Abstract
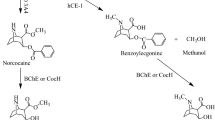
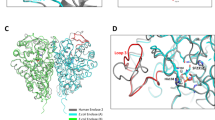
Here we report the first structure of a cocaine-degrading enzyme. The bacterial esterase, cocE, hydrolyzes pharmacologically active (−)-cocaine to a nonpsychoactive metabolite with a rate faster than any other reported cocaine esterase (kcat = 7.8 s−1 and KM = 640 nM). Because of the high catalytic proficiency of cocE, it is an attractive candidate for novel protein-based therapies for cocaine overdose. The crystal structure of cocE, solved by multiple anomalous dispersion (MAD) methods, reveals that cocE is a serine esterase composed of three domains: (i) a canonical α/β hydrolase fold (ii) an α-helical domain that caps the active site and (iii) a jelly-roll-like β-domain that interacts extensively with the other two domains. The active site was identified within the interface of all three domains by analysis of the crystal structures of transition state analog adduct and product complexes, which were refined at 1.58 Å and 1.63 Å resolution, respectively. These structural studies suggest that substrate recognition arises partly from interactions between the benzoyl moiety of cocaine and a highly evolved specificity pocket.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bresler, M.M., Rosser, S.J., Basran, A. & Bruce, N.C. Appl. Environ. Microbiol. 66, 904–908 (2000).
Inaba, T., Stewart, D.J. & Kalow, W. Clin. Pharmacol. Ther. 23, 547–552 (1978).
Brzezinski, M.R., Abraham, T.L., Stone, C.L., Dean, R.A. & Bosron, W.F. Biochem. Pharmacol. 48, 1747–1755 (1994).
Pindel, E.V. et al. J. Biol. Chem. 272, 14769–14775 (1997).
Britt, A.J., Bruce, N.C. & Lowe, C.R. J. Bacteriol. 174, 2087–2094 (1992).
Carroll, F.L., Howell, L.L. & Kuhar, M.J. J. Med. Chem. 42, 2721–2736 (1999).
Carrera, M.R.A. et al. Nature 378, 727–730 (1995).
Mets, B. et al. Proc. Natl. Acad. Sci. USA 95, 10176–10181 (1998).
Landry, D.W., Zhao, K., Yang, G.X.Q., Glickman, M. & Georgiadis, T.M. Science 259, 1899–1901 (1993).
Lu, G. DOMID http://bioinfol.mbfys.lu.se/Domid/ (1999).
Ollis, D.L. et al. Protein Eng. 5, 197–211 (1992).
Nardini, M. & Dijkstra, B.W. Curr. Opin. Struct. Biol. 9, 732–737 (1999).
Heikinheimo, P., Goldman, A., Jeffries, C. & Ollis, D.L. Structure 7, R141–R146 (1999).
Connolly, M.L. J. Appl. Crystallogr. 16, 548–558 (1983).
Sheriff, S., Hendrickson, W.A. & Smith, J.L. J. Mol. Biol. 197, 273–296 (1987).
Derewenda, Z.S. & Wei, Y. J. Am. Chem. Soc. 117, 2104–2105 (1995).
Holm, L. & Sander, C. J. Mol. Biol. 233, 123–138 (1993).
Wei, Y. et al. Structure 6, 511–519 (1997).
Fulop, V., Bocskei Z., & Polgar, L. Cell 94, 161–170 (1998).
Egloff, M.P. et al. Biochemistry 34, 2751–2762 (1995).
Wirsching, P., Ashley, J.A., Lo, C.-H.L., Janda, K.D. & Lerner, R.A. Science 270, 1775–1782 (1995).
Melton, R.G. & Sherwood, R.F. J. Natl. Cancer Inst. 88, 153–165 (1996).
Abuchowski, A., Davis, F. & Davis, S. Cancer Treat. Rep. 65, 1077–1081 (1981).
Xie, W. et al. Mol. Pharmacol. 55, 83–91 (1999).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Terwilliger, T.C. & Berendzen, J. Acta Crystallogr. D 53, 571–579 (1997).
Terwilliger, T.C. Acta Crystallogr. D 56, 965–972 (2000).
Cowtan, K.D. & Main, P. Acta Crystallogr. D 52, 43–48 (1996).
Lamzin, V.S. & Wilson, K.S. Acta Crystallogr. D 49, 129–149 (1993).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Sheldrick, G.M. & Schneider, T.R. Methods Enzymol. 277B, 319–343 (1997).
Esnouf, R.M. Acta Crystallogr. D 55, 938–940 (1999).
Merritt, E.A. & Murphy, M.E.P. Acta Crystallogr. D 50, 869–873 (1994).
Read, R.J. Acta Crystallogr. A 42, 140–149 (1986).
Acknowledgements
The authors thank the Advanced Light Source at Berkeley and the Stanford Synchrotron Radiation Laboratory (SSRL), in particular A. Gonzalez, for advice on MAD data collection. We are also indebted to R. Stanfield for assistance on synchrotron trips and M. Elsliger for helpful discussions. Support was provided by the National Institutes of Health (I.A.W.), the Biotechnology and Biological Sciences Research Council (N.C.B.), the Howard Hughes Medical Institute (J.M.T.), and the Bernie Gilula Foundation (N.A.L.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Larsen, N., Turner, J., Stevens, J. et al. Crystal structure of a bacterial cocaine esterase. Nat Struct Mol Biol 9, 17–21 (2002). https://doi.org/10.1038/nsb742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb742
This article is cited by
-
A thermostable bacterial cocaine esterase rapidly eliminates cocaine from brain in nonhuman primates
Translational Psychiatry (2014)
-
Improving Pseudomonas alcaligenes lipase’s diastereopreference in hydrolysis of diastereomeric mixture of menthyl propionate by site-directed mutagenesis
Biotechnology and Bioprocess Engineering (2014)
-
Long-Lasting Effects of a PEGylated Mutant Cocaine Esterase (CocE) on the Reinforcing and Discriminative Stimulus Effects of Cocaine in Rats
Neuropsychopharmacology (2012)
-
Amelioration of the Cardiovascular Effects of Cocaine in Rhesus Monkeys by a Long-Acting Mutant Form of Cocaine Esterase
Neuropsychopharmacology (2011)
-
A Cocaine Hydrolase Engineered from Human Butyrylcholinesterase Selectively Blocks Cocaine Toxicity and Reinstatement of Drug Seeking in Rats
Neuropsychopharmacology (2008)