Abstract
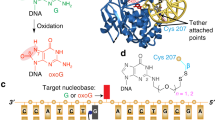
The DNA glycosylase MutY, which is a member of the Helix-hairpin-Helix (HhH) DNA glycosylase superfamily, excises adenine from mispairs with 8-oxoguanine and guanine. High-resolution crystal structures of the MutY catalytic core (cMutY), the complex with bound adenine, and designed mutants reveal the basis for adenine specificity and glycosyl bond cleavage chemistry. The two cMutY helical domains form a positively-charged groove with the adenine-specific pocket at their interface. The Watson-Crick hydrogen bond partners of the bound adenine are substituted by protein atoms, confirming a nucleotide flipping mechanism, and supporting a specific DNA binding orientation by MutY and structurally related DNA glycosylases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lindahl, T. Nature 362, 709–715 ( 1993).
Gilchrest, B. A. & Bohr, V. A. FASEB J. 11, 322–330 (1997).
Beckman, K. B. & Ames, B. N. J. Biol. Chem. 272, 19633–19636 (1997).
Shibutani, S., Takeshita, M. & Grollman, A. P. Nature 349, 431– 434 (1991).
Michaels, M. L. & Miller, J. H. J. Bacteriol. 174, 6321–6325 ( 1992).
Cunningham, R. P. Mutat. Res. 383, 189–196 (1997).
Taddei, F. et al. Science 278, 128–130 (1997).
Nghiem, Y., Cabrera, M., Cupples, C. G., & Miller, J. H. Proc. Natl. Acad. Sci. USA 85, 2709– 2713 (1988).
Lu, R., Nash, H. M., & Verdine, G. L. Curr. Biol. 7, 397– 407 (1997).
Slupska, M.M. et al. J. Bacteriol. 178, 3885– 3892 (1996).
Parikh, S.S., Mol, C.D. & Tainer, J.A. Structure 5, 1543– 1550 (1997).
Michaels, M.L., Pham, L., Nghiem, Y., Cruz, C. & Miller, J.H. Nucleic Acids Res. 18, 3841– 3845 (1990).
Nash, H.M. et al. Curr. Biol. 6, 968– 980 (1996).
Kuo, C.F. et al. Science 258, 434–440 (1994).
Yamagata, Y. et al. Cell 86, 311–319 (1996).
Labahn, J. et al. Cell 86, 321–329 (1996).
Thayer, M.M., Ahern, H., Xing, D., Cunningham, R.P., & Tainer, J.A. EMBO J. 14, 4108– 4120 (1995).
Manuel, R. C. & Lloyd, R. S. Biochemistry 36, 11140–11152 (1997).
Dodson, M. L., Michaels, M. L. & Lloyd, R. S. J. Biol. Chem. 269, 32709– 32712 (1994).
Manuel, R. C., Czerwinski, E. W. & Lloyd, R. S. J. Biol. Chem. 271, 16218– 16226 (1996).
Pelletier, H., Sawaya, M. R., Wolfe, W., Wilson, S. H., & Kraut, J. Biochemistry 35, 12742– 12761 (1996).
Parikh, S.S., Mol, C.D., Slupphaug, G. Bharati, S. Krokan, H.E., & Tainer, J.A. EMBO J. 17, 5214–5226 (1998).
McAuley-Hecht, K.E. et al. Biochemistry 33, 10266– 10270 (1994).
Berdal, K.G., Johansen, R.F. & Seeberg, E. EMBO J. 15, 363– 367 (1998).
Otwinowski, Z. & Minor, W. Meth. Enz. 276, 307–326 (1997).
McRee, D.E. J. Mol. Graphics 10, 44–46 (1992).
Furey, W. & Swaminathan, S. Meth. Enz. 277, 590–629 (1997).
Cowtan, K. Joint CCP4 and ESF-EACBM newsletter on protein crystallography 31, 34–38 (1994).
Read, R.J. Acta Crystallogr. A 42, 140–149 (1986).
Brünger, A.T., Kuriyan, J. & Karplus, M. Science 235, 458– 460 (1987).
Sheldrick, G.M. & Schneider, T.R. Meth. Enz. 277, 319–343 ( 1997).
Acknowledgements
We thank C.D. Putnam for aid with data collection, T.P. Lo, K.E. Morgan, P. Frosst, M. Nelson, and M.L. Dodson for helpful discussions, and M. Pique for help with figures. This work was supported by grants from the National Institutes of Health to J.A.T., R.S.L. and J.H.M., a Distinguished Chair in Environmental Toxicology from The Houston Endowment to R.S.L., a Robert Welch Foundation grant to R.S.L., an NIH Postdoctoral Fellowship grant and an American Cancer Society fellowship to Y.G., a Special Fellowship from the Leukemia Society of America to C.D.M., and a National Science Foundation Graduate Research fellowship to S.S.P.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Guan, Y., Manuel, R., Arvai, A. et al. MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nat Struct Mol Biol 5, 1058–1064 (1998). https://doi.org/10.1038/4168
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/4168
This article is cited by
-
The study of a barley epigenetic regulator, HvDME, in seed development and under drought
BMC Plant Biology (2013)
-
Insights into Chi recognition from the structure of an AddAB-type helicase-nuclease complex
The EMBO Journal (2012)
-
DNA glycosylases: in DNA repair and beyond
Chromosoma (2012)
-
Loss of genes for DNA recombination and repair in the reductive genome evolution of thioautotrophic symbionts of Calyptogena clams
BMC Evolutionary Biology (2011)
-
Evolutionary loss of 8-oxo-G repair components among eukaryotes
Genome Integrity (2010)