Abstract
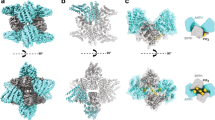
Multisubunit tethering complexes of the CATCHR (complexes associated with tethering containing helical rods) family are proposed to mediate the initial contact between an intracellular trafficking vesicle and its membrane target. To begin elucidating the molecular architecture of one well-studied example, the conserved oligomeric Golgi (COG) complex, we reconstituted its essential subunits (Cog1, Cog2, Cog3 and Cog4) and used single-particle electron microscopy to reveal a y-shaped structure with three flexible, highly extended legs. Labeling experiments established that the N termini of all four subunits interact along the proximal segment of one leg, whereas three of the four C termini are located at the tips of the legs. Our results suggest that the central region of the Cog1–Cog2–Cog3–Cog4 complex, as well as the distal regions of at least two legs, all participate in interactions with other components of the intracellular trafficking machinery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Whyte, J.R. & Munro, S. Vesicle tethering complexes in membrane traffic. J. Cell Sci. 115, 2627–2637 (2002).
Cai, H., Reinisch, K. & Ferro-Novick, S. Coats, tethers, Rabs, and SNAREs work together to mediate the intracellular destination of a transport vesicle. Dev. Cell 12, 671–682 (2007).
Sztul, E. & Lupashin, V. Role of vesicle tethering factors in the ER-Golgi membrane traffic. FEBS Lett. 583, 3770–3783 (2009).
Yu, I. & Hughson, F.M. Tethering factors as organizers of intracellular vesicular traffic. Annu. Rev. Cell Dev. Biol. 26, 137–156 (2010).
Ungar, D., Oka, T., Krieger, M. & Hughson, F.M. Retrograde transport on the COG railway. Trends Cell Biol. 16, 113–120 (2006).
Ungar, D. et al. Characterization of a mammalian Golgi-localized protein complex, COG, that is required for normal Golgi morphology and function. J. Cell Biol. 157, 405–415 (2002).
Walter, D.M., Paul, K.S. & Waters, M.G. Purification and characterization of a novel 13 S hetero–oligomeric protein complex that stimulates in vitro Golgi transport. J. Biol. Chem. 273, 29565–29576 (1998).
Whyte, J.R. & Munro, S. The Sec34/35 Golgi transport complex is related to the exocyst, defining a family of complexes involved in multiple steps of membrane traffic. Dev. Cell 1, 527–537 (2001).
VanRheenen, S.M., Cao, X., Lupashin, V.V., Barlowe, C. & Waters, M.G. Sec35p, a novel peripheral membrane protein, is required for ER to Golgi vesicle docking. J. Cell Biol. 141, 1107–1119 (1998).
VanRheenen, S.M. et al. Sec34p, a protein required for vesicle tethering to the yeast Golgi apparatus, is in a complex with Sec35p. J. Cell Biol. 147, 729–742 (1999).
Wuestehube, L.J. et al. New mutants of Saccharomyces cerevisiae affected in the transport of proteins from the endoplasmic reticulum to the Golgi complex. Genetics 142, 393–406 (1996).
Ram, R.J., Li, B. & Kaiser, C.A. Identification of Sec36p, Sec37p, and Sec38p: components of yeast complex that contains Sec34p and Sec35p. Mol. Biol. Cell 13, 1484–1500 (2002).
Suvorova, E.S., Duden, R. & Lupashin, V.V. The Sec34/Sec35p complex, a Ypt1p effector required for retrograde intra-Golgi trafficking, interacts with Golgi SNAREs and COPI vesicle coat proteins. J. Cell Biol. 157, 631–643 (2002).
Luo, Z. & Gallwitz, D. Biochemical and genetic evidence for the involvement of yeast Ypt6-GTPase in protein retrieval to different Golgi compartments. J. Biol. Chem. 278, 791–799 (2003).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Shestakova, A., Suvorova, E., Pavliv, O., Khaidakova, G. & Lupashin, V. Interaction of the conserved oligomeric Golgi complex with t-SNARE Syntaxin5a/Sed5 enhances intra-Golgi SNARE complex stability. J. Cell Biol. 179, 1179–1192 (2007).
Zolov, S.N. & Lupashin, V.V. Cog3p depletion blocks vesicle-mediated Golgi retrograde trafficking in HeLa cells. J. Cell Biol. 168, 747–759 (2005).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Deutschbauer, A.M. et al. Mechanisms of haploinsufficiency revealed by genome-wide profiling in yeast. Genetics 169, 1915–1925 (2005).
Fotso, P., Koryakina, Y., Pavliv, O., Tsiomenko, A.B. & Lupashin, V.V. Cog1p plays a central role in the organization of the yeast conserved oligomeric Golgi complex. J. Biol. Chem. 280, 27613–27623 (2005).
Ungar, D., Oka, T., Vasile, E., Krieger, M. & Hughson, F.M. Subunit architecture of the conserved oligomeric Golgi complex. J. Biol. Chem. 280, 32729–32735 (2005).
Ren, Y. et al. A structure-based mechanism for vesicle capture by the multisubunit tethering complex Dsl1. Cell 139, 1119–1129 (2009).
Tripathi, A., Ren, Y., Jeffrey, P.D. & Hughson, F.M. Structural characterization of Tip20p and Dsl1p, subunits of the Dsl1p vesicle tethering complex. Nat. Struct. Mol. Biol. 16, 114–123 (2009).
Cavanaugh, L.F. et al. Structural analysis of conserved oligomeric Golgi complex subunit 2. J. Biol. Chem. 282, 23418–23426 (2007).
Richardson, B.C. et al. Structural basis for a human glycosylation disorder caused by mutation of the COG4 gene. Proc. Natl. Acad. Sci. USA 106, 13329–13334 (2009).
Dong, G., Hutagalung, A.H., Fu, C., Novick, P. & Reinisch, K.M. The structures of exocyst subunit Exo70p and the Exo84p C-terminal domains reveal a common motif. Nat. Struct. Mol. Biol. 12, 1094–1100 (2005).
Sivaram, M.V., Furgason, M.L., Brewer, D.N. & Munson, M. The structure of the exocyst subunit Sec6p defines a conserved architecture with diverse roles. Nat. Struct. Mol. Biol. 13, 555–556 (2006).
Wu, S., Mehta, S.Q., Pichaud, F., Bellen, H.J. & Quiocho, F.A. Sec15 interacts with Rab11 via a novel domain and affects Rab11 localization in vivo. Nat. Struct. Mol. Biol. 12, 879–885 (2005).
Scheich, C., Kummel, D., Soumailakakis, D., Heinemann, U. & Bussow, K. Vectors for co-expression of an unrestricted number of proteins. Nucleic Acids Res. 35, e43 (2007).
Wu, X. et al. Mutation of the COG complex subunit gene COG7 causes a lethal congenital disorder. Nat. Med. 10, 518–523 (2004).
Foulquier, F. et al. A new inborn error of glycosylation due to a Cog8 deficiency reveals a critical role for the Cog1-Cog8 interaction in COG complex formation. Hum. Mol. Genet. 16, 717–730 (2007).
Kranz, C. et al. COG8 deficiency causes new congenital disorder of glycosylation type IIh. Hum. Mol. Genet. 16, 731–741 (2007).
Foulquier, F. et al. Conserved oligomeric Golgi complex subunit 1 deficiency reveals a previously uncharacterized congenital disorder of glycosylation type II. Proc. Natl. Acad. Sci. USA 103, 3764–3769 (2006).
Hsu, S.C. et al. Subunit composition, protein interactions, and structures of the mammalian brain sec6/8 complex and septin filaments. Neuron 20, 1111–1122 (1998).
Sutton, R.B., Fasshauer, D., Jahn, R. & Brunger, A.T. Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 Å resolution. Nature 395, 347–353 (1998).
Reynders, E. et al. Golgi function and dysfunction in the first COG4-deficient CDG type II patient. Hum. Mol. Genet. 18, 3244–3256 (2009).
Laufman, O., Kedan, A., Hong, W. & Lev, S. Direct interaction between the COG complex and the SM protein, Sly1, is required for Golgi SNARE pairing. EMBO J. 28, 2006–2017 (2009).
Semenza, J.C., Hardwick, K.G., Dean, N. & Pelham, H.R. ERD2, a yeast gene required for the receptor-mediated retrieval of luminal ER proteins from the secretory pathway. Cell 61, 1349–1357 (1990).
Voth, W.P., Jiang, Y.W. & Stillman, D.J. New 'marker swap' plasmids for converting selectable markers on budding yeast gene disruptions and plasmids. Yeast 20, 985–993 (2003).
Longtine, M.S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Rose, M.D., Misra, L.M. & Vogel, J.P. KAR2, a karyogamy gene, is the yeast homolog of the mammalian BiP/GRP78 gene. Cell 57, 1211–1221 (1989).
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification—powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004).
Li, Z., Hite, R.K., Cheng, Y. & Walz, T. Evaluation of imaging plates as recording medium for images of negatively stained single particles and electron diffraction patterns of two-dimensional crystals. J. Electron Microsc. (Tokyo) 59, 53–63 (2010).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Acknowledgements
We gratefully acknowledge M. Rose and C. Ydenberg (Princeton University) for providing strains, reagents, technical assistance and valuable advice, as well as S. Silverman and L. Schepis (Princeton University) for providing reagents and technical assistance. We also thank V. Lupashin, B. Richardson, D. Ungar and M. Paul for useful discussions. This work was supported by a grant to F.M.H. from the US National Institutes of Health (GM071574). C.K.Y. acknowledges fellowships from the Jane Coffin-Childs Memorial Fund and the Canadian Institutes for Health Research. T.W. is an investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.A.L. conducted yeast experiments and prepared samples for EM. C.K.Y. conducted EM experiments and analyzed the data. All authors discussed the results and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 10209 kb)
Rights and permissions
About this article
Cite this article
Lees, J., Yip, C., Walz, T. et al. Molecular organization of the COG vesicle tethering complex. Nat Struct Mol Biol 17, 1292–1297 (2010). https://doi.org/10.1038/nsmb.1917
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1917
This article is cited by
-
Systematic analysis reveals the prevalence and principles of bypassable gene essentiality
Nature Communications (2019)
-
Chaperoning SNARE assembly and disassembly
Nature Reviews Molecular Cell Biology (2016)
-
Molecular architecture of the complete COG tethering complex
Nature Structural & Molecular Biology (2016)
-
CATCHR, HOPS and CORVET tethering complexes share a similar architecture
Nature Structural & Molecular Biology (2016)
-
Subunit connectivity, assembly determinants and architecture of the yeast exocyst complex
Nature Structural & Molecular Biology (2016)