Abstract
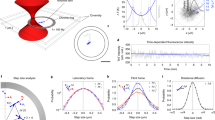
Amyloid fibrils are important in diverse cellular functions, feature in many human diseases and have potential applications in nanotechnology. Here we describe methods that combine optical trapping and fluorescent imaging to characterize the forces that govern the integrity of amyloid fibrils formed by a yeast prion protein. A crucial advance was to use the self-templating properties of amyloidogenic proteins to tether prion fibrils, enabling their manipulation in the optical trap. At normal pulling forces the fibrils were impervious to disruption. At much higher forces (up to 250 pN), discontinuities occurred in force-extension traces before fibril rupture. Experiments with selective amyloid-disrupting agents and mutations demonstrated that such discontinuities were caused by the unfolding of individual subdomains. Thus, our results reveal unusually strong noncovalent intermolecular contacts that maintain fibril integrity even when individual monomers partially unfold and extend fibril length.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Nelson, R. & Eisenberg, D. Structural models of amyloid-like fibrils. Adv. Protein Chem. 73, 235–282 (2006).
Selkoe, D.J. Folding proteins in fatal ways. Nature 426, 900–904 (2003).
Cheng, I.H. et al. Accelerating amyloid-β fibrillization reduces oligomer levels and functional deficits in Alzheimer disease mouse models. J. Biol. Chem. 282, 23818–23828 (2007).
Douglas, P.M. et al. Chaperone-dependent amyloid assembly protects cells from prion toxicity. Proc. Natl. Acad. Sci. USA 105, 7206–7211 (2008).
Shorter, J. & Lindquist, S. Prions as adaptive conduits of memory and inheritance. Nat. Rev. Genet. 6, 435–450 (2005).
Dobson, C.M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Serio, T.R. et al. Nucleated conformational conversion and the replication of conformational information by a prion determinant. Science 289, 1317–1321 (2000).
Glover, J.R. et al. Self-seeded fibers formed by Sup35, the protein determinant of [PSI+], a heritable prion-like factor of S. cerevisiae. Cell 89, 811–819 (1997).
Paushkin, S.V., Kushnirov, V.V., Smirnov, V.N. & TerAvanesyan, M.D. In vitro propagation of the prion-like state of yeast Sup35 protein. Science 277, 381–383 (1997).
Satpute-Krishnan, P., Langseth, S.X. & Serio, T.R. Hsp104-dependent remodeling of prion complexes mediates protein-only inheritance. PLoS Biol. 5, 251–262 (2007).
Kryndushkin, D.S., Alexandrov, I.M., Ter-Avanesyan, M.D. & Kushnirov, V.V. Yeast [PSI+] prion aggregates are formed by small Sup35 polymers fragmented by Hsp104. J. Biol. Chem. 278, 49636–49643 (2003).
Chernoff, Y.O., Lindquist, S.L., Ono, B., Ingevechtomov, S.G. & Liebman, S.W. Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [psi+]. Science 268, 880–884 (1995).
Tanaka, M., Collins, S.R., Toyama, B.H. & Weissman, J.S. The physical basis of how prion conformations determine strain phenotypes. Nature 442, 585–589 (2006).
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J.S. Conformational variations in an infectious protein determine prion strain differences. Nature 428, 323–328 (2004).
Uptain, S.M., Sawicki, G.J., Caughey, B. & Lindquist, S. Strains of [PSI+] are distinguished by their efficiencies of prion-mediated conformational conversion. EMBO J. 20, 6236–6245 (2001).
Woodside, M.T., Garcia-Garcia, C. & Block, S.M. Folding and unfolding single RNA molecules under tension. Curr. Opin. Chem. Biol. 12, 640–646 (2008).
Block, S.M. Kinesin motor mechanics: binding, stepping, tracking, gating, and limping. Biophys. J. 92, 2986–2995 (2007).
Bechtluft, P. et al. Direct observation of chaperone-induced changes in a protein folding pathway. Science 318, 1458–1461 (2007).
Cecconi, C., Shank, E.A., Bustamante, C. & Marqusee, S. Direct observation of the three-state folding of a single protein molecule. Science 309, 2057–2060 (2005).
Kellermayer, M.S.Z., Smith, S.B., Granzier, H.L. & Bustamante, C. Folding-unfolding transitions in single titin molecules characterized with laser tweezers. Science 276, 1112–1116 (1997).
Karsai, A. et al. Mechanical manipulation of Alzheimer's amyloid β1–42 fibrils. J. Struct. Biol. 155, 316–326 (2006).
Kellermayer, M.S.Z. et al. Reversible mechanical unzipping of amyloid β-fibrils. J. Biol. Chem. 280, 8464–8470 (2005).
Raman, E.P., Takeda, T., Barsegov, V. & Klimov, D.K. Mechanical unbinding of Aβ peptides from amyloid fibrils. J. Mol. Biol. 373, 785–800 (2007).
Liu, J.J., Sondheimer, N. & Lindquist, S.L. Changes in the middle region of Sup35 profoundly alter the nature of epigenetic inheritance for the yeast prion [PSI+]. Proc. Natl. Acad. Sci. USA 99 (Suppl. 4), 16446–16453 (2002).
Krishnan, R. & Lindquist, S.L. Structural insights into a yeast prion illuminate nucleation and strain diversity. Nature 435, 765–772 (2005).
Tessier, P.M. & Lindquist, S. Prion recognition elements govern nucleation, strain specificity and species barriers. Nature 447, 556–561 (2007).
Toyama, B.H., Kelly, M.J., Gross, J.D. & Weissman, J.S. The structural basis of yeast prion strain variants. Nature 449, 233–237 (2007).
Scheibel, T. et al. Conducting nanowires built by controlled self-assembly of amyloid fibers and selective metal deposition. Proc. Natl. Acad. Sci. USA 100, 4527–4532 (2003).
Tskhovrebova, L., Trinick, J., Sleep, J.A. & Simmons, R.M. Elasticity and unfolding of single molecules of the giant muscle protein titin. Nature 387, 308–312 (1997).
Collins, S.R., Douglass, A., Vale, R.D. & Weissman, J.S. Mechanism of prion propagation: amyloid growth occurs by monomer addition. PLoS Biol. 2, 1582–1590 (2004).
DePace, A.H. & Weissman, J.S. Origins and kinetic consequences of diversity in Sup35 yeast prion fibers. Nat. Struct. Biol. 9, 389–396 (2002).
Santoso, A., Chien, P., Osherovich, L.Z. & Weissman, J.S. Molecular basis of a yeast prion species barrier. Cell 100, 277–288 (2000).
Dudko, O.K., Hummer, G. & Szabo, A. Theory, analysis, and interpretation of single-molecule force spectroscopy experiments. Proc. Natl. Acad. Sci. USA 105, 15755–15760 (2008).
Lang, M.J., Asbury, C.L., Shaevitz, J.W. & Block, S.M. An automated two-dimensional optical force clamp for single molecule studies. Biophys. J. 83, 491–501 (2002).
Cao, Y. & Li, H. How do chemical denaturants affect the mechanical folding and unfolding of proteins? J. Mol. Biol. 375, 316–324 (2008).
Andersen, C.B. et al. Branching in amyloid fibril growth. Biophys. J. 96, 1529–1536 (2009).
Wang, H. et al. Direct and selective elimination of specific prions and amyloids by 4,5-dianilinophthalimide and analogs. Proc. Natl. Acad. Sci. USA 105, 7159–7164 (2008).
Tessier, P.M. & Lindquist, S. Unraveling infectious structures, strain variants and species barriers for the yeast prion [PSI+]. Nat. Struct. Mol. Biol. 16, 598–605 (2009).
Liu, J.J. & Lindquist, S. Oligopeptide-repeat expansions modulate 'protein-only' inheritance in yeast. Nature 400, 573–576 (1999).
Greene, D.N. et al. Single-molecule force spectroscopy reveals a stepwise unfolding of Caenorhabditis elegans giant protein kinase domains. Biophys. J. 95, 1360–1370 (2008).
Oberhauser, A.F., Hansma, P.K., Carrion-Vazquez, M. & Fernandez, J.M. Stepwise unfolding of titin under force-clamp atomic force microscopy. Proc. Natl. Acad. Sci. USA 98, 468–472 (2001).
Cao, Y. & Li, H.B. Polyprotein of GB1 is an ideal artificial elastomeric protein. Nat. Mater. 6, 109–114 (2007).
Bell, G.I. Models for specific adhesion of cells to cells. Science 200, 618–627 (1978).
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl. Acad. Sci. USA 96, 3694–3699 (1999).
Gebhardt, J.C.M., Bornschlogla, T. & Rief, M. Full distance-resolved folding energy landscape of one single protein molecule. Proc. Natl. Acad. Sci. USA 107, 2013–2018 (2010).
Kellermayer, M.S.Z., Smith, S., Bustamante, C. & Granzier, H.L. Mechanical manipulation of single titin molecules with laser tweezers. Adv. Exp. Med. Biol. 481, 111–126 (2000).
Zhang, J. & Muthukumar, M. Simulations of nucleation and elongation of amyloid fibrils. J. Chem. Phys. 130, 035102 (2009).
Pallitto, M.M. & Murphy, R.M. A mathematical model of the kinetics of β-amyloid fibril growth from the denatured state. Biophys. J. 81, 1805–1822 (2001).
Lee, C.C., Nayak, A., Sethuraman, A., Belfort, G. & Mcrae, G.J. A three-stage kinetic model of amyloid fibrillation. Biophys. J. 92, 3448–3458 (2007).
Kishimoto, A. et al. β-helix is a likely core structure of yeast prion Sup35 amyloid fibers. Biochem. Biophys. Res. Commun. 315, 739–745 (2004)[anna].
Lazo, N.D. & Downing, D.T. Amyloid fibrils may be assembled from β-helical protofibrils. Biochemistry 37, 1731–1735 (1998).
Shewmaker, F., Wickner, R.B. & Tycko, R. Amyloid of the prion domain of Sup35p has an in-register parallel β-sheet structure. Proc. Natl. Acad. Sci. USA 103, 19754–19759 (2006).
Keten, S. & Buehler, M.J. Large deformation and fracture mechanics of a beta-helical protein nanotube: atomistic and continuum modeling. Comput. Methods Appl. Mech. Eng. 197, 3203–3214 (2008).
Brockwell, D.J. et al. Pulling geometry defines the mechanical resistance of a β-sheet protein. Nat. Struct. Biol. 10, 731–737 (2003).
Bustamante, C., Chemla, Y.R., Forde, N.R. & Izhaky, D. Mechanical processes in biochemistry. Annu. Rev. Biochem. 73, 705–748 (2004).
Yang, S.C., Levine, H., Onuchic, J.N. & Cox, D.L. Structure of infectious prions: stabilization by domain swapping. FASEB J. 19, 1778–1782 (2005).
White, H.E. et al. Globular tetramers of β2-microglobulin assemble into elaborate amyloid fibrils. J. Mol. Biol. 389, 48–57 (2009).
Sondheimer, N. & Lindquist, S. Rnq1: an epigenetic modifier of protein function in yeast. Mol. Cell 5, 163–172 (2000).
Neuman, K.C. & Block, S.M. Optical trapping. Rev. Sci. Instrum. 75, 2787–2809 (2004).
Palmer, J.S. & Boyce, M.C. Constitutive modeling of the stress-strain behavior of F-actin filament networks. Acta Biomater. 4, 597–612 (2008).
Acknowledgements
We thank members of the Lindquist laboratory for comments on the manuscript and S. Block, W. Hwang and K. Allendoerfer for their critical reading. S.L. is an investigator of the Howard Hughes Medical Institute. This work was supported by US National Institutes of Health grant GM025874 to S.L., a US National Science Foundation Career Award (0643745) to M.J.L. and an American Heart Association fellowship to J.D. (0725849T). The project described was also supported by a US National Institute of Biomedical Imaging and Bioengineering grant (T32EB006348). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of Biomedical Imaging and Bioengineering or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.D. initiated the project and carried out sample preparation and yeast biology; J.D. and C.E.C. designed and carried out the optical-trapping experiments and data analysis; M.C.B., M.J.L. and S.L. supervised the projects and interpreted the results; J.D. and S.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Methods (PDF 603 kb)
Supplementary Movie 1
Movie S1. The NM fibrils were tethered to the glass surface. The movie was taken from a standard inverted fluorescence microscope with a 100× objective lens (field of observation 60 μm × 60 μm). The main bodies of the tethered fibrils were often out of focus as seen at the beginning of the movie. In the middle of the experiment, flow was introduced to straighten the fibrils for better visualization. The direction of the flow was changed during the experiments to demonstrate that these fibrils were indeed attached to the surface at one end while the main bodies were mobile. (MOV 593 kb)
Supplementary Movie 2
Movie S2. Fluorescence image of tethered NM fibrils on the optical trapping instrument (field of observation 15 μm × 15 μm). The distribution of tethered fibrils in the movie was illustrated below. Fibrils with one end attached to the glass surface are highlighted by green arrows, while fibrils with both ends attached to the glass surface are highlighted by red arrows. We optimized the procedure for each experiment to reduce the number of fibrils double-tethered to the glass surface. (MOV 12803 kb)
Supplementary Movie 3
Movie S3. The NM prion domain from Candida albicans Sup35 homolog did not capture S. cerevisiae NM fibrils. When CaNM monomers were pre-deposited to the glass surface followed by casein blocking, ScNM fibrils were rarely recruited and tethered to the surface. Typically no more than three ScNM fibrils were observed in the field of observation (60 μm × 60 μm) and the movie showed the extreme. (MOV 9471 kb)
Supplementary Movie 4
Movie S4 – 6. Cleavage of NM monomers at the junction between the fibril and the glass surface released tethered NM fibrils. S4. Before the TEV treatment to release tethered NM fibrils. The fluorescent channel was opened transiently for imaging to minimize fluorescence bleaching. (MOV 1054 kb)
Supplementary Movie 5
Movie S4 – 6. Cleavage of NM monomers at the junction between the fibril and the glass surface released tethered NM fibrils. S5. After the TEV treatment to release tethered NM fibrils. 20 μl of 20 unit μl−1 TEV protease was introduced into the chamber by flow. This fluorescent image was recorded after 10-minute incubation of TEV protease. The bead that was previously attached to one end of the fibril and was trapped with the optical laser is visible in the image. However, the previously tethered fibril shown in movie S4 was no longer detected after the TEV protease treatment. We suspected that, after release from the glass surface, the flexible fibril might collapse and be hidden from view by the bead. (MOV 1437 kb)
Supplementary Movie 6
Movie S4 – 6. Cleavage of NM monomers at the junction between the fibril and the glass surface released tethered NM fibrils. S6. After the TEV treatment to release tethered NM fibrils, imaged with a constant flow. To test if the released fibril was hidden from view by the bead, we asked if we could extend the hidden fibril with a constant flow of 1xCRBB buffer through the flow channel. The flow is readily detectable in this movie by the free objects in the flow channel flowing from the left lower corner to the upper right corner in the movie. Indeed, in the presence of a constant flow, the fibril that had been released from the glass surface by the TEV cleavage was now visible, aligning itself with the flow. (MOV 1438 kb)
Supplementary Movie 7
Movie S7 – 8. Typical pulling behavior of tethered NM fibrils. Prior to each pulling experiment, the tethered fibril and the attached bead were fluorescently imaged to confirm the nature of the tethers. The instrument we employed to achieve low trapping forces allowed an interlaced optical force and fluorescence (IOFF). In short, fluorescence and excitation lasers were cycled out of phase at 50 kHz with a 10% duty cycle lag time in between to allow excited electrons to return to their ground state. This method prolonged the fluorescence signal from the bead and the fibril and enabled imaging of the fibril morphology throughout the force-extension experiment. S7. Fluorescent image of the tethered NM fibril when the trapping laser was not turned on. (MOV 1865 kb)
Supplementary Movie 8
Movie S7 – 8. Typical pulling behavior of tethered NM fibrils. Prior to each pulling experiment, the tethered fibril and the attached bead were fluorescently imaged to confirm the nature of the tethers. The instrument we employed to achieve low trapping forces allowed an interlaced optical force and fluorescence (IOFF). In short, fluorescence and excitation lasers were cycled out of phase at 50 kHz with a 10% duty cycle lag time in between to allow excited electrons to return to their ground state. This method prolonged the fluorescence signal from the bead and the fibril and enabled imaging of the fibril morphology throughout the force-extension experiment. S8. The trapping laser was turned on to perform the pulling experiments. (MOV 7783 kb)
Supplementary Movie 9
Movie S9– 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S9 – 10. Rupture occurred in the middle of this fibril. S9, before rupture; S10, after rupture. The portion of the fibril left on the glass surface moved freely relative to the trapped bead after rupture. The trapped bead was able to move freely with pulling force, confirming that the connection to the fibril had been ruptured. (MOV 2490 kb)
Supplementary Movie 10
Movie S9 – 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S9 – 10.Rupture occurred in the middle of this fibril. S9, before rupture; S10, after rupture. The portion of the fibril left on the glass surface moved freely relative to the trapped bead after rupture. The trapped bead was able to move freely with pulling force, confirming that the connection to the fibril had been ruptured. (MOV 27001 kb)
Supplementary Movie 11
Movie S9 – 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S11 – 12.Rupture occurred close to the bead. S11, before rupture; S12, after rupture. (MOV 2869 kb)
Supplementary Movie 12
Movie S9 – 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S11 – 12.Rupture occurred close to the bead. S11, before rupture; S12, after rupture. (MOV 13611 kb)
Supplementary Movie 13
Movie S9 – 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S13 – 14.Rupture occurred close to the glass surface. S13, before rupture; S14, after rupture. The small fragment remaining on the surface couldn't be identified but the fibril, still attached to the bead, could be visualized. To ensure that rupture of the tether had occurred we then flowed buffer through the chamber. This reoriented the fibril – now tethered only to the bead – relative to its original orientation. (MOV 3461 kb)
Supplementary Movie 14
Movie S9 – 14. Direct observation of the rupture of tethered NM fibrils by fluorescent imaging. We trapped the bead at the extended position and manually aligned the fibril with the x-axis of the piezo stage without centering. Without the prolonged process of centering, trap-accelerated photo bleaching was minimized and fluorescent images could be recorded stably before rupture. Note however that the fragment of the fibril closest to the bead still became photobleached during this process. Constant high force was then applied to the bead in the absence of fluorescent imaging until the tether was ruptured. To establish the position of rupture, fluorescent imaging was re-initiated. In movies recorded after rupture, fragments of the previously tethered fibrils could be seen still attached to the surface, and these represented varying fractions of the initial fibril length. Other objects in the field of view provide a point of reference for the ruptured fibril. S13 – 14. Rupture occurred close to the glass surface. S13, before rupture; S14, after rupture. The small fragment remaining on the surface couldn't be identified but the fibril, still attached to the bead, could be visualized. To ensure that rupture of the tether had occurred we then flowed buffer through the chamber. This reoriented the fibril – now tethered only to the bead – relative to its original orientation. (MOV 22718 kb)
Rights and permissions
About this article
Cite this article
Dong, J., Castro, C., Boyce, M. et al. Optical trapping with high forces reveals unexpected behaviors of prion fibrils. Nat Struct Mol Biol 17, 1422–1430 (2010). https://doi.org/10.1038/nsmb.1954
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1954
This article is cited by
-
Molecular dynamics simulation-based understanding of the structure and property of amyloid proteins at multiple length scales
JMST Advances (2023)
-
Myosin II Adjusts Motility Properties and Regulates Force Production Based on Motor Environment
Cellular and Molecular Bioengineering (2022)
-
Extracellular protein isolation from the matrix of anammox biofilm using ionic liquid extraction
Applied Microbiology and Biotechnology (2020)
-
Mechanical Deformation Mechanisms and Properties of Prion Fibrils Probed by Atomistic Simulations
Nanoscale Research Letters (2017)
-
Trapping and manipulation of nanoparticles using multifocal optical vortex metalens
Scientific Reports (2017)