Abstract
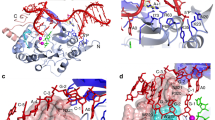
Family X polymerases such as DNA polymerase λ (Pol λ) are well suited for filling short gaps during DNA repair because they simultaneously bind both the 5′ and 3′ ends of short gaps. DNA binding and gap filling are well characterized for 1-nucleotide (nt) gaps, but the location of yet-to-be-copied template nucleotides in longer gaps is unknown. Here we present crystal structures revealing that, when bound to a 2-nt gap, Pol λ scrunches the template strand and binds the additional uncopied template base in an extrahelical position within a binding pocket that comprises three conserved amino acids. Replacing these amino acids with alanine results in less processive gap filling and less efficient NHEJ when 2-nt gaps are involved. Thus, akin to scrunching by RNA polymerase during transcription initiation, scrunching occurs during gap filling DNA synthesis associated with DNA repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Moon, A.F. et al. The X family portrait: structural insights into biological functions of X family polymerases. DNA Repair (Amst.) 6, 1709–1725 (2007).
Beard, W.A. & Wilson, S.H. Structure and mechanism of DNA polymerase β. Chem. Rev. 106, 361–382 (2006).
García-Díaz, M. et al. DNA polymerase λ, a novel DNA repair enzyme in human cells. J. Biol. Chem. 277, 13184–13191 (2002).
García-Díaz, M., Bebenek, K., Kunkel, T.A. & Blanco, L. Identification of an intrinsic 5′-deoxyribose-5-phosphate lyase activity in human DNA polymerase λ: a possible role in base excision repair. J. Biol. Chem. 276, 34659–34663 (2001).
Braithwaite, E.K. et al. DNA polymerase λ protects mouse fibroblasts against oxidative DNA damage and is recruited to sites of DNA damage/repair. J. Biol. Chem. 280, 31641–31647 (2005).
Tano, K. et al. Interplay between DNA polymerases β and λ in repair of oxidation DNA damage in chicken DT40 cells. DNA Repair (Amst.) 6, 869–875 (2007).
Nick McElhinny, S.A. et al. A gradient of template dependence defines distinct biological roles for family X polymerases in nonhomologous end joining. Mol. Cell 19, 357–366 (2005).
Lee, J.W. et al. Implication of DNA polymerase λ in alignment-based gap filling for nonhomologous DNA end joining in human nuclear extracts. J. Biol. Chem. 279, 805–811 (2004).
Lieber, M.R., Lu, H., Gu, J. & Schwarz, K. Flexibility in the order of action and in the enzymology of the nuclease, polymerases, and ligase of vertebrate non-homologous DNA end joining: relevance to cancer, aging, and the immune system. Cell Res. 18, 125–133 (2008).
Weterings, E. & Chen, D.J. The endless tale of non-homologous end-joining. Cell Res. 18, 114–124 (2008).
Garcia-Diaz, M., Bebenek, K., Krahn, J.M., Kunkel, T.A. & Pedersen, L.C. A closed conformation for the Pol λ catalytic cycle. Nat. Struct. Mol. Biol. 12, 97–98 (2005).
Singhal, R.K. & Wilson, S.H. Short gap-filling synthesis by DNA polymerase β is processive. J. Biol. Chem. 268, 15906–15911 (1993).
Sawaya, M.R., Prasad, R., Wilson, S.H., Kraut, J. & Pelletier, H. Crystal structures of human DNA polymerase β complexed with gapped and nicked DNA: evidence for an induced fit mechanism. Biochemistry 36, 11205–11215 (1997).
Moon, A.F. et al. Structural insight into the substrate specificity of DNA polymerase μ. Nat. Struct. Mol. Biol. 14, 45–53 (2007).
van Gent, D.C. & van der Burg, M. Non-homologous end-joining, a sticky affair. Oncogene 26, 7731–7740 (2007).
Klungland, A. & Lindahl, T. Second pathway for completion of human DNA base excision-repair: reconstitution with purified proteins and requirement for DNase IV (FEN1). EMBO J. 16, 3341–3348 (1997).
Davis, B.J., Havener, J.M. & Ramsden, D.A. End-bridging is required for Pol μ to efficiently promote repair of noncomplementary ends by nonhomologous end joining. Nucleic Acids Res. 36, 3085–3094 (2008).
Brissett, N.C. et al. Structure of a NHEJ polymerase-mediated DNA synaptic complex. Science 318, 456–459 (2007).
Garcia-Diaz, M. et al. A structural solution for the DNA polymerase λ-dependent repair of DNA gaps with minimal homology. Mol. Cell 13, 561–572 (2004).
Cheetham, G.M. & Steitz, T.A. Structure of a transcribing T7 RNA polymerase initiation complex. Science 286, 2305–2309 (1999).
Kapanidis, A.N. et al. Initial transcription by RNA polymerase proceeds through a DNA-scrunching mechanism. Science 314, 1144–1147 (2006).
Ikeda, R.A. & Richardson, C.C. Interactions of the RNA polymerase of bacteriophage T7 with its promoter during binding and initiation of transcription. Proc. Natl. Acad. Sci. USA 83, 3614–3618 (1986).
Bebenek, K., Garcia-Diaz, M., Blanco, L. & Kunkel, T.A. The frameshift infidelity of human DNA polymerase λ. Implications for function. J. Biol. Chem. 278, 34685–34690 (2003).
Garcia-Diaz, M., Bebenek, K., Krahn, J.M., Pedersen, L.C. & Kunkel, T.A. Structural analysis of strand misalignment during DNA synthesis by a human DNA polymerase. Cell 124, 331–342 (2006).
Bebenek, K. et al. Substrate-induced DNA strand misalignment during catalytic cycling by DNA polymerase λ. EMBO Rep. 9, 459–464 (2008).
Garcia-Diaz, M. & Kunkel, T.A. Mechanism of a genetic glissando: structural biology of indel mutations. Trends Biochem. Sci. 31, 206–214 (2006).
Tang, G.Q., Roy, R., Ha, T. & Patel, S.S. Transcription initiation in a single-subunit RNA polymerase proceeds through DNA scrunching and rotation of the N-terminal subdomains. Mol. Cell 30, 567–577 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Bebenek, K., Garcia-Diaz, M., Patishall, S.R. & Kunkel, T.A. Biochemical properties of Saccharomyces cerevisiae DNA polymerase IV. J. Biol. Chem. 280, 20051–20058 (2005).
Kokoska, R.J., McCulloch, S.D. & Kunkel, T.A. The efficiency and specificity of apurinic/apyrimidinic site bypass by human DNA polymerase η and Sulfolobus solfataricus Dpo4. J. Biol. Chem. 278, 50537–50545 (2003).
Osheroff, W.P., Jung, H.K., Beard, W.A., Wilson, S.H. & Kunkel, T.A. The fidelity of DNA polymerase β during distributive and processive DNA synthesis. J. Biol. Chem. 274, 3642–3650 (1999).
Pérez, A. et al. Refinement of the AMBER force field for nucleic acids: improving the description of α/γ conformers. Biophys. J. 92, 3817–3829 (2007).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Case, D.A. et al. AMBER 10 (University of California, San Francisco, CA, 2008).
Acknowledgements
We thank L. Pedersen, W. Beard and B. Van Houten for critical reading of the manuscript. This work was partly supported by Project Z01 ES065070 to T.A.K. from the Division of Intramural Research of the US National Institutes of Health (NIH), National Institute of Environmental Health Sciences and by NIH grant CA 097096 to D.A.R. Use of the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. W-31-109-Eng-38.
Author information
Authors and Affiliations
Contributions
T.A.K. and D.A.R. supervised the study, designed experiments and analyzed data; M.G.-D., K.B., A.A.L. and J.M.H. performed experiments and analyzed data; L.C.P. contributed to structure refinement; J.M.K. and L.P. performed molecular dynamics simulations; J.M.K. prepared the Supplementary Video 1; all authors contributed to writing the manuscript.
Corresponding author
Supplementary information
Supplementary Video 1
Model of gap filling by Pol λ. The first incoming dNTP and matching template base are in green. The second incoming and template bases are in magenta. Residues at the scrunched binding site are in orange (Leu277, His511, Arg514). Side chains are shown for Arg517 near the template strand, and the following residues, from left to right: Asp490, Asp427, Asp429, Arg386, Tyr505 and Arg420. The dRP domain is light purple, the fingers domain is light cyan, the palm domain is light green and the thumb domain is beige. The animation shows transitions between nine models, each using experimental coordinates except for DNA base substitutions modeled to maintain the same sequence. The catalytic transitions use coordinates from superimposed models to produce the complete set of coordinates. Transitions were modeled with CNS, using stepped distance restraints as a tractor-beam, along with standard geometry restraints. This is meant as a tool to aid in visualizing the current series of structures. It is not a rigorous dynamics simulation, and may be missing intermediate details not present in the known structures. (AVI 5578 kb)
1) Scrunching and incoming dNTP (starts from 1RZT, ends at current structure)
2) Activation #1 of scrunched structure (active site with Mg from 2PFO)
3) Catalysis #1 (active site from 1XSP1)
4) Translocation from scrunched product to 1nt-gap (1XSL), PPi+Mg release
5) dNTP binding to 1-nt gap with DNA template-strand docking (1XSN)
6) Activation #2 of 1nt-gap (active site with Mg from 2PFO)
7) Catalysis #2 (active site from 1XSP1)
8) Relaxation after Catalysis #2 (ending at 1XSP2), and PPi+Mg release
Rights and permissions
About this article
Cite this article
Garcia-Diaz, M., Bebenek, K., Larrea, A. et al. Template strand scrunching during DNA gap repair synthesis by human polymerase λ. Nat Struct Mol Biol 16, 967–972 (2009). https://doi.org/10.1038/nsmb.1654
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1654
This article is cited by
-
Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints
Nature Communications (2022)
-
Analysis of diverse double-strand break synapsis with Polλ reveals basis for unique substrate specificity in nonhomologous end-joining
Nature Communications (2022)
-
Molecular basis for DNA repair synthesis on short gaps by mycobacterial Primase-Polymerase C
Nature Communications (2020)
-
Structure and function relationships in mammalian DNA polymerases
Cellular and Molecular Life Sciences (2020)
-
PAXX and its paralogs synergistically direct DNA polymerase λ activity in DNA repair
Nature Communications (2018)