Abstract
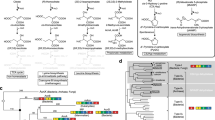
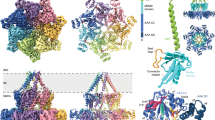
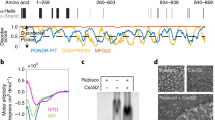
The major bifunctional aconitase of Escherichia coli (AcnB) serves as either an enzymic catalyst or a mRNA-binding post-transcriptional regulator, depending on the status of its iron–sulfur cluster. AcnB represents a large, distinct group of Gram-negative bacterial aconitases that have an altered domain organization relative to mitochondrial aconitase and other aconitases. Here the 2.4 Å structure of E. coli AcnB reveals a high degree of conservation at the active site despite its domain reorganization. It also reveals that the additional domain, characteristic of the AcnB subfamily, is a HEAT-like domain, implying a role in protein–protein recognition. This domain packs against the remainder of the protein to form a tunnel leading to the aconitase active site, potentially for substrate channeling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Robbins, A.H. & Stout, C.D. Proteins Struct. Funct. Genet. 5, 289–312 (1989).
Gruer, M.J., Artymiuk, P.J. & Guest, J.R. Trends Biochem. Sci. 22, 3–6 (1997).
Bradbury, A.J., Gruer, M.J., Rudd, K.E. & Guest, J.R. Microbiology 142, 389–400 (1996).
Cunningham, L., Gruer, M.J. & Guest, J.R. Microbiology 143, 3795–3805 (1997).
Beinert, H. & Kennedy, M.C. FASEB J. 7, 1442–1449 (1993).
Hentze, M.M. & Kühn, L.C. Proc. Natl. Acad. Sci. USA 93, 8175–8182 (1996).
Alén, C. & Sonenshein, A.L. Proc. Natl. Acad. Sci. USA 96, 10412–10417 (1999).
Tang, Y. & Guest, J.R. Microbiology 145, 3069–3079 (1999).
Tang, Y., Quail, M.A., Artymiuk, P.J., Guest, J.R. & Green, J. Microbiology 148, 1027–1037 (2002).
Butt, J. et al. Proc. Natl. Acad. Sci. USA 93, 4345–4349 (1996).
Todd, A.E., Orengo, C.A. & Thornton, J.M. J. Mol. Biol. 307, 1113–1143 (2001).
Polekhina, G., Board, P.G., Gali, R.R., Rossjohn, J. & Parker, M.W. EMBO J. 18, 3204–3213 (1999).
Lauble, H., Kennedy, M.C., Beinert, H. & Stout, C.D. Biochemistry 31, 2735–2748 (1992).
Lauble, H., Kennedy, M.C., Beinert, H. & Stout, C.D. J. Mol. Biol. 237, 437–451 (1994).
Beinert, H., Kennedy, M.C. & Stout, C.D. Chem. Rev. 96, 2335–2373 (1996).
Grindley, H.M., Artymiuk, P.J., Rice, D.W. & Willett, P. J. Mol. Biol. 229, 707–721 (1993).
Groves, M.R., Hanlon, N., Turowski, P., Hemmings, B.A. & Barford, D. Cell 96, 99–110 (1999).
Conti, E., Uy, M., Leighton, L., Blobel, G. & Kuriyan, J. Cell 94, 193–204 (1998).
Andrade, M.A., Petosa, C., O'Donoghue, S.I., Muller, C.W. & Bork, P. J. Mol. Biol. 309, 1–18 (2001).
Edwards, T.A., Pyle, S.E., Wharton, R.P. & Aggarwal, A.K. cell 105, 281–289 (2001).
Wang, X.Q., Zamore, P.D. & Hall, T.M.T. Mol. Cell 7, 855–865 (2001).
Wharton, R.P., Sonoda, J., Lee, T., Patterson, M. & Murata, Y. Mol. Cell 1, 863–872 (1998).
Basilion, J.P., Rouault, T.A., Massinople, C.M., Klausner, R.D. & Burgess, W.H. Proc. Natl. Acad. Sci. USA 91, 574–578 (1994).
Addess, K.J., Basilion, J.P., Klausner, R.D., Rouault, T.A. & Pardi, A. J. Mol. Biol. 274, 72–83 (1997).
Kaldy, P., Menotti, E., Moret, R. & Kühn, L.C. EMBO J. 18, 6073–6083 (1999).
Ruediger, R. et al. Mol. Cell. Biol. 12, 4872–4882 (1992).
Jordan, P.A., Tang, Y., Bradbury, A., Thomson, A.J. & Guest, J.R. Biochem. J. 344, 739–746 (1999).
Hyde, C.C., Ahmed, S.A., Padlan, E.A., Miles, E.W. & Davies, D.R. J. Biol. Chem. 263, 17857–17871 (1988).
Thoden, J.B., Holden, H.M., Wesenberg, G., Raushel, F.M. & Rayment, I. Biochemistry 36, 6305–6316 (1997).
Srere, P.A. Trends Biochem. Sci. 10, 109–110 (1985).
Srere, P.A. Annu. Rev. Biochem. 56, 89–124 (1987).
Barnes, S.J. & Weitzman, P.D.J. FEBS Lett. 201, 267–271 (1986).
Nguyen, N.T. et al. Biochemistry 40, 13177–13187 (2001).
Hurley, J.H. et al. Proc. Natl. Acad. Sci. USA 86, 8635–8639 (1989).
Gruer, M.J., Bradbury, A.J. & Guest, J.R. Microbiology 143, 1837–1846 (1997).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Bailey, S. Acta Crystallogr. D 50, 760–763 (1994).
Cowtan, K. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994).
Roussel, A. & Cambillau, C. Silicon Graphics Geometry Partners Directory 86, 86 (1991).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Acta Crystallogr. D 53, 240–255 (1997).
Brunger, A.T. Acta Crystallogr. D 49, 24–36 (1993).
Ferrin, T.E., Huang, C.C., Jarvis, L.E. & Langridge, R. J. Mol. Graph. 6, 13–27 (1988).
Barton, G.L. Protein Eng. 6, 13–27 (1993).
Esnouf, R.M. J. Mol. Graph. 15, 132–134 (1997).
Nicholls, A., Bharadwaj, R. & Honig, B. Biophys. J. 64, 166 (1993).
Acknowledgements
We thank BBSRC, EPSRC and the Wellcome Trust for their support, CCLRC Daresbury for synchrotron radiation facilities, and the Royal Society and Wolfson Foundation for computing facilities. The Krebs Institute is a BBSRC-designated Biomolecular Sciences center and a member of the North of England Structural Biology Centre.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Williams, C., Stillman, T., Barynin, V. et al. E. coli aconitase B structure reveals a HEAT-like domain with implications for protein–protein recognition. Nat Struct Mol Biol 9, 447–452 (2002). https://doi.org/10.1038/nsb801
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb801
This article is cited by
-
Kinetic compartmentalization by unnatural reaction for itaconate production
Nature Communications (2022)
-
Maturation strategy influences expression levels and cofactor occupancy in Fe–S proteins
JBIC Journal of Biological Inorganic Chemistry (2022)
-
Extensive regulation of enzyme activity by phosphorylation in Escherichia coli
Nature Communications (2021)
-
Crystal structures of aconitase X enzymes from bacteria and archaea provide insights into the molecular evolution of the aconitase superfamily
Communications Biology (2021)
-
Selection and validation of reference genes for RT-qPCR indicates that juice of sugarcane varieties modulate the expression of C metabolism genes in the endophytic diazotrophic Herbaspirillum rubrisubalbicans strain HCC103
Antonie van Leeuwenhoek (2017)