Abstract
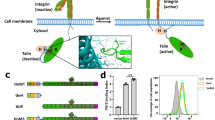
PINCH is an adaptor protein found in focal adhesions, large cellular complexes that link extracellular matrix to the actin cytoskeleton. PINCH, which contains an array of five LIM domains, has been implicated as a platform for multiple protein–protein interactions that mediate integrin signaling within focal adhesions. We had previously characterized the LIM1 domain of PINCH, which functions in focal adhesions by binding specifically to integrin-linked kinase. Using NMR spectroscopy, we show here that the PINCH LIM4 domain, while maintaining the conserved LIM scaffold, recognizes the third SH3 domain of another adaptor protein, Nck2 (also called Nckβ or Grb4), in a manner distinct from that of the LIM1 domain. Point mutation of LIM residues in the SH3-binding interface disrupted LIM–SH3 interaction and substantially impaired localization of PINCH to focal adhesions. These data provide novel structural insight into LIM domain–mediated protein–protein recognition and demonstrate that the PINCH-Nck2 interaction is an important component of the focal adhesion assembly during integrin signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Martin, K.H., Slack, J.K., Boerner, S.A., Martin, C.C. & Parsons, J.T. Integrin connections map: to infinity and beyond. Science 296, 1652–1653 (2002).
Hynes, R. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Rearden, A. A new LIM protein containing an autoepitope homologous to 'senescent cell antigen'. Biochem. Biophys. Res. Commun. 201, 1124–1131 (1994).
Tu, Y., Li, F., Goicoechea, S. & Wu, C. The LIM-only protein PINCH directly interacts with integrin-linked kinase and is recruited to integrin-rich sites in spreading cells. Mol. Cell. Biol. 19, 2425–2434 (1999).
Wu, C. & Dedhar, S. Integrin-linked kinase (ILK) and its interactors: a new paradigm for the coupling of extracellular matrix to actin cytoskeleton and signaling complexes. J. Cell Biol. 155, 505–510 (2001).
Hobert, O., Moerman, D.G., Clark, K.A., Beckerle, M.C. & Ruvkun, G. A conserved LIM protein that affects muscular adherens junction integrity and mechanosensory function in Caenorhabditis elegans. J. Cell Biol. 144, 45–57 (1999).
Clark, K.A., McGrall, M. & Beckerle, M.C. Analysis of PINCH function in Drosophila demonstrates its requirement in integrin-dependent cellular processes. Development 130, 2611–2621 (2003).
Tu, Y., Li, F. & Wu, C. Nck-2, a novel Src homology2/3-containing adaptor protein that interacts with the LIM-only protein PINCH and components of growth factor receptor kinase-signaling pathways. Mol. Biol. Cell 9, 3367–3382 (1998).
Li, F., Zhang, Y. & Wu, C. Integrin-linked kinase is localized to cell-matrix focal adhesions but not cell-cell adhesion sites and the focal adhesion localization of integrin-linked kinase is regulated by the PINCH-binding ANK repeats. J. Cell Sci. 112, 4589–4599 (1999).
Velyvis, A., Yang, Y., Wu, C. & Qin, J. Solution structure of the focal adhesion adaptor PINCH LIM1 domain and characterization of its interaction with the integrin-linked kinase ankyrin repeat domain. J. Biol. Chem. 276, 4932–4939 (2001).
Bax, A. & Grzesiek, S. Methodological advances in protein NMR. Acc. Chem. Res. 26, 131–138 (1993).
Clore, G.M. & Gronenborn, A.M. Determining the structures of large proteins and protein complexes by NMR. Trends Biotechnol. 16, 22–34 (1998).
Perez-Alvarado, G.C. et al. Structure of the carboxy-terminal LIM domain from the cysteine rich protein CRP. Nat. Struct. Biol. 1, 388–398 (1994).
Perez-Alvarado, G.C. et al. Structure of the cysteine-rich intestinal protein, CRIP. J. Mol. Biol. 257, 153–174 (1996).
Konrat, R., Krautler, B., Weiskirchen, R. & Bister, K. Structure of cysteine- and glycine-rich protein CRP2. Backbone dynamics reveal motional freedom and independent spatial orientation of the LIM domains. J. Biol. Chem. 273, 23233–23240 (1998).
Yao, X. et al. Solution structure of the chicken cysteine-rich protein, CRP1, a double-LIM protein implicated in muscle differentiation. Biochemistry 38, 5701–5713 (1999).
Spera, S. & Bax, A. Correlations of Cα/β chemical shifts to the protein secondary structure. J. Am. Chem. Soc. 113, 5490–5492 (1991).
Wüthrich, K. NMR of Proteins and Nucleic Acids (John Wiley & Sons, New York; 1986).
Gorelick, R.J. et al. Strict conservation of the retroviral nucleocapsid protein zinc finger is strongly influenced by its role in viral infection processes: characterization of HIV-1 particles containing mutant nucleocapsid zinc-coordinating sequences. Virology 256, 92–104 (1999).
Guo, J. et al. Subtle alterations of the native zinc finger structures have dramatic effects on the nucleic acid chaperone activity of human immunodeficiency virus type 1 nucleocapsid protein. J. Virol. 76, 4370–4378 (2002).
Michelsen, J.W. et al. Mutational analysis of the metal sites in an LIM domain. J. Biol. Chem. 269,11108–11113 (1994).
Schuler, W., Kloiber, K., Matt, T., Bister, K. & Konrat, R. Application of cross- correlated NMR spin relaxation to the zinc-finger protein CRP2(LIM2): evidence for collective motions in LIM domains. Biochemistry 40, 9596–9604 (2001).
Qin, J., Vinogradova, O. & Gronenborn, A. Protein-protein interactions probed by NMR spectroscopy. Methods Enzymol. 339, 377–389 (2001).
Vinogradova, O. et al. A structural mechanism of integrin αIIbβ3 'inside-out' activation as regulated by its cytoplasmic face. Cell 110, 587–597 (2002).
Cavanagh, J., Fairbrother, W.J., Palmer, A.G. & Skelton, N.J. Protein NMR Spectroscopy: Principles and Practice (Academic Press, San Diego, 1996).
Li, S.C. et al. Structure of a Numb PTB domain-peptide complex suggests a basis for diverse binding specificity. Nat. Struct. Biol. 5, 1075–1083 (1998).
Zwahlen, C, Li, S.C., Kay, L.E., Pawson, T. & Forman-Kay, J.D. Multiple modes of peptide recognition by the PTB domain of the cell fate determinant Numb. EMBO J. 19, 1505–1515 (2000).
Dhalluin, C. et al. Structural basis of SNT PTB domain interactions with distinct neurotrophic receptors. Mol. Cell 6, 921–929 (2000).
Mallis, R.J., Brazin, K.N., Fulton, D.B. & Andreotti, A.H. Structural characterization of a proline-driven conformational switch within the Itk SH2 domain. Nat. Struct. Biol. 9, 900–905 (2002).
Sadler, I., Crawford, A.W., Michelsen, J.W. & Beckerle, M.C. Zyxin and cCRP: two interactive LIM domain proteins associated with the cytoskeleton. J. Cell. Biol. 119, 1573–1587 (1992).
Schmeichel, K.L. & Beckerle, M.C. Molecular dissection of a LIM domain. Mol. Biol. Cell 8, 219–230 (1997).
Wu R, et al. Specificity of LIM domain interactions with receptor tyrosine kinases. J. Biol. Chem. 271, 15934–15941 (1996).
Zhang, Y. et al. Assembly of the PINCH-ILK-CH-ILKBP complex precedes and is essential for localization of each component to cell-matrix adhesion sites. J. Cell Sci. 115, 4777–4786 (2002).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Garrett, D.S., Powers, R., Gronenborn, A.M. & Clore, G.M. A common-sense approach to peak picking in two-, three-, and four-dimensional spectra using automatic computer analysis of contour diagrams. J. Magn. Reson. 95, 214–220 (1991).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Laskowski, R.A., Rullmann, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association — insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
Zhang, Y., Guo, L., Chen, K. & Wu, C. A critical role of the PINCH-integrin-linked kinase interaction in the regulation of cell shape change and migration. J. Biol. Chem. 277, 318–326 (2002).
Acknowledgements
We thank F. Delaglio for NMRPipe software and D. Garrett for PIPP. The work was supported by US National Institutes of Health grants to J.Q. and C.W.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Velyvis, A., Vaynberg, J., Yang, Y. et al. Structural and functional insights into PINCH LIM4 domain–mediated integrin signaling. Nat Struct Mol Biol 10, 558–564 (2003). https://doi.org/10.1038/nsb938
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb938