Abstract
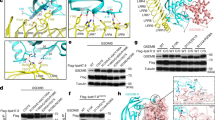
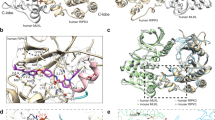
Granzyme A (GzmA) belongs to a family of trypsin-like serine proteases localized in cytoplasmic granules of activated lymphocytes and natural killer (NK) cells. In contrast to the related granzyme B (GzmB), GzmA forms a stable disulfide-linked homodimer and triggers target-cell death in a caspase-independent way. Limited proteolysis of a high-molecular-mass complex containing SET (also named putative HLA-associated protein II or PHAPII), PHAPI (pp32, leucine-rich acidic nuclear protein) and HMG2 by GzmA liberates NM23-H1, a Mg2+-dependent DNase that causes single-stranded breaks in nuclear DNA. By analyzing the dimeric GzmA structure at a resolution of 2.5 Å, we determined the substrate-binding constraints and selective advantages of the two domains arranged as a unique functional tandem. The active sites of the two subunits point in opposite directions and the nearby noncatalytic surfaces can function as exosites, presenting substrates to the active site region of the adjacent partner in a manner analogous to staphylokinase or streptokinase, which present plasminogen to the cofactor–plasmin and cofactor–plasminogen complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pasternack, M.S. & Eisen, H.N. A novel serine esterase expressed by cytotoxic T lymphocytes. Nature 314, 743–745 (1985).
Pasternack, M.S., Verret, C.R., Liu, M.A. & Eisen, H.N. Serine esterase in cytolytic T lymphocytes. Nature 322, 740–743 (1986).
Simon, M.M., Hoschutzky, H., Fruth, U., Simon, H.G. & Kramer, M.D. Purification and characterization of a T cell specific serine proteinase (TSP-1) from cloned cytolytic T lymphocytes. EMBO J. 5, 3267–3274 (1986).
Young, J.D. et al. Isolation and characterization of a serine esterase from cytolytic T cell granules. Cell 47, 183–194 (1986).
Masson, D. & Tschopp, J. A family of serine esterases in lytic granules of cytolytic T lymphocytes. Cell 49, 679–685 (1987).
Fruth, U. et al. A novel serine proteinase (HuTSP) isolated from a cloned human CD8+ cytolytic T cell line is expressed and secreted by activated CD4+ and CD8+ lymphocytes. Eur. J. Immunol. 17, 1625–1633 (1987).
Krähenbühl, O. et al. Characterization of granzymes A and B isolated from granules of cloned human cytotoxic T lymphocytes. J. Immunol. 141, 3471–3477 (1988).
Poe, M. et al. Human cytotoxic lymphocyte tryptase. Its purification from granules and the characterization of inhibitor and substrate specificity. J. Biol. Chem. 263, 13215–13222 (1988).
Gershenfeld, H.K., Hershberger, R.J., Shows, T.B. & Weissman, I.L. Cloning and chromosomal assignment of a human cDNA encoding a T cell- and natural killer cell-specific trypsin-like serine protease. Proc. Natl. Acad. Sci. USA 85, 1184–118 (1988).
Shi, L., Kam, C.M., Powers, J.C., Aebersold, R. & Greenberg, A.H. Purification of three cytotoxic lymphocyte granule serine proteases that induce apoptosis through distinct substrate and target cell interactions. J. Exp. Med. 176, 1521–1529 (1992).
Shiver, J.W., Su, L. & Henkart, P.A. Cytotoxicity with target DNA breakdown by rat basophilic leukemia cells expressing both cytolysin and granzyme A. Cell 71, 315–322 (1992).
Beresford, P.J., Xia, Z., Greenberg, A.H. & Lieberman, J. Granzyme A loading induces rapid cytolysis and a novel form of DNA damage independently of caspase activation. Immunity 10, 585–594 (1999).
Beresford, P.J., Kam, C.M., Powers, J.C. & Lieberman, J. Recombinant human granzyme A binds to two putative HLA-associated proteins and cleaves one of them. Proc. Natl. Acad. Sci. USA 94, 9285–9290 (1997).
Fan, Z., Beresford, P.J., Zhang, D. & Lieberman, J. HMG2 interacts with the nucleosome assembly protein SET and is a target of the cytotoxic T-lymphocyte protease granzyme A. Mol. Cell Biol. 22, 2810–2820 (2002).
Fan, Z. et al. Cleaving the oxidative repair protein Ape1 enhances cell death mediated by granzyme A. Nat. Immunol. 4, 145–153 (2003).
Pinkoski, M.J. & Green, D.R. Granzyme A: the road less traveled. Nat. Immunol. 4, 106–108 (2003).
Chakravarti, D. & Hong, R. SET-ting the stage for life and death. Cell 112, 589–591 (2003).
Fan, Z., Beresford, P.J., Oh, D.Y., Zhang, D. & Lieberman, J. Tumor suppressor NM23-H1 is a granzyme A–activated DNase during CTL-mediated apoptosis, and the nucleosome assembly protein SET is its inhibitor. Cell 112, 659–672 (2003).
Beresford, P.J. et al. Granzyme A activates an endoplasmic reticulum–associated caspase-independent nuclease to induce single-stranded DNA nicks. J. Biol. Chem. 276, 43285–43293 (2001).
Zhang, D., Beresford, P.J., Greenberg, A.H. & Lieberman, J. Granzymes A and B directly cleave lamins and disrupt the nuclear lamina during granule-mediated cytolysis. Proc. Natl. Acad. Sci. USA 98, 5746–5751 (2001).
Zhang, D. et al. Induction of rapid histone degradation by the cytotoxic T lymphocyte protease granzyme A. J. Biol. Chem. 276, 3683–3690 (2001).
Simon, M.M., Fruth, U., Simon, H.G., Gay, S. & Kramer, M.D. Evidence for multiple functions of T-lymphocytes associated serine proteinases. Adv. Exp. Med. Biol. 247A, 609–613 (1989).
Kam, C.M., Hudig, D. & Powers, J.C. Granzymes (lymphocyte serine proteases): characterization with natural and synthetic substrates and inhibitors. Biochim. Biophys. Acta 1477, 307–323 (2000).
Wilharm, E. et al. Generation of catalytically active granzyme K from Escherichia coli inclusion bodies and identification of efficient granzyme K inhibitors in human plasma. J. Biol. Chem. 274, 27331–27337 (1999).
Hink-Schauer, C. et al. The 2.2-Å crystal structure of human pro-granzyme K reveals a rigid zymogen with unusual features. J. Biol. Chem. 277, 50923–50933 (2002).
Jackson, D.S. et al. Synthesis and evaluation of diphenyl phosphonate esters as inhibitors of the trypsin-like granzymes A and K and mast cell tryptase. J. Med. Chem. 41, 2289–2301 (1998).
Bode, W. et al. X-ray crystal structure of the complex of human leukocyte elastase (PMN elastase) and the third domain of the turkey ovomucoid inhibitor. EMBO J. 5, 2453–2458 (1986).
Chandrasekharan, U.M., Sanker, S., Glynias, M.J., Karnik, S.S. & Husain, A. Angiotensin II-forming activity in a reconstructed ancestral chymase. Science 271, 502–505 (1996).
Parry, M.A. et al. The ternary microplasmin-staphylokinase-microplasmin complex is a proteinase–cofactor–substrate complex in action. Nat. Struct. Biol. 5, 917–923 (1998).
Parry, M.A., Zhang, X.C. & Bode, I. Molecular mechanisms of plasminogen activation: bacterial cofactors provide clues. Trends Biochem. Sci. 25, 53–59 (2000).
Bell, J.K. et al. The oligomeric structure of human granzyme A is a determinant of its extended substrate specificity. Nat. Struct. Biol. 10, 527–534 (2003).
Navaza, J. Implementation of molecular replacement in AMoRe. Acta Cryst. 57, 1367–1372 (2001).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D. Biol. Crystallogr. 54 (Pt 5), 905–921 (1998).
Collaborative Computational Project 4. The CCP4 suite: programs for protein crystallography. Acta Cryst. D 50, 760–763 (1994).
Laskowski, R., MacArthur, M., Hutchinson, E. & Thorton, J.J. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993).
Estébanez-Perpiná, E. et al. Crystal structure of the caspase activator human granzyme B, a proteinase highly specific for an Asp-P1 residue. Biol. Chem. 381, 1203–1214 (2000).
Nicholls, A., Bharadwaj, R. & Honig, B. GRASP - graphical representation and analysis of surface-properties. Biophys. J. 64, A166 (2003).
Acknowledgements
We thank R. Friedrich, P. Fuentes-Prior, S. Steinbacher and W. Klinkert for helpful discussions and H. Wekerle and R. Huber for their continuous interest in the project. Recombinant SET was provided by J. Lieberman. This work was supported by grants of the German Research Council and the European Union.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Hink-Schauer, C., Estébanez-Perpiñá, E., Kurschus, F. et al. Crystal structure of the apoptosis-inducing human granzyme A dimer. Nat Struct Mol Biol 10, 535–540 (2003). https://doi.org/10.1038/nsb945
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb945
This article is cited by
-
Identification of pyroptosis-related genes and long non-coding RNAs signatures in osteosarcoma
Cancer Cell International (2022)
-
Direct Cytosolic Delivery of Proteins Using Lyophilized and Reconstituted Polymer-Protein Assemblies
Pharmaceutical Research (2022)
-
Identification and annotation of bovine granzyme genes reveals a novel granzyme encoded within the trypsin-like locus
Immunogenetics (2018)
-
Lassa and Marburg viruses elicit distinct host transcriptional responses early after infection
BMC Genomics (2014)
-
Granzyme M: characterization with sites of post-translational modification and specific sites of interaction with substrates and inhibitors
Molecular Biology Reports (2011)