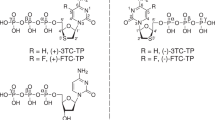
Abstract
Human immunodeficiency virus (HIV-1) develops resistance to 3′-azido-2′,3′-deoxythymidine (AZT, zidovudine) by acquiring mutations in reverse transcriptase that enhance the ATP-mediated excision of AZT monophosphate from the 3′ end of the primer. The excision reaction occurs at the dNTP-binding site, uses ATP as a pyrophosphate donor, unblocks the primer terminus and allows reverse transcriptase to continue viral DNA synthesis. The excision product is AZT adenosine dinucleoside tetraphosphate (AZTppppA). We determined five crystal structures: wild-type reverse transcriptase–double-stranded DNA (RT–dsDNA)–AZTppppA; AZT-resistant (AZTr; M41L D67N K70R T215Y K219Q) RT–dsDNA–AZTppppA; AZTr RT–dsDNA terminated with AZT at dNTP- and primer-binding sites; and AZTr apo reverse transcriptase. The AMP part of AZTppppA bound differently to wild-type and AZTr reverse transcriptases, whereas the AZT triphosphate part bound the two enzymes similarly. Thus, the resistance mutations create a high-affinity ATP-binding site. The structure of the site provides an opportunity to design inhibitors of AZT-monophosphate excision.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Fauci, A.S. 25 years of HIV/AIDS science: reaching the poor with research advances. Cell 131, 429–432 (2007).
Kohlstaedt, L.A., Wang, J., Friedman, J.M., Rice, P.A. & Steitz, T.A. Crystal structure at 3.5 Å resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 256, 1783–1790 (1992).
Jacobo-Molina, A. et al. Crystal structure of human immunodeficiency virus type 1 reverse transcriptase complexed with double-stranded DNA at 3.0 Å resolution shows bent DNA. Proc. Natl. Acad. Sci. USA 90, 6320–6324 (1993).
Huang, H., Chopra, R., Verdine, G.L. & Harrison, S.C. Structure of a covalently trapped catalytic complex of HIV-1 reverse transcriptase: implications for drug resistance. Science 282, 1669–1675 (1998).
Sarafianos, S.G. et al. Lamivudine (3TC) resistance in HIV-1 reverse transcriptase involves steric hindrance with β-branched amino acids. Proc. Natl. Acad. Sci. USA 96, 10027–10032 (1999).
Das, K. et al. Structural basis for the role of the K65R mutation in HIV-1 reverse transcriptase polymerization, excision antagonism, and tenofovir resistance. J. Biol. Chem. 284, 35092–35100 (2009).
Larder, B.A. & Kemp, S.D. Multiple mutations in HIV-1 reverse transcriptase confer high-level resistance to zidovudine (AZT). Science 246, 1155–1158 (1989).
Arion, D., Kaushik, N., McCormick, S., Borkow, G. & Parniak, M.A. Phenotypic mechanism of HIV-1 resistance to 3′-azido-3′-deoxythymidine (AZT): increased polymerization processivity and enhanced sensitivity to pyrophosphate of the mutant viral reverse transcriptase. Biochemistry 37, 15908–15917 (1998).
Meyer, P.R., Matsuura, S.E., So, A.G. & Scott, W.A. Unblocking of chain-terminated primer by HIV-1 reverse transcriptase through a nucleotide-dependent mechanism. Proc. Natl. Acad. Sci. USA 95, 13471–13476 (1998).
Meyer, P.R., Matsuura, S.E., Mian, A.M., So, A.G. & Scott, W.A. A mechanism of AZT resistance: an increase in nucleotide-dependent primer unblocking by mutant HIV-1 reverse transcriptase. Mol. Cell 4, 35–43 (1999).
Boyer, P.L., Sarafianos, S.G., Arnold, E. & Hughes, S.H. Selective excision of AZTMP by drug-resistant human immunodeficiency virus reverse transcriptase. J. Virol. 75, 4832–4842 (2001).
Jeeninga, R.E., Keulen, W., Boucher, C., Sanders, R.W. & Berkhout, B. Evolution of AZT resistance in HIV-1: the 41–70 intermediate that is not observed in vivo has a replication defect. Virology 283, 294–305 (2001).
Mas, A. et al. Role of a dipeptide insertion between codons 69 and 70 of HIV-1 reverse transcriptase in the mechanism of AZT resistance. EMBO J. 19, 5752–5761 (2000).
Boyer, P.L., Sarafianos, S.G., Arnold, E. & Hughes, S.H. Nucleoside analog resistance caused by insertions in the fingers of human immunodeficiency virus type 1 reverse transcriptase involves ATP-mediated excision. J. Virol. 76, 9143–9151 (2002).
Meyer, P.R., Lennerstrand, J., Matsuura, S.E., Larder, B.A. & Scott, W.A. Effects of dipeptide insertions between codons 69 and 70 of human immunodeficiency virus type 1 reverse transcriptase on primer unblocking, deoxynucleoside triphosphate inhibition, and DNA chain elongation. J. Virol. 77, 3871–3877 (2003).
Meyer, P.R. et al. Relationship between 3′-azido-3′-deoxythymidine resistance and primer unblocking activity in foscarnet-resistant mutants of human immunodeficiency virus type 1 reverse transcriptase. J. Virol. 77, 6127–6137 (2003).
Sluis-Cremer, N. et al. Molecular mechanism by which the K70E mutation in human immunodeficiency virus type 1 reverse transcriptase confers resistance to nucleoside reverse transcriptase inhibitors. Antimicrob. Agents Chemother. 51, 48–53 (2007).
Dharmasena, S., Pongracz, Z., Arnold, E., Sarafianos, S.G. & Parniak, M.A. 3′-Azido-3′-deoxythymidine-(5′)-tetraphospho-(5′)-adenosine, the product of ATP-mediated excision of chain-terminating AZTMP, is a potent chain-terminating substrate for HIV-1 reverse transcriptase. Biochemistry 46, 828–836 (2007).
Han, Q., Gaffney, B.L. & Jones, R.A. One-flask synthesis of dinucleoside tetra- and pentaphosphates. Org. Lett. 8, 2075–2077 (2006).
Sarafianos, S.G. et al. Structures of HIV-1 reverse transcriptase with pre- and post-translocation AZTMP-terminated DNA. EMBO J. 21, 6614–6624 (2002).
Carfí, A., Smith, C.A., Smolak, P.J., McGrew, J. & Wiley, D.C. Structure of a soluble secreted chemokine inhibitor vCCI (p35) from cowpox virus. Proc. Natl. Acad. Sci. USA 96, 12379–12383 (1999).
Kleywegt, G.J. Use of non-crystallographic symmetry in protein structure refinement. Acta Crystallogr. D Biol. Crystallogr. 52, 842–857 (1996).
Carroll, S.S. et al. Sensitivity of HIV-1 reverse transcriptase and its mutants to inhibition by azidothymidine triphosphate. Biochemistry 33, 2113–2120 (1994).
Kerr, S.G. & Anderson, K.S. Pre-steady-state kinetic characterization of wild type and 3′-azido-3′-deoxythymidine (AZT) resistant human immunodeficiency virus type 1 reverse transcriptase: implication of RNA directed DNA polymerization in the mechanism of AZT resistance. Biochemistry 36, 14064–14070 (1997).
Krebs, R., Immendorfer, U., Thrall, S.H., Wohrl, B.M. & Goody, R.S. Single-step kinetics of HIV-1 reverse transcriptase mutants responsible for virus resistance to nucleoside inhibitors zidovudine and 3-TC. Biochemistry 36, 10292–10300 (1997).
Hsiou, Y. et al. Structure of unliganded HIV-1 reverse transcriptase at 2.7 Å resolution: implications of conformational changes for polymerization and inhibition mechanisms. Structure 4, 853–860 (1996).
Ueno, T. & Mitsuya, H. Comparative enzymatic study of HIV-1 reverse transcriptase resistant to 2′,3′-dideoxynucleotide analogs using the single-nucleotide incorporation assay. Biochemistry 36, 1092–1099 (1997).
Deval, J. et al. The molecular mechanism of multidrug resistance by the Q151M human immunodeficiency virus type 1 reverse transcriptase and its suppression using alpha-boranophosphate nucleotide analogues. J. Biol. Chem. 277, 42097–42104 (2002).
Sluis-Cremer, N. et al. The 3′-azido group is not the primary determinant of 3′-azido-3′-deoxythymidine (AZT) responsible for the excision phenotype of AZT-resistant HIV-1. J. Biol. Chem. 280, 29047–29052 (2005).
Meyer, P.R., Smith, A.J., Matsuura, S.E. & Scott, W.A. Chain-terminating dinucleoside tetraphosphates are substrates for DNA polymerization by human immunodeficiency virus type 1 reverse transcriptase with increased activity against thymidine analogue-resistant mutants. Antimicrob. Agents Chemother. 50, 3607–3614 (2006).
Ray, A.S. et al. Probing the molecular mechanisms of AZT drug resistance mediated by HIV-1 reverse transcriptase using a transient kinetic analysis. Biochemistry 42, 8831–8841 (2003).
Selmi, B. et al. The Y181C substitution in 3′-azido-3′-deoxythymidine-resistant human immunodeficiency virus, type 1, reverse transcriptase suppresses the ATP-mediated repair of the 3′-azido-3′-deoxythymidine 5′-monophosphate-terminated primer. J. Biol. Chem. 278, 40464–40472 (2003).
White, K.L. et al. Molecular mechanisms of tenofovir resistance conferred by human immunodeficiency virus type 1 reverse transcriptase containing a diserine insertion after residue 69 and multiple thymidine analog-associated mutations. Antimicrob. Agents Chemother. 48, 992–1003 (2004).
Boyer, P.L., Imamichi, T., Sarafianos, S.G., Arnold, E. & Hughes, S.H. Effects of the Delta67 complex of mutations in human immunodeficiency virus type 1 reverse transcriptase on nucleoside analog excision. J. Virol. 78, 9987–9997 (2004).
Hooker, D.J. et al. An in vivo mutation from leucine to tryptophan at position 210 in human immunodeficiency virus type 1 reverse transcriptase contributes to high-level resistance to 3′-azido-3′-deoxythymidine. J. Virol. 70, 8010–8018 (1996).
Harrigan, P.R. et al. Significance of amino acid variation at human immunodeficiency virus type 1 reverse transcriptase residue 210 for zidovudine susceptibility. J. Virol. 70, 5930–5934 (1996).
Romano, L. et al. Broad nucleoside-analogue resistance implications for human immunodeficiency virus type 1 reverse-transcriptase mutations at codons 44 and 118. J. Infect. Dis. 185, 898–904 (2002).
Marcelin, A.G. et al. Thymidine analogue reverse transcriptase inhibitors resistance mutations profiles and association to other nucleoside reverse transcriptase inhibitors resistance mutations observed in the context of virological failure. J. Med. Virol. 72, 162–165 (2004).
Girouard, M., Diallo, K., Marchand, B., McCormick, S. & Gotte, M. Mutations E44D and V118I in the reverse transcriptase of HIV-1 play distinct mechanistic roles in dual resistance to AZT and 3TC. J. Biol. Chem. 278, 34403–34410 (2003).
White, K.L. et al. The K65R reverse transcriptase mutation in HIV-1 reverses the excision phenotype of zidovudine resistance mutations. Antivir. Ther. 11, 155–163 (2006).
Parikh, U.M., Zelina, S., Sluis-Cremer, N. & Mellors, J.W. Molecular mechanisms of bidirectional antagonism between K65R and thymidine analog mutations in HIV-1 reverse transcriptase. AIDS 21, 1405–1414 (2007).
Gu, Z., Fletcher, R.S., Arts, E.J., Wainberg, M.A. & Parniak, M.A. The K65R mutant reverse transcriptase of HIV-1 cross-resistant to 2′, 3′-dideoxycytidine, 2′,3′-dideoxy-3′-thiacytidine, and 2′,3′-dideoxyinosine shows reduced sensitivity to specific dideoxynucleoside triphosphate inhibitors in vitro. J. Biol. Chem. 269, 28118–28122 (1994).
White, K.L. et al. A combination of decreased NRTI incorporation and decreased excision determines the resistance profile of HIV-1 K65R RT. AIDS 19, 1751–1760 (2005).
Das, K. et al. Crystal structures of clinically relevant Lys103Asn/Tyr181Cys double mutant HIV-1 reverse transcriptase in complexes with ATP and non-nucleoside inhibitor HBY 097. J. Mol. Biol. 365, 77–89 (2007).
Mansky, L.M. & Bernard, L.C. 3′-Azido-3′-deoxythymidine (AZT) and AZT-resistant reverse transcriptase can increase the in vivo mutation rate of human immunodeficiency virus type 1. J. Virol. 74, 9532–9539 (2000).
Das, K. et al. Molecular modeling and biochemical characterization reveal the mechanism of hepatitis B virus polymerase resistance to lamivudine (3TC) and emtricitabine (FTC). J. Virol. 75, 4771–4779 (2001).
Urban, S., Fischer, K.P. & Tyrrell, D.L. Efficient pyrophosphorolysis by a hepatitis B virus polymerase may be a primer-unblocking mechanism. Proc. Natl. Acad. Sci. USA 98, 4984–4989 (2001).
Deval, J., Powdrill, M.H., D'Abramo, C.M., Cellai, L. & Gotte, M. Pyrophosphorolytic excision of nonobligate chain terminators by hepatitis C virus NS5B polymerase. Antimicrob. Agents Chemother. 51, 2920–2928 (2007).
Acosta-Hoyos, A.J. & Scott, W.A. The role of nucleotide excision by reverse transcriptase in HIV drug resistance. Viruses 2, 372–394 (2010).
Kellam, P., Boucher, C.A. & Larder, B.A. Fifth mutation in human immunodeficiency virus type 1 reverse transcriptase contributes to the development of high-level resistance to zidovudine. Proc. Natl. Acad. Sci. USA 89, 1934–1938 (1992).
Deval, J. et al. Mechanistic basis for reduced viral and enzymatic fitness of HIV-1 reverse transcriptase containing both K65R and M184V mutations. J. Biol. Chem. 279, 509–516 (2004).
Sluis-Cremer, N., Arion, D., Kaushik, N., Lim, H. & Parniak, M.A. Mutational analysis of Lys65 of HIV-1 reverse transcriptase. Biochem. J. 348, 77–82 (2000).
Arion, D. & Parniak, M.A. HIV resistance to zidovudine: the role of pyrophosphorolysis. Drug Resist. Updat. 2, 91–95 (1999).
Yahi, N. et al. Mutation patterns of the reverse transcriptase and protease genes in human immunodeficiency virus type 1-infected patients undergoing combination therapy: survey of 787 sequences. J. Clin. Microbiol. 37, 4099–4106 (1999).
Cozzi-Lepri, A. et al. Thymidine analogue mutation profiles: factors associated with acquiring specific profiles and their impact on the virological response to therapy. Antivir. Ther. 10, 791–802 (2005).
Sarafianos, S.G. et al. Trapping HIV-1 reverse transcriptase before and after translocation on DNA. J. Biol. Chem. 278, 16280–16288 (2003).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Brünger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Acknowledgements
We acknowledge personnel at the Cornell High Energy Synchrotron Source and the Advanced Photon Source for support of data collection, the members of our laboratories, including R. Bandwar and S. Martinez, for valuable conversations and assistance, and P. Clark for assistance with protein preparation. We are grateful to the US National Institutes of Health (NIH; grants R37 MERIT Award AI 27690 to E.A. and P01 GM 066671 to E.A. and R.A.J.) for support of reverse transcriptase structural studies. S.H.H. was supported by the Intramural Research Program of NIH, US National Cancer Institute (NCI), Center for Cancer Research and US National Institute of General Medical Sciences. This research was supported, in part, by the Intramural Research Program of the NIH, NCI, Center for Cancer Research.
Author information
Authors and Affiliations
Contributions
X.T. carried out the research, did analysis and helped with writing the manuscript. K.D. advised on crystallographic studies and did analysis, writing and editing of the manuscript. Q.H., J.D.B., X.H., B.L.G. and R.A.J. contributed special reagents. A.D.C. and Y.V.F. carried out parts of the research. P.L.B. contributed special reagents and manuscript editing. S.H.H. carried out analysis and manuscript writing and editing. S.G.S. designed the research and advised on experiments and manuscript editing. E.A. designed the research, supervised the project and did analysis, writing and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 570 kb)
Rights and permissions
About this article
Cite this article
Tu, X., Das, K., Han, Q. et al. Structural basis of HIV-1 resistance to AZT by excision. Nat Struct Mol Biol 17, 1202–1209 (2010). https://doi.org/10.1038/nsmb.1908
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1908
This article is cited by
-
Structural features in common of HBV and HIV-1 resistance against chirally-distinct nucleoside analogues entecavir and lamivudine
Scientific Reports (2020)
-
Elucidating molecular interactions of L-nucleotides with HIV-1 reverse transcriptase and mechanism of M184V-caused drug resistance
Communications Biology (2019)
-
AZT resistance alters enzymatic properties and creates an ATP-binding site in SFVmac reverse transcriptase
Retrovirology (2015)
-
Prediction of HIV drug resistance from genotype with encoded three-dimensional protein structure
BMC Genomics (2014)
-
Common and unique features of viral RNA-dependent polymerases
Cellular and Molecular Life Sciences (2014)