Abstract
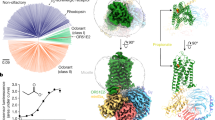
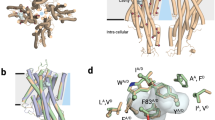
In mammals, odorant binding proteins may play an important role in the transport of odors towards specific olfactory receptors on sensory neurones across the aqueous compartment of the nasal mucus. We have solved the X-ray structure of such a transport protein, bovine odorant binding protein (OBP) at 2.0 Å resolution. The β-barrel of OBP is similar to that of lipocalins, but OBP dimer association results from domain swapping, an observation unique among the lipocalins. The α-helix of each monomer stacks against the β-barrel of the other monomer. Contrary to previous reports, each monomer has an internal buried cavity which could accommodate a naturally occurring molecule. Besides this cavity, an open cavity is located at the dimer interface. Data in solution suggest that this central cavity may be a binding site created by domain swapping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Buck, L. & Axel, R. A novel multigene family may encode odorant receptors: a molecular basis for odor recognition. Cell 65, 175–187 (1991).
Pelosi, P. Odorant-binding proteins. Crit. Rev. Molec. Biol. 29, 199–228 (1994).
Pevsner, J., Sklar, P.B., Hwang, P.M. & Snyder, S.H. in Chemical Senses vol. 1 (eds J.G. Brand, J.H. Teeter, R.H. Cagan & M.R. Kare) 227–242 (MDI Publishers, 1989).
Pelosi, P, Baldaccini, N.E.. & Pisanelli, A.M. Identification of a specific olfactory receptor for 2-isobutyl-3-methoxypyrazine. Biochem. J. 201, 245–248. (1982)
Bignetti, E. et al. Purification and characterization of an odorant-binding protein from cow nasal tissue. Eur. J. Biochem. 149, 227–231 (1985).
Pevsner, J., Trifiletti, R.R., Strittmatter, S.M. & Snyder, S.H. Isolation and characterization of an olfactory receptor protein for odorant pyrazines. Proc. Natl. Acad. Sci. USA 82, 3050–3054 (1985).
Krieger, J., Ganssel, H., Raming, K. & Breer, H. Odorant binding proteins of Heliothis virescens. Insect Biochem. Molec. Biol. 23, 449–456 (1993).
Cavaggioni, A., Sorbi, R.T., Keen, J.N., Pappin, D.J. & Findlay, J.B. Homology between the pyrazine binding protein from nasal mucosa and major urinary proteins. FEBS Letters 212, 225–228 (1987).
Böcskei, Z. et al. Pheromone binding to two rodent urinary proteins revealed by X-ray crystallography. Nature 360, 186–188 (1992).
Banaszak, L. et al. Lipid-binding proteins: a family of fatty acid and retinoid transport proteins. Advances in Protein Chemistry 45, 89–151 (1994).
Cowan, S.W., Newcomer, M.E. & Jones, T.A. Crystallographic refinement of human serum retinol binding protein at 2 Å resolution. Proteins 8, 44–61 (1990).
Monaco, H.L. & Zanotti, G. Three-dimensional structure and active site of three hydrophobic molecule-binding proteins with significant amino acid sequence similarity. Biopolymers 32, 457–465 (1992).
Bussolati, L., Ramoni, R., Grolli, S., Donofrio, G. & Bignetti, E. Preparation of an affinity resin for odorants by coupling odorant binding protein from bovine nasal mucosa to Sepharose 4B. J. Biotechnol. 30, 225–230 (1993).
Bennett, M.J., Choe, S. & Eisenberg, D. Domain swapping: Entangling alliances between proteins. Proc. Nat. Acad. Sci. USA 91, 3127–3131 (1994).
Bennet, J.M., Schlunegger, M.P. & Eisenberg, D. 3D Domain swapping: a mechanism for oligomer assembly. Prot. Sci. 4, 2455–(Author: page range?) (1995)
Connolly, M.L. Surface-accessible surfaces of proteins and nucleic acids. Science 306, 287–290 (1983).
Singer, A.G. & Macrides, F. Aphrodisin: pheromone or transducer? Chem. Senses 15, 199–203 (1990).
Pevsner, J., Reed, R.R., Feinstein, P.G. & Snyder, S.H. Molecular cloning of odorant-binding protein: member of a ligand carrier family. Science 241, 336–339 (1988).
Dal Monte, M., Andreini, I., Revoltella, R. & Pelosi, P. Purification and characterization of two odorant binding proteins from nasal tissue of rabbit and pig. Comp. Biochem. Physiol. 99B, 445–451 (1991).
Pes, D., Dal Monte, M., Ganni, M. & Pelosi, P. Isolation of two odorant-binding proteins from mouse nasal tissue. Comp. Biochem. Physiol. 103B, 1011–1017 (1992).
Bignetti, E. et al. Crystallization of an odorant-binding protein from cow nasal mucosa. J.Mol.Biol. 186, 211–212 (1985).
Otwinovsky, Z. DENZO: an oscillation data processing program for macromolecular crystallography. Yale University, New Haven, CT, USA (1993).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A50, 157–163 (1994).
Brünger, A.T. X-PLOR version 3.1 Manual. Yale University, New Haven, CT, USA (1993).
Howard, E.I., Grigera, J.R., Grigera, T.S. & Podjarny, A.D. in CCP4 Newsletter, June, 1995 17–19 (1995).
Laskowski, R. MacArthur, M, Moss, D, & Thornton, J. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 91–97 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Tegoni, M., Ramoni, R., Bignetti, E. et al. Domain swapping creates a third putative combining site in bovine odorant binding protein dimer. Nat Struct Mol Biol 3, 863–867 (1996). https://doi.org/10.1038/nsb1096-863
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1096-863
This article is cited by
-
A new non-classical fold of varroa odorant-binding proteins reveals a wide open internal cavity
Scientific Reports (2021)
-
OBPred: feature-fusion-based deep neural network classifier for odorant-binding protein prediction
Neural Computing and Applications (2021)
-
Probe-dependence of competitive fluorescent ligand binding assays to odorant-binding proteins
Analytical and Bioanalytical Chemistry (2020)
-
The olfactory secretome varies according to season in female sheep and goat
BMC Genomics (2019)
-
Buffalo nasal odorant-binding protein (bunOBP) and its structural evaluation with putative pheromones
Scientific Reports (2018)