Abstract
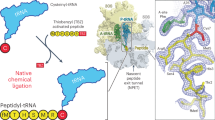
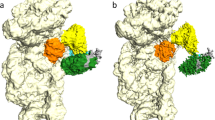
The large ribosomal subunit catalyzes peptide bond formation during protein synthesis. Its peptidyl transferase activity has often been studied using a 'fragment assay' that depends on high concentrations of methanol or ethanol. Here we describe a version of this assay that does not require alcohol and use it to show, both crystallographically and biochemically, that crystals of the large ribosomal subunits from Haloarcula marismortui are enzymatically active. Addition of these crystals to solutions containing substrates results in formation of products, which ceases when crystals are removed. When substrates are diffused into large subunit crystals, the subsequent structure shows that products have formed. The CC-puromycin-peptide product is found bound to the A-site and the deacylated CCA is bound to the P-site, with its 3′ OH near N3 A2486 (Escherichia coli A2451). Thus, this structure represents a state that occurs after peptide bond formation but before the hybrid state of protein synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Monro, R.E. Catalysis of peptide bond formation by 50S ribosomal subunits from Escherchia coli. J. Mol. Biol. 26, 147–151 (1967).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289, 905–920 (2000).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Polacek, N., Gaynor, M., Yassin, A. & Mankin, A.S. Ribosomal peptidyl transferase can withstand mutations at the putative catalytic nucleotide. Nature 411, 498–501 (2001).
Xiong, L., Polacek, H., Sander, P., Boettger, E.G. & Mankin, A.S. pKa of adenine 2451 in the ribosomal peptidyl transferase center remains elusive. RNA 7, 1365–1369 (2001).
Bayfield, M.A., Dahlberg, A.E., Schulmeister, U., Dorner, S., & Barta, A. A conformational change in the ribosomal peptidyl transferase center upon active/inactive transition. Proc. Natl. Acad. Sci. USA 98, 10096–10101 (2001).
Thompson, J. et al. Analysis of mutations at residues A2451 and G2447 of 23S rRNA in the peptidyltransferase active site of the 50S ribosomal subunit. Proc. Natl. Acad. Sci. USA 98, 9002–9007 (2001).
Monro, R.E. & Marker, K.A. Ribosome-catalysed reaction of puromycin with a formylmethionine-containing oligonucleotide. J. Mol. Biol. 25, 347–350 (1967).
Traut, R.R. & Monro, R.E. The puromycin reaction and its relation to protein synthesis. J. Mol. Biol. 10, 63–71 (1964).
Quiggle, K. & Chladek, S. The role of the cytidine residues of the tRNA 3′-terminus at the peptidyltransferase A- and P-sites. FEBS Lett. 118, 172–175 (1980).
Maden, B.E. & Monro, R.E. Ribosome-catalyzed peptidyl transfer. Effects of cations and pH value. Eur. J. Biochem. 6, 309–316 (1968).
Fernandez-Munoz, R., Monro, R.E., Torres-Pinedo, R. & Vazquez, D. Substrate- and antibiotic-binding sites at the peptidyl-transferase centre of Escherichia coli ribosomes. Studies on the chloramphenicol, lincomycin and erythromycin sites. Eur. J. Biochem. 23, 185–193 (1971).
Pestka, S. Studies on the formation of transfer ribonucleic acid–ribosome complexes. Phenylalanyl-oligonucleotide binding to ribosomes and the mechanism of chloramphenicol action. Biochem. Biophys. Res. Commun. 36, 589–595 (1969).
Dosher, M. S. & Richards, F. M. The activity of an enzyme in the crystalline state: ribonuclease S*. J. Biol. Chem. 238, 2399–2406 (1963).
Ban, N. et al. A 9 Å resolution X-ray crystallographic map of the large ribosomal subunit. Cell 93, 1105–1115 (1998).
Bashan, A. et al. in Cold Spring Harbor symp. on quantitative biology 66, (Cold Spring Harbor Press, New York, in the press).
Marin, I., Sanz, J.L., Sanchez, M.E. & Amils, R. Archaea, A laboratory manual, Halophiles (eds Robb, F.T. et al.) (Cold Spring Harbor Press, New York; 1995).
Moazed, D. & Noller, H.F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Wilson, K.S. & Noller, H.F. Molecular movement inside the translational engine. Cell 92, 337–349 (1998).
Borowski, C., Rodnina, M.V. & Wintermeyer W. Truncated elongation factor G lacking the G domain promotes translocation of the 3′ end but not the anticodon domain of peptidyl–tRNA. Proc. Natl. Acad. Sci. USA 93, 4202–4206 (1996).
Green, R., Switzer, C. & Noller, H.F. Ribosome-catalyzed peptide-bond formation with an A-site substrate covalently linked to 23S ribosomal RNA. Science 280, 286–290 (1998).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T. A., Zou, J. A., Cowan, S. & Kjeldgaard, M. Improved method for binding protein models in electron density maps and the locations of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Yusupov, M. M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001).
Acknowledgements
We thank F.M. Richards, A.K. Oylere, D. Kitchen and S.P. Ryder for advice concerning experimental design and execution; D.J. Klein, A. Ray and A.A. Szewczak for helpful discussions; S. Kamtekar for critical reading of the manuscript; and J. Wang and A.R. Curran for assistance with computation. We are indebted to A. Joachimiak and the staff of 19-ID at the Advanced Photon Source (Argonne National Laboratory). This research was supported by NIH grants to T.A.S., P.B.M. and S.A.S., and an Agouron Institute grant to T.A.S. and P.B.M.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Schmeing, T., Seila, A., Hansen, J. et al. A pre-translocational intermediate in protein synthesis observed in crystals of enzymatically active 50S subunits. Nat Struct Mol Biol 9, 225–230 (2002). https://doi.org/10.1038/nsb758
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb758