Abstract
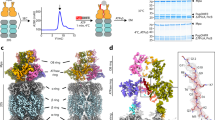
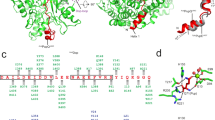
Mycobacterium tuberculosis uses a proteasome system that is analogous to the eukaryotic ubiquitin-proteasome pathway and is required for pathogenesis. However, the bacterial analog of ubiquitin, prokaryotic ubiquitin-like protein (Pup), is an intrinsically disordered protein that bears little sequence or structural resemblance to the highly structured ubiquitin. Thus, it was unknown how pupylated proteins were recruited to the proteasome. Here, we show that the Mycobacterium proteasomal ATPase (Mpa) has three pairs of tentacle-like coiled coils that recognize Pup. Mpa bound unstructured Pup through hydrophobic interactions and a network of hydrogen bonds, leading to the formation of an α-helix in Pup. Our work describes a binding-induced folding recognition mechanism in the Pup-proteasome system that differs mechanistically from substrate recognition in the ubiquitin-proteasome system. This key difference between the prokaryotic and eukaryotic systems could be exploited for the development of a small molecule-based treatment for tuberculosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 78, 477–513 (2009).
Lupas, A. et al. Eubacterial proteasomes. Mol. Biol. Rep. 24, 125–131 (1997).
Lin, G. et al. Mycobacterium tuberculosis prcBA genes encode a gated proteasome with broad oligopeptide specificity. Mol. Microbiol. 59, 1405–1416 (2006).
Hu, G. et al. Structure of the Mycobacterium tuberculosis proteasome and mechanism of inhibition by a peptidyl boronate. Mol. Microbiol. 59, 1417–1428 (2006).
Wang, T. et al. Structural insights on the Mycobacterium tuberculosis proteasomal ATPase Mpa. Structure 17, 1377–1385 (2009).
Gandotra, S., Schnappinger, D., Monteleone, M., Hillen, W. & Ehrt, S. In vivo gene silencing identifies the Mycobacterium tuberculosis proteasome as essential for the bacteria to persist in mice. Nat. Med. 13, 1515–1520 (2007).
Voges, D., Zwickl, P. & Baumeister, W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (1999).
Nathan, C. et al. A philosophy of anti-infectives as a guide in the search for new drugs for tuberculosis. Tuberculosis (Edinb.) 88 Suppl 1, S25–S33 (2008).
Darwin, K.H., Lin, G., Chen, Z., Li, H. & Nathan, C.F. Characterization of a Mycobacterium tuberculosis proteasomal ATPase homologue. Mol. Microbiol. 55, 561–571 (2005).
Zhang, F. et al. Structural insights into the regulatory particle of the proteasome from Methanocaldococcus jannaschii. Mol. Cell 34, 473–484 (2009).
Djuranovic, S. et al. Structure and activity of the N-terminal substrate recognition domains in proteasomal ATPases. Mol. Cell 34, 580–590 (2009).
Pearce, M.J., Mintseris, J., Ferreyra, J., Gygi, S.P. & Darwin, K.H. Ubiquitin-like protein involved in the proteasome pathway of Mycobacterium tuberculosis. Science 322, 1104–1107 (2008).
Burns, K.E., Liu, W.T., Boshoff, H.I., Dorrestein, P.C. & Barry, C.E. III. Proteasomal protein degradation in Mycobacteria is dependent upon a prokaryotic ubiquitin-like protein. J. Biol. Chem. 284, 3069–3075 (2009).
Sutter, M., Damberger, F.F., Imkamp, F., Allain, F.H. & Weber-Ban, E. Prokaryotic ubiquitin-like protein (Pup) is coupled to substrates via the side chain of its C-terminal glutamate. J. Am. Chem. Soc. 132, 5610–5612 (2010).
Striebel, F., Hunkeler, M., Summer, H. & Weber-Ban, E. The mycobacterial Mpa-proteasome unfolds and degrades pupylated substrates by engaging Pup's N-terminus. EMBO J. 29, 1262–1271 (2010).
Burns, K.E., Pearce, M.J. & Darwin, K.H. Prokaryotic ubiquitin-like protein provides a two-part degron to Mycobacterium proteasome substrates. J. Bacteriol. 192, 2933–2935 (2010).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Zhou, F.X., Merianos, H.J., Brunger, A.T. & Engelman, D.M. Polar residues drive association of polyleucine transmembrane helices. Proc. Natl. Acad. Sci. USA 98, 2250–2255 (2001).
Meindl-Beinker, N.M., Lundin, C., Nilsson, I., White, S.H. & von Heijne, G. Asn- and Asp-mediated interactions between transmembrane helices during translocon-mediated membrane protein assembly. EMBO Rep. 7, 1111–1116 (2006).
Sutter, M., Striebel, F., Damberger, F.F., Allain, F.H. & Weber-Ban, E. A distinct structural region of the prokaryotic ubiquitin-like protein (Pup) is recognized by the N-terminal domain of the proteasomal ATPase Mpa. FEBS Lett. 583, 3151–3157 (2009).
Chen, X. et al. Prokaryotic ubiquitin-like protein pup is intrinsically disordered. J. Mol. Biol. 392, 208–217 (2009).
Martin, A., Baker, T.A. & Sauer, R.T. Pore loops of the AAA+ ClpX machine grip substrates to drive translocation and unfolding. Nat. Struct. Mol. Biol. 15, 1147–1151 (2008).
Burns, K.E. et al. “Depupylation” of prokaryotic ubiquitin-like protein from mycobacterial proteasome substrates. Mol. Cell 39, 821–827 (2010).
Darwin, K.H. & Hofmann, K. SAMPyling proteins in archaea. Trends Biochem. Sci. 35, 348–351 (2010).
Humbard, M.A. et al. Ubiquitin-like small archaeal modifier proteins (SAMPs) in Haloferax volcanii. Nature 463, 54–60 (2010).
Dyson, H.J. & Wright, P.E. Coupling of folding and binding for unstructured proteins. Curr. Opin. Struct. Biol. 12, 54–60 (2002).
Shoemaker, B.A., Portman, J.J. & Wolynes, P.G. Speeding molecular recognition by using the folding funnel: the fly-casting mechanism. Proc. Natl. Acad. Sci. USA 97, 8868–8873 (2000).
Horton, R.M., Hunt, H.D., Ho, S.N., Pullen, J.K. & Pease, L.R. Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene 77, 61–68 (1989).
Hatfull, G.F. & Jacobs, J.W.R. Molecular Genetics of Mycobacteria (ASM, Washington, DC, 2000).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Potterton, E., McNicholas, S., Krissinel, E., Cowtan, K. & Noble, M. The CCP4 molecular-graphics project. Acta Crystallogr. D Biol. Crystallogr. 58, 1955–1957 (2002).
Acknowledgements
We thank K. Burns for reviewing this manuscript and C. Nathan for advice and encouragement. X-ray diffraction data for this study were collected at beamlines X25 and X29 of the National Synchrotron Light Source. Financial support was principally from the Offices of Biological and Environmental Research and of Basic Energy Sciences of the US Department of Energy, and from the National Center for Research Resources of the US National Institutes of Health (NIH). This work was supported by NIH grant AI070285 and Brookhaven National Laboratory LDRD grant 10-016 to H.L. and by NIH grant HL092774 to K.H.D. K.H.D. was also supported by a Burroughs Wellcome Investigator in the Pathogenesis of Infectious Diseases award.
Author information
Authors and Affiliations
Contributions
T.W. performed protein purification, crystallization and structure determination; K.H.D. performed mutagenesis and in vivo degradation assays; T.W., K.H.D. and H.L. designed experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1 and 2 (PDF 2054 kb)
Rights and permissions
About this article
Cite this article
Wang, T., Darwin, K. & Li, H. Binding-induced folding of prokaryotic ubiquitin-like protein on the Mycobacterium proteasomal ATPase targets substrates for degradation. Nat Struct Mol Biol 17, 1352–1357 (2010). https://doi.org/10.1038/nsmb.1918
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1918
This article is cited by
-
Active metabolism unmasks functional protein–protein interactions in real time in-cell NMR
Communications Biology (2020)
-
Prokaryotic ubiquitin-like protein remains intrinsically disordered when covalently attached to proteasomal target proteins
BMC Structural Biology (2018)
-
Genetic heterogeneity among slow acetylator N-acetyltransferase 2 phenotypes in cryopreserved human hepatocytes
Archives of Toxicology (2017)
-
Catalytic properties and heat stabilities of novel recombinant human N-acetyltransferase 2 allozymes support existence of genetic heterogeneity within the slow acetylator phenotype
Archives of Toxicology (2017)
-
Bacteria-host relationship: ubiquitin ligases as weapons of invasion
Cell Research (2016)