Abstract
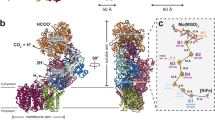
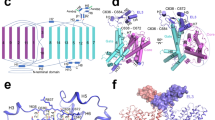
The major facilitator superfamily (MFS) represents one of the largest classes of evolutionarily related membrane transporter proteins. Here we present the three-dimensional structure at 6.5 Å resolution of a bacterial member of this superfamily, OxlT. The structure, derived from an electron crystallographic analysis of two-dimensional crystals, reveals that the 12 helices in the OxlT molecule are arranged around a central cavity, which is widest at the center of the membrane. The helices divide naturally into three groups: a peripheral set comprising helices 3, 6, 9 and 12; a second set comprising helices 2, 5, 8 and 11 that faces the central substrate transport pathway across most of the length of the membrane; and a third set comprising helices 1, 4, 7 and 10 that participate in the pathway either on the cytoplasmic side (4 and 10) or on the periplasmic side (1 and 7). Overall, the architecture of the protein is remarkably symmetric, providing a compelling molecular explanation for the ability of such transporters to carry out bi-directional substrate transport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Paulsen, I.T., Sliwinski, M.K. & Saier, M.H. Jr J. Mol. Biol. 277, 573–592 (1998).
Doyle, D.A. et al. Science 280, 69–77 (1998).
Weiss, M.S. et al. Science 254, 1627–1630 (1991).
Murata, K. et al. Nature 407, 599–605 (2000).
Fu, D. et al. Science 290, 481–486 (2000).
Henderson, R. et al. J. Mol. Biol. 213, 899–929 (1990).
Chang, G. & Roth, C.B. Science 293, 1793–1800 (2001).
Anantharam, V., Allison, M.J. & Maloney, P.C. J. Biol. Chem. 264, 7244–7250 (1989).
Heymann, J.A. et al. EMBO J. 20, 4408–4413 (2001).
Subramaniam, S. et al. J. Mol. Biol. 287, 145–161 (1999).
Dubochet, J. et al. Q. Rev. Biophys. 21, 129–228 (1988).
Crowther, R.A., Henderson, R. & Smith, J.M. J. Struct. Biol. 116, 9–16 (1996).
Henderson, P.J. & Maiden, M.C. Phil. Trans. R. Soc. Lond. B Biol. Sci. 326, 391–410 (1990).
Williams, K.A. Nature 403, 112–115 (2000).
Goswitz, V.C. & Brooker, R.J. Protein Sci. 4, 534–537 (1995).
Frillingos, S., Sahin-Toth, M., Wu, J. & Kaback, H.R. FASEB J. 12, 1281–1299 (1998).
Tamura, N. et al. J. Biol. Chem. 276, 20330–20339 (2001).
Amos, L.A., Henderson, R. & Unwin, P.N. Prog. Biophys. Mol. Biol. 39, 183–231 (1982).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta. Crystallogr. A 47, 110–119 (1991).
Kleywegt, G.J. & Jones, T.A. Structure 4, 1395–1400 (1996).
Acknowledgements
We thank L. Ye for generous assistance with purification of OxIT and D. Bliss for assistance with preparation of figures. This work was supported by grants to S.S. from the intramural program at the National Institutes of Health and to P.C.M. from the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Hirai, T., Heymann, J., Shi, D. et al. Three-dimensional structure of a bacterial oxalate transporter. Nat Struct Mol Biol 9, 597–600 (2002). https://doi.org/10.1038/nsb821
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb821
This article is cited by
-
Plant glucose transporter structure and function
Pflügers Archiv - European Journal of Physiology (2020)
-
Genome mining and homologous comparison strategy for digging exporters contributing self-resistance in natamycin-producing Streptomyces strains
Applied Microbiology and Biotechnology (2020)
-
pH-induced structural change in a sodium/proton antiporter from Methanococcus jannaschii
The EMBO Journal (2005)