Abstract
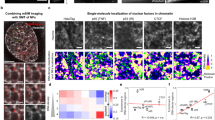
DNA-binding proteins survey genomes for targets using facilitated diffusion, which typically includes a one-dimensional (1D) scanning component for sampling local regions. Eukaryotic proteins must accomplish this task while navigating through chromatin. Yet it is unknown whether nucleosomes disrupt 1D scanning or eukaryotic DNA-binding factors can circumnavigate nucleosomes without falling off DNA. Here we use single-molecule microscopy in conjunction with nanofabricated curtains of DNA to show that the postreplicative mismatch repair protein complex Mlh1–Pms1 diffuses in 1D along DNA via a hopping/stepping mechanism and readily bypasses nucleosomes. This is the first experimental demonstration that a passively diffusing protein can traverse stationary obstacles. In contrast, Msh2–Msh6, a mismatch repair protein complex that slides while maintaining continuous contact with DNA, experiences a boundary upon encountering nucleosomes. These differences reveal important mechanistic constraints affecting intranuclear trafficking of DNA-binding proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
von Hippel, P.H. & Berg, O. Facilitated target location in biological systems. J. Biol. Chem. 264, 675–678 (1989).
Elf, J., Li, G.-W. & Xie, X. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Hager, G.L., McNally, J. & Misteli, T. Transcription dynamics. Mol. Cell 35, 741–753 (2009).
Li, G.-W., Berg, O. & Elf, J. Effects of macromolecular crowding and DNA looping on gene regulation kinetics. Nat. Phys. 5, 294–297 (2009).
Blainey, P.C., van Oijen, A.M., Banerjee, A., Verdine, G.L. & Xie, X.S. A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA. Proc. Natl. Acad. Sci. USA 103, 5752–5757 (2006).
Gorman, J. & Greene, E.C. Visualizing one-dimensional diffusion of proteins along DNA. Nat. Struct. Mol. Biol. 15, 768–774 (2008).
Gorski, S.A., Dundr, M. & Misteli, T. The road much traveled: trafficking in the cell nucleus. Curr. Opin. Cell Biol. 18, 284–290 (2006).
Kampmann, M. Facilitated diffusion in chromatin lattices: mechanistic diversity and regulatory potential. Mol. Microbiol. 57, 889–899 (2005).
Hodges, C., Bintu, L., Lubkowska, L., Kashlev, M. & Bustamante, C. Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II. Science 325, 626–628 (2009).
Studitsky, V.M., Clark, D. & Felsenfeld, G. Overcoming a nucleosomal barrier to transcription. Cell 83, 19–27 (1995).
Jiricny, J. The multifaceted mismatch-repair system. Nat. Rev. Mol. Cell Biol. 7, 335–346 (2006).
Modrich, P. Mechanisms in eukaryotic mismatch repair. J. Biol. Chem. 281, 30305–30309 (2006).
Kunkel, T.A. & Erie, D.A. DNA mismatch repair. Annu. Rev. Biochem. 74, 681–710 (2005).
Gorman, J. et al. Dynamic basis for one-dimensional DNA scanning by the mismatch repair complex Msh2-Msh6. Mol. Cell 28, 359–370 (2007).
Gorman, J., Fazio, T., Wang, F., Wind, S. & Greene, E. Nanofabricated racks of aligned and anchored DNA substrates for single-molecule imaging. Langmuir 26, 1372–1379 (2010).
Hall, M.C., Wang, H., Erie, D.A. & Kunkel, T.A. High affinity cooperative DNA binding by the yeast Mlh1-Pms1 heterodimer. J. Mol. Biol. 312, 637–647 (2001).
Sacho, E.J., Kadyrov, F.A., Modrich, P., Kunkel, T.A. & Erie, D.A. Direct visualization of asymmetric adenine-nucleotide-induced conformational changes in MutL α. Mol. Cell 29, 112–121 (2008).
Guarné, A., Junop, M. & Yang, W. Structure and function of the N-terminal 40 kDa fragment of human PMS2: a monomeric GHL ATPase. EMBO J. 20, 5521–5531 (2001).
Guarné, A. et al. Structure of the MutL C-terminal domain: a model of intact MutL and its roles in mismatch repair. EMBO J. 23, 4134–4145 (2004).
Kosinski, J., Steindorf, I., Bujnicki, J., Giron-Monzon, L. & Friedhoff, P. Analysis of the quaternary structure of the MutL C-terminal domain. J. Mol. Biol. 351, 895–909 (2005).
Komazin-Meredith, G., Mirchev, R., Golan, D., van Oijen, A. & Coen, D. Hopping of a processivity factor on DNA revealed by single-molecule assays of diffusion. Proc. Natl. Acad. Sci. USA 105, 10721–10726 (2008).
Hall, M.C. et al. DNA binding by yeast Mlh1 and Pms1: implications for DNA mismatch repair. Nucleic Acids Res. 31, 2025–2034 (2003).
Visnapuu, M.-L. & Greene, E. Single-molecule imaging of DNA curtains reveals intrinsic energy landscapes for nucleosome deposition. Nat. Struct. Mol. Biol. 16, 1056–1062 (2009).
Warren, J.J. et al. Structure of the human MutSα DNA lesion recognition complex. Mol. Cell 26, 579–592 (2007).
Li, F., Tian, L., Gu, L. & Li, G. Evidence that nucleosomes inhibit mismatch repair in eukaryotic cells. J. Biol. Chem. 284, 33056–33061 (2009).
Gradia, S. et al. hMSH2-hMSH6 forms a hydrolysis-independent sliding clamp on mismatched DNA. Mol. Cell 3, 255–261 (1999).
Pluciennik, A. & Modrich, P. Protein roadblocks and helix discontinuities are barriers to the initiation of mismatch repair. Proc. Natl. Acad. Sci. USA 104, 12709–12713 (2007).
Mendillo, M.L., Mazur, D.J. & Kolodner, R.D. Analysis of the interaction between the Saccharomyces cerevisiae MSH2–MSH6 and MLH1–PMS1 complexes with DNA using a reversible DNA end-blocking system. J. Biol. Chem. 280, 22245–22257 (2005).
Berg, O.G., Winter, R. & von Hippel, P. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Winter, R.B., Berg, O. & von Hippel, P. Diffusion-driven mechanisms of protein translocation on nucleic acids. 3. The Escherichia coli lac repressor–operator interaction: kinetic measurements and conclusions. Biochemistry 20, 6961–6977 (1981).
Winter, R.B. & von Hippel, P. Diffusion-driven mechanisms of protein translocation on nucleic acids. 2. The Escherichia coli repressor–operator interaction: equilibrium measurements. Biochemistry 20, 6948–6960 (1981).
Riggs, A.D., Bourgeois, S. & Cohn, M. The lac repressor-operator interaction. 3. Kinetic studies. J. Mol. Biol. 53, 401–417 (1970).
Doucleff, M. & Clore, G. Global jumping and domain-specific intersegment transfer between DNA cognate sites of the multidomain transcription factor Oct-1. Proc. Natl. Acad. Sci. USA 105, 13871–13876 (2008).
Iwahara, J. & Clore, G. Direct observation of enhanced translocation of a homeodomain between DNA cognate sites by NMR exchange spectroscopy. J. Am. Chem. Soc. 128, 404–405 (2006).
Iwahara, J., Zweckstetter, M. & Clore, G. NMR structural and kinetic characterization of a homeodomain diffusing and hopping on nonspecific DNA. Proc. Natl. Acad. Sci. USA 103, 15062–15067 (2006).
Roy, R., Kozlov, A., Lohman, T. & Ha, T. SSB protein diffusion on single-stranded DNA stimulates RecA filament formation. Nature 461, 1092–1097 (2009).
Liu, S., Abbondanzieri, E., Rausch, J., Le Grice, S. & Zhuang, X. Slide into action: dynamic shuttling of HIV reverse transcriptase on nucleic acid substrates. Science 322, 1092–1097 (2008).
Tafvizi, A. et al. Tumor suppressor p53 slides on DNA with low friction and high stability. Biophys. J. 95, L01–L03 (2008).
Halford, S.E. & Szczelkun, M. How to get from A to B: strategies for analysing protein motion on DNA. Eur. Biophys. J. 31, 257–267 (2002).
Mirny, L. et al. How a protein searches for its site on DNA: the mechanism of facilitated diffusion. J. Phys. A: Math. Theor. 42, 434013 (2009).
Groth, A., Rocha, W., Verreault, A. & Almouzni, G. Chromatin challenges during DNA replication and repair. Cell 128, 721–733 (2007).
Umar, A. et al. Requirement for PCNA in DNA mismatch repair at a step preceding DNA resynthesis. Cell 87, 65–73 (1996).
Kolodner, R.D., Mendillo, M.L. & Putnam, C.D. Coupling distant sites in DNA during mismatch repair. Proc. Natl. Acad. Sci. USA 104, 12953–12954 (2007).
Bagchi, B., Blainey, P.C. & Xie, X.S. Diffusion constant of a nonspecifically bound protein undergoing curvilinear motion along DNA. J. Phys. Chem. B 112, 6282–6284 (2008).
Luger, K., Mäder, A., Richmond, R., Sargent, D. & Richmond, T. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Pathak, S., Davidson, M. & Silva, G. Characterization of the functional binding properties of antibody conjugated quantum dots. Nano Lett. 7, 1839–1845 (2007).
Acknowledgements
We thank members of our laboratories for carefully reading the manuscript and providing suggestions throughout this study. This work was supported by the Howard Hughes Medical Institute, by a US National Science Foundation Presidential Early Career Award for Scientists and Engineers, by US National Institutes of Health (NIH) grant GM082848 to E.C.G. and by NIH grant GM53085 to E.A. J.G. is supported by an NIH training grant for Cellular and Molecular Foundations of Biomedical Sciences (T32GM00879807). A.J.P. was supported by a State University of New York fellowship. This work was partially supported by the Initiatives in Science and Engineering program through Columbia University, the Nanoscale Science and Engineering Initiative of the National Science Foundation under US National Science Foundation Award Number CHE-0641523 and by the New York State Office of Science, Technology, and Academic Research.
Author information
Authors and Affiliations
Contributions
J.G. designed the TIRFM experiments and collected and analyzed the single-molecule data; A.J.P. designed and engineered all constructs, expressed and purified the proteins, did all ensemble-level characterization and conducted the immunoprecipitation experiments; M.-L.V. made and characterized all chromatin substrates; J.G., A.J.P., M.-L.V., E.A. and E.C.G. discussed the data and cowrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Table 1, Supplementary Discussion and Supplementary Methods (PDF 3298 kb)
Supplementary Video 1
Double-tethered DNA curtains. (MOV 811 kb)
Supplementary Video 2
Double-tethered DNA curtains with diffusing Mlh1-Pms1. (MOV 16938 kb)
Rights and permissions
About this article
Cite this article
Gorman, J., Plys, A., Visnapuu, ML. et al. Visualizing one-dimensional diffusion of eukaryotic DNA repair factors along a chromatin lattice. Nat Struct Mol Biol 17, 932–938 (2010). https://doi.org/10.1038/nsmb.1858
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1858
This article is cited by
-
MutS and MutL sliding clamps in DNA mismatch repair
Genome Instability & Disease (2022)
-
A mini-review of the diffusion dynamics of DNA-binding proteins: experiments and models
Journal of the Korean Physical Society (2021)
-
Inferring quantity and qualities of superimposed reaction rates from single molecule survival time distributions
Scientific Reports (2020)
-
Dynamical reorganization of the pluripotency transcription factors Oct4 and Sox2 during early differentiation of embryonic stem cells
Scientific Reports (2020)
-
MutL sliding clamps coordinate exonuclease-independent Escherichia coli mismatch repair
Nature Communications (2019)