Abstract
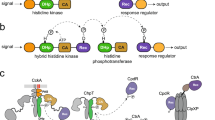
The mechanistic understanding of how membrane-embedded sensor kinases recognize signals and regulate kinase activity is currently limited. Here we report structure-function relationships of the multidomain membrane sensor kinase DcuS using solid-state NMR, structural modeling and mutagenesis. Experimental data of an individual cytoplasmic Per-Arnt-Sim (PAS) domain were compared to structural models generated in silico. These studies, together with previous NMR work on the periplasmic PAS domain, enabled structural investigations of a membrane-embedded 40-kDa construct by solid-state NMR, comprising both PAS segments and the membrane domain. Structural alterations are largely limited to protein regions close to the transmembrane segment. Data from isolated and multidomain constructs favor a disordered N-terminal helix in the cytoplasmic domain. Mutations of residues in this region strongly influence function, suggesting that protein flexibility is related to signal transduction toward the kinase domain and regulation of kinase activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Szurmant, H., White, R.A. & Hoch, J.A. Sensor complexes regulating two-component signal transduction. Curr. Opin. Struct. Biol. 17, 706–715 (2007).
Mascher, T., Helmann, J.D. & Unden, G. Stimulus perception in bacterial signal-transducing histidine kinases. Microbiol. Mol. Biol. Rev. 70, 910–938 (2006).
Zientz, E., Bongaerts, J. & Unden, G. Fumarate regulation of gene expression in Escherichia coli by the DcuSR (dcuSR genes) two-component regulatory system. J. Bacteriol. 180, 5421–5425 (1998).
Golby, P., Davies, S., Kelly, D.J., Guest, J.R. & Andrews, S.C. Identification and characterization of a two-component sensor-kinase and response-regulator system (DcuS-DcuR) controlling gene expression in response to C4-dicarboxylates in Escherichia coli. J. Bacteriol. 181, 1238–1248 (1999).
Janausch, I.G., Garcia-Moreno, I. & Unden, G. Function of DcuS from Escherichia coli as a fumarate-stimulated histidine protein kinase in vitro. J. Biol. Chem. 277, 39809–39814 (2002).
Bader, M.W. et al. Recognition of antimicrobial peptides by a bacterial sensor kinase. Cell 122, 461–472 (2005).
Neiditch, M.B. et al. Ligand-induced asymmetry in histidine sensor kinase complex regulates quorum sensing. Cell 126, 1095–1108 (2006).
Rossmann, M.G., Morais, M.C., Leiman, P.G. & Zhang, W. Combining X-ray crystallography and electron microscopy. Structure 13, 355–362 (2005).
Alber, F. et al. Determining the architectures of macromolecular assemblies. Nature 450, 683–694 (2007).
Pickford, A.R. & Campbell, I.D. NMR studies of modular protein structures and their interactions. Chem. Rev. 104, 3557–3566 (2004).
Topf, M. & Sali, A. Combining electron microscopy and comparative protein structure modeling. Curr. Opin. Struct. Biol. 15, 578–585 (2005).
Qian, B. et al. High-resolution structure prediction and the crystallographic phase problem. Nature 450, 259–264 (2007).
Meiler, J. & Baker, D. Rapid protein fold determination using unassigned NMR data. Proc. Natl. Acad. Sci. USA 100, 15404–15409 (2003).
Cavalli, A., Salvatella, X., Dobson, C.M. & Vendruscolo, M. Protein structure determination from NMR chemical shifts. Proc. Natl. Acad. Sci. USA 104, 9615–9620 (2007).
Schueler-Furman, O., Wang, C., Bradley, P., Misura, K. & Baker, D. Progress in modeling of protein structures and interactions. Science 310, 638–642 (2005).
Sali, A., Glaeser, R., Earnest, T. & Baumeister, W. From words to literature in structural proteomics. Nature 422, 216–225 (2003).
Lee, A.G. How lipids affect the activities of integral membrane proteins. Biochim. Biophys. Acta 1666, 62–87 (2004).
Cierpicki, T., Bushweller, J.H. & Derewenda, Z.S. Probing the supramodular architecture of a multidomain protein: the structure of syntenin in solution. Structure 13, 319–327 (2005).
Thompson, L.K. Solid-state NMR studies of the structure and mechanisms of proteins. Curr. Opin. Struct. Biol. 12, 661–669 (2002).
Patel, A.B. et al. Coupling of retinal isomerization to the activation of rhodopsin. Proc. Natl. Acad. Sci. USA 101, 10048–10053 (2004).
Kim, D.E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Pappalardo, L. et al. The NMR structure of the sensory domain of the membranous two-component fumarate sensor (histidine protein kinase) DcuS of Escherichia coli. J. Biol. Chem. 278, 39185–39188 (2003).
Etzkorn, M., Böckmann, A., Penin, F., Riedel, D. & Baldus, M. Characterization of folding intermediates of a domain-swapped protein by solid-state NMR spectroscopy. J. Am. Chem. Soc. 129, 169–175 (2007).
Vuister, G.W., Kim, S.J., Wu, C. & Bax, A. 2D and 3D NMR-study of phenylalanine residues in proteins by reverse isotopic labeling. J. Am. Chem. Soc. 116, 9206–9210 (1994).
Heise, H. et al. Molecular-level secondary structure, polymorphism, and dynamics of full-length α-synuclein fibrils studied by solid-state NMR. Proc. Natl. Acad. Sci. USA 102, 15871–15876 (2005).
Baldus, M. Molecular interactions investigated by multi-dimensional solid-state NMR. Curr. Opin. Struct. Biol. 16, 618–623 (2006).
Watts, K.J., Johnson, M.S. & Taylor, B.L. Minimal requirements for oxygen sensing by the aerotaxis receptor Aer. Mol. Microbiol. 59, 1317–1326 (2006).
Taylor, B.L. & Zhulin, I.B. PAS domains: internal sensors of oxygen, redox potential, and light. Microbiol. Mol. Biol. Rev. 63, 479–506 (1999).
Jones, D.T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999).
Wang, Y. & Jardetzky, O. Probability-based protein secondary structure identification using combined NMR chemical-shift data. Protein Sci. 11, 852–861 (2002).
Wishart, D.S. & Sykes, B.D. Chemical-shifts as a tool for structure determination. Methods Enzymol. 239, 363–392 (1994).
Neal, S., Nip, A.M., Zhang, H. & Wishart, D.S. Rapid and accurate calculation of protein 1H, 13C and 15N chemical shifts. J. Biomol. NMR 26, 215–240 (2003).
Lange, A. et al. A concept for rapid protein-structure determination by solid-state NMR spectroscopy. Angew. Chem. Int. Edn. Engl. 44, 2089–2092 (2005).
Vreede, J., van der Horst, M.A., Hellingwerf, K.J., Crielaard, W. & van Aalten, D.M.F. PAS domains. Common structure and common flexibility. J. Biol. Chem. 278, 18434–18439 (2003).
Pandini, A. & Bonati, L. Conservation and specialization in PAS domain dynamics. Protein Eng. Des. Sel. 18, 127–137 (2005).
Lange, A., Luca, S. & Baldus, M. Structural constraints from proton-mediated rare-spin correlation spectroscopy in rotating solids. J. Am. Chem. Soc. 124, 9704–9705 (2002).
Seidel, K., Etzkorn, M., Heise, H., Becker, S. & Baldus, M. High-resolution solid-state NMR studies on uniformly [13C,15N]-labeled ubiquitin. ChemBioChem 6, 1638–1647 (2005).
Sevvana, M. et al. A ligand-induced switch in the periplasmic domain of sensor histidine kinase CitA. J. Mol. Biol. 377, 512–523 (2008).
Kneuper, H. et al. The nature of the stimulus and of the fumarate binding site of the fumarate sensor DcuS of Escherichia coli. J. Biol. Chem. 280, 20596–20603 (2005).
Key, J., Hefti, M., Purcell, E.B. & Moffat, K. Structure of the redox sensor domain of Azotobacter vinelandii NifL at atomic resolution: signaling, dimerization, and mechanism. Biochemistry 46, 3614–3623 (2007).
Repik, A. et al. PAS domain residues involved in signal transduction by the Aer redox sensor of Escherichia coli. Mol. Microbiol. 36, 806–816 (2000).
Six, S., Andrews, S.C., Unden, G. & Guest, J.R. Escherichia coli possesses two homologous anaerobic C4-dicarboxylate membrane transporters (DcuA and DcuB) distinct from the aerobic dicarboxylate transport system (Dct). J. Bacteriol. 176, 6470–6478 (1994).
Janausch, I.G., Zientz, E., Tran, Q.H., Kröger, A. & Unden, G. C4-dicarboxylate carriers and sensors in bacteria. Biochim. Biophys. Acta 1553, 39–56 (2002).
Marina, A., Waldburger, C.D. & Hendrickson, W.A. Structure of the entire cytoplasmic portion of a sensor histidine-kinase protein. EMBO J. 24, 4247–4259 (2005).
Zoltowski, B.D. et al. Conformational switching in the fungal light sensor Vivid. Science 316, 1054–1057 (2007).
Harper, S.M., Neil, L.C. & Gardner, K.H. Structural basis of a phototropin light switch. Science 301, 1541–1544 (2003).
Watts, K.J., Sommer, K., Fry, S.L., Johnson, M.S. & Taylor, B.L. Function of the N-terminal cap of the PAS domain in signaling by the aerotaxis receptor Aer. J. Bacteriol. 188, 2154–2162 (2006).
Ma, X., Sayed, N., Baskaran, P., Beuve, A. & van den Akker, F. PAS-mediated dimerization of soluble guanylyl cyclase revealed by signal transduction histidine kinase domain crystal structure. J. Biol. Chem. 283, 1167–1178 (2008).
Lee, J. et al. Changes at the KinA PAS-A dimerization interface influence histidine kinase function. Biochemistry 47, 4051–4064 (2008).
Abo-Amer, A.E. et al. DNA interaction and phosphotransfer of the C4-dicarboxylate-responsive DcuS-DcuR two-component regulatory system from Escherichia coli. J. Bacteriol. 186, 1879–1889 (2004).
Martin, R.W. & Zilm, K.W. Preparation of protein nanocrystals and their characterization by solid state NMR. J. Magn. Reson. 165, 162–174 (2003).
Miroux, B. & Walker, J.E. Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J. Mol. Biol. 260, 289–298 (1996).
Miller, J.H. A Short Course In Bacterial Genetics (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1992).
Rigaud, J.L., Pitard, B. & Levy, D. Reconstitution of membrane proteins into liposomes: application to energy-transducing membrane proteins. Biochim. Biophys. Acta 1231, 223–246 (1995).
Fung, B.M., Khitrin, A.K. & Ermolaev, K. An improved broadband decoupling sequence for liquid crystals and solids. J. Magn. Reson. 142, 97–101 (2000).
Schwieters, C.D., Kuszewski, J.J., Tjandra, N. & Clore, G.M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Stein, E.G., Rice, L.M. & Brunger, A.T. Torsion-angle molecular dynamics as a new efficient tool for NMR structure calculation. J. Magn. Reson. 124, 154–164 (1997).
Linge, J.P., Williams, M.A., Spronk, C.A.E.M., Bonvin, A.M.J.J. & Nilges, M. Refinement of protein structures in explicit solvent. Proteins Struct. Funct. Genet. 50, 496–506 (2003).
Acknowledgements
Technical assistance by B. Angerstein, K. Sabagh and A.-K. Brückner is gratefully acknowledged. This work was funded by the DFG (BA 1700/6-2; UN 49/6 and UN49/8).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1 and 2 and Supplementary Methods. (PDF 1275 kb)
Rights and permissions
About this article
Cite this article
Etzkorn, M., Kneuper, H., Dünnwald, P. et al. Plasticity of the PAS domain and a potential role for signal transduction in the histidine kinase DcuS. Nat Struct Mol Biol 15, 1031–1039 (2008). https://doi.org/10.1038/nsmb.1493
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1493