol. Biol. 12, 233–237 (2005).
Villebeck, L., Moparthi, S.B., Lindgren, M., Hammarstrom, P. & Jonsson, B.H. Domain-specific chaperone-induced expansion is required for beta-actin folding: a comparison of beta-actin conformations upon interactions with GroEL and tail-less complex polypeptide 1 ring complex (TRiC). Biochemistry 46, 12639–12647 (2007).
Lin, P. & Sherman, F. The unique hetero-oligomeric nature of the subunits in the catalytic cooperativity of the yeast Cct chaperonin complex. Proc. Natl. Acad. Sci. USA 94, 10780–10785 (1997).
Melki, R., Batelier, G., Soulie, S. & Williams, R.C. Jr. Cytoplasmic chaperonin containing TCP-1: structural and functional characterization. Biochemistry 36, 5817–5826 (1997).
Llorca, O., Carrascosa, J.L. & Valpuesta, J.M. Biochemical characterization of symmetric GroEL-GroES complexes. Evidence for a role in protein folding. J. Biol. Chem. 271, 68–76 (1996).
Clare, D.K., Bakkes, P.J., van Heerikhuizen, H., van der Vies, S.M. & Saibil, H.R. Chaperonin complex with a newly folded protein encapsulated in the folding chamber. Nature 457, 107–110 (2009).
Ritco-Vonsovici, M. & Willison, K.R. Defining the eukaryotic cytosolic chaperonin-binding sites in human tubulins. J. Mol. Biol. 304, 81–98 (2000).
Oubridge, C., Krummel, D.A., Leung, A.K., Li, J. & Nagai, K. Interpreting a low resolution map of human U1 snRNP using anomalous scatterers. Structure 17, 930–938 (2009).
Booth, C.R. et al. Mechanism of lid closure in the eukaryotic chaperonin TRiC/CCT. Nat. Struct. Mol. Biol. 15, 746–753 (2008).
Iizuka, R. et al. Role of the helical protrusion in the conformational change and molecular chaperone activity of the archaeal group II chaperonin. J. Biol. Chem. 279, 18834–18839 (2004).
Reissmann, S., Parnot, C., Booth, C.R., Chiu, W. & Frydman, J. Essential function of the built-in lid in the allosteric regulation of eukaryotic and archaeal chaperonins. Nat. Struct. Mol. Biol. 14, 432–440 (2007).
Heller, M. et al. NMR studies on the substrate-binding domains of the thermosome: structural plasticity in the protrusion region. J. Mol. Biol. 336, 717–729 (2004).
Camasses, A., Bogdanova, A., Shevchenko, A. & Zachariae, W. The CCT chaperonin promotes activation of the anaphase-promoting complex through the generation of functional Cdc20. Mol. Cell 12, 87–100 (2003).
Liu, X. et al. CCT chaperonin complex is required for the biogenesis of functional Plk1. Mol. Cell. Biol. 25, 4993–5010 (2005).
Won, K.A., Schumacher, R.J., Farr, G.W., Horwich, A.L. & Reed, S.I. Maturation of human cyclin E requires the function of eukaryotic chaperonin CCT. Mol. Cell. Biol. 18, 7584–7589 (1998).
Frydman, J. et al. Function in protein folding of TRiC, a cytosolic ring complex containing TCP-1 and structurally related subunits. EMBO J. 11, 4767–4778 (1992).
Gao, Y., Thomas, J.O., Chow, R.L., Lee, G.H. & Cowan, N.J. A cytoplasmic chaperonin that catalyzes beta-actin folding. Cell 69, 1043–1050 (1992).
Grantham, J., Brackley, K.I. & Willison, K.R. Substantial CCT activity is required for cell cycle progression and cytoskeletal organization in mammalian cells. Exp. Cell Res. 312, 2309–2324 (2006).
Amit, M. et al. Equivalent mutations in the eight subunits of the chaperonin CCT produce dramatically different cellular and gene expression phenotypes. J. Mol. Biol. 401, 532–543 (2010).
Boisvert, D.C., Wang, J., Otwinowski, Z., Horwich, A.L. & Sigler, P.B. The 2.4 Å crystal structure of the bacterial chaperonin GroEL complexed with ATP gamma S. Nat. Struct. Biol. 3, 170–177 (1996).
Martín-Benito, J. et al. Structure of eukaryotic prefoldin and of its complexes with unfolded actin and the cytosolic chaperonin CCT. EMBO J. 21, 6377–6386 (2002).
Kabsch, W. Automatic indexing of rotation diffraction patterns. J. Appl. Crystallogr. 21, 67–71 (1988).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
de la Fortelle, E. & Bricogne, G. Macromolecular Crystallography 472–494 (Academic, New York, 1997).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Cowtan, K.D., Zhang, K.Y.J. & Main, P. Crystallography of Biological Macromolecules 705–710 (Kluwer, 2001).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Sorzano, C.O. et al. XMIPP: a new generation of an open-source image processing package for electron microscopy. J. Struct. Biol. 148, 194–204 (2004).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Scheres, S.H., Valle, M., Grob, P., Nogales, E. & Carazo, J.M. Maximum likelihood refinement of electron microscopy data with normalization errors. J. Struct. Biol. 166, 234–240 (2009).
Braig, K., Menz, R.I., Montgomery, M.G., Leslie, A.G. & Walker, J.E. Structure of bovine mitochondrial F(1)-ATPase inhibited by Mg2+ ADP and aluminium fluoride. Structure 8, 567–573 (2000).
Acknowledgements
We thank the Swiss Light Source and European Synchrotron Radiation Facility beamline staff for their support. Funding was obtained through Ministerio de Ciencia e Innovación (MICINN) grants BFU2008-01344/BMC to G.M., BFU2007-62382/BMC to J.M.V. and CSD2006-20642 to G.M. and J.M.V., Comunidad Autónoma de Madrid grants CAM-P2006/Gen-0166 to G.M. and S2009MAT-1507 to J.M.V., and EU 3D-Repertoire grants LSHG-CT-2005-512028 to G.M., J.M.V., M.Z. and C.V.R. and HEALTH-F4-2008-201648 to C.V.R. M.M. acknowledges the financial support of SAF2009-07973 and CSD2007-00017 (MICINN) and S-BIO-0283-2006 (CAM).
Author information
Authors and Affiliations
Contributions
H.Y. and A.B. isolated the complex, I.G.M. and H.Y. obtained the crystals, I.G.M. and G.M. solved the structure, H.Y. and J.M.V. performed the cryo-EM, H.Y. and M.Z. carried out the protease digestion and substrate cleaning experiments, M.Z., A.Y.P. and C.V.R. performed the mass spectrometry and proteomic analysis, P.M., G.d.C., E.B.-N., M.M. and M.S. performed the shRNA assay, all the authors analyzed the data and J.M.V. and G.M. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Notes, Supplementary Figures 1–11, Supplementary Table 1 and Supplementary Methods (PDF 2004 kb)
Supplementary Movie 1
CCT subunits conformational changes inside the ring. This movie summarizes the relative movements that take place inside one of the CCT rings (only the equatorial domain and part of the intermediate domain are shown here). One of the CCT subunits alternates between the different conformations present in the ring (all the other subunits were fitted on that position superimposing their structurally conserved equatorial domains). A dashed circle is depicted as a reference to display the variable sensor loop conformations, which apparently produce a retractile movement towards the centre of the CCT inner cavity (the frame sequence has been ordered to show this “movement”). This movie also depicts the relative movement of the intermediate domain with respect the equatorial domain, which produces the opening and closure of the nucleotide-binding site (here represented with an AMPPNP molecule modelled from the thermosome structure 1Q3Q). (MOV 1073 kb)
Supplementary Movie 2
The molecular forge. A combination of the different conformation of the CCT subunits has been mounted in a random sequence (each subunit varies between eight different resolved conformations; see Supplementary Movie 1) depicting a possible mechanism for CCT mediated folding. In this representation, the piston-like sensor loops modify the shape of a white rubber band that represents a theoretical substrate. If this substrate is bound to several sensor loops at the same time, the substrate could experience local compressions, expansions and torsions that force its folding process. This input of kinetic energy, driven by the ATP binding-hydrolysis cycles, would be used to overcome folding-pathway barriers. Eventually a properly folded product is obtained. (MOV 7014 kb)
Rights and permissions
About this article
Cite this article
Muñoz, I., Yébenes, H., Zhou, M. et al. Crystal structure of the open conformation of the mammalian chaperonin CCT in complex with tubulin. Nat Struct Mol Biol 18, 14–19 (2011). https://doi.org/10.1038/nsmb.1971
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1971
This article is cited by
-
Pathway and mechanism of tubulin folding mediated by TRiC/CCT along its ATPase cycle revealed using cryo-EM
Communications Biology (2023)
-
The role of the molecular chaperone CCT in protein folding and mediation of cytoskeleton-associated processes: implications for cancer cell biology
Cell Stress and Chaperones (2019)
-
TRiC controls transcription resumption after UV damage by regulating Cockayne syndrome protein A
Nature Communications (2018)
-
Development of a yeast internal-subunit eGFP labeling strategy and its application in subunit identification in eukaryotic group II chaperonin TRiC/CCT
Scientific Reports (2018)
-
Chaperonin CCT checkpoint function in basal transcription factor TFIID assembly
Nature Structural & Molecular Biology (2018)