Abstract
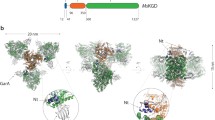
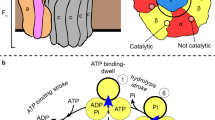
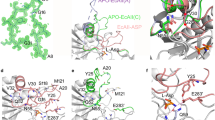
The molecular architecture of the Class II E. coli fructose 1,6-bisphosphate aldolase dimer was determined to 1.6 Å resolution. The subunit fold corresponds to a singly wound α/β-barrel with an active site located on the β-barrel carboxyl side of each subunit. In each subunit there are two mutually exclusive zinc metal ion binding sites, 3.2 Å apart; the exclusivity is mediated by a conformational transition involving side-chain rotations by chelating histidine residues. A binding site for K+ and NH4+ activators was found near the β-barrel centre. Although Class I and Class II aldolases catalyse identical reactions, their active sites do not share common amino acid residues, are structurally dissimilar, and from sequence comparisons appear to be evolutionary distinct.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Horecker, B.L., Tsolas, O. & Lai, C.Y. In The Enzymes 3rd ed. Vol 7 (ed. RD. Boyer) 213–258 (New York: Academic Press, 1972).
Sygusch, J., Beaudry, D. & Allaire, M. Molecular architecture of rabbit skeletal muscle aldolase at 2.7 Å resolution. Proc. Natl. Acad. Sci. USA 84, 7846–7850 (1987).
Gamblin, S.J. et al. Activity and specificity of human aldolases. J. Molec. Biol. 219, 573–576 (1991).
Hester, G. et al. The crystal structure of fructose-1,6-bisphosphate aldolase from Drosophila melanogaster at 2.5 Å resolution. FEBS Lett. 292, 237–242 (1991).
Rutter, W.J. Evolution of aldolase. FASEB. J. 23, 1248–1257 (1964).
Naismith, J.H. et al. Initiating a crystallographic study of a class II fructose-1,6-bisphosphate aldolase, J. Molec. Biol. 225, 1137–1141 (1992).
Bergmeyer, H.U. In Methods of Enzymatic Analysis 4 (eds H.U. Bergmeyer & K. Gewahn) 2272–2274 (Weinheim,Verlag Chemie, 1974).
den Hollander, J.A., Brown, T.R., Ugurbil, K. & Shulman, R.G. 13C nuclear magnetic resonance studues of anaerobic glycolysis in suspension of yeast cells. Proc. Natl. Acad. Sci. USA 76, 6096–6100 (1979).
Lowry, O.H., Carter, J., Ward, J.B. & Galser, L. The effect of carbon and nitorgen sources on the level of metabolic intermediates in Escherichia coli. J. Biol Chem. 246, 6511–6521 (1971).
Ugurbil, K., Brown, T.R., den Hollander, J.A., Glynn, P. & Shulman, R.G. High-resolution 13C nuclear magnetic resonance studies of glucose metabolism in Escherichia coli. Proc. Natl. Acd. Sci. USA 75, 3742–3746 (1978).
Szwergold, B.S., Ugurbil, K. & Brown, T.R. Propeties of fructose-1,6-bisphosphate aldolase from Escherichia coli: An NMR anlysis. Arch. Biochem. Biophys. 317, 224–252 (1995).
Kitagawa, Y., Leonard, G.A., Harrop, S.J., Peterson, M.R. & Hunter, W.N. Additional crystal forms of the E. coli class II fructose-1,6-bisphosphate aldolase. Acta Crystallogr. D51, 833–834 (1995).
International Tables for X-ray Crystallography Vol III, 257–259 (Birmingham:TheKynoch Press (IUCR), 1968).
Brünger, A.T. The free R factor value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–474 (1992).
Richardson, J.S. Describing patterns of protein tertiary structure. Metha. Enzymol. 115, 341–358 (1985).
Ren, J., Stuart, D.I. & Acharya, K.R. α-Lactalbumin possesses a distinct zinc binding site. J. Biol. Chem. 268, 19292–19298 (1993).
Kobes, R.D., Simpson, R.T., Vallee, B.L. & Rutter, W.J. A functional role of metal ions in class II aldolase. Biochemistry 8, 585–588 (1969).
Smith, G.M. & Mildvan, A.S. Nuclear magnetic resonance and chemical modification studies of the role of the metal in yeast aldolase Biochemistry 20, 4340–4346 (1981).
Belasco, J.G. & Knowles, J.R. Polarization of substrate carbonyl groups by yeast aldolase: investigation by Fourier transform infrared spectroscopy Biochemistry 22, 122–129 (1983).
Baldwin, S.A., Perham, R.N. & Stribling, D. Purification and characterization of the class II D-fructose 1,6-bisphosphate aldolase from Escherichia coli (Crookes'strain), Biochem. J. 169, 633–641 (1978).
Sobieszek, A. & Bremel, R.D. Preparation and properties of vertebrate smooth-muscle myofibrils and actomyosin. Eur. J. Biochem. 55, 49–60 (1975).
Takio, K., Blumenthal, D.K., Walsh, K.A., Titani, K. & Krebs, E.G. Amino acid sequence of rabbit skeletal muscle myosin light chain kinase. Biochemistry 25, 8049–8057 (1986).
Qamar, S., March, K. & Berry, A. Identification of arginine 331 as an important active site residue in the class II fructose-1,6-bisphosphate aldolase of Escherichia coli. Prot. Sci. 5, 154–161 (1996).
Vallee, B.L. & Auld, D.S. Zinc coordination, function, and structure of zinc enzymes and other proteins. Biochemistry 29, 5647–5659 (1990).
Chakravarty, S. & Kannan, K.K. Drug-protein interactions. Refined structures of three sulfonamide drug complexes of human carbonic anhydrase I enzyme. J. Molec. Biol. 243, 298–309 (1994).
Teplyakov, A., Wilson, K.S., Orioli, P. & Mangani, S. The high resolution crystal structure of the complex between carboxypeptidase A and L-phenyl lactate. Acta Crystallogr. D49, 534–540 (1993).
Dreyer, M.K. & Schulz, G.E. The spatial structure of the class II L-fuculose-1-phosphate aldolase from Escherichia coli. J. Molec. Biol. 231, 549–553 (1993).
Wilson, D.K., Rudolph, F.B. & Quiocho, F. Atomic structure of adenosine deaminase complex with a transition-state analog: understanding catalysis and immunodeficiency mutations. Science 252, 1278–1284 (1991).
Jabri, E., Carr, M., Hausinger, R. & Karplus, P.A. The crystal structure of urease from Klebsiella aerogenes. Science 268, 998–1004 (1995).
Chothia, C. & Lesk, A.M. The relation between the divergence of sequence and structure in proteins, EMBO J. 5, 823–826 (1986).
Freeze, H & Brock, T.D. Thermostable aldolase from Thermus aquaticus. J. Bact. 101, 541–550 (1970).
Yoshino, M. & Murakami, K. AMP deaminase reaction as a control system of glycolysis in yeast. Role of ammonium ion in the interaction of phosphofructokinase and purivate kinase activity with the adenylate energy charge. J. Biol. Chem. 260, 4729–4732 (1985).
Meury, J. & Kepes, A. The regulation of potassium fluxes for the adjustment and maintenance of potassium levels in Escherichia coli. Eur. J. Biochem. 119, 165–170 (1981).
Alefounder, P.R., Baldwin, S.A., Perham, R.N. & Short, N.J. Cloning, sequence analysis and over-expression of the gene for the class II fructose 1,6-bisphosphate aldolase from Escherichia coli. Biochem. J. 257, 529–534 (1989).
Rondeau, J.M. et al. Novel NADPH-binding domain revealed by the crystal structure of aldolase reductase. Nature 355, 469–472 (1992).
Klein, C. & Schulz, G.E. Structure of cyclodextrin glycosyltransferase refined at 2.0 Å resolution. J. Mol. Biol. 217, 737–750 (1991).
Rao, V., Guan, C. & van Roey, P. Endo-β-N-acetylglucosaminidase H at 1.9 Å resolution: active site geometry and substrate recognition. Structure 3, 449–457 (1995).
Lolis, E., Alber, T., Davenport, R.C., Rose, D., Hartman, F.C. & Petsko, G.A. Structure of Yeast triosephosphate isomerase at 1.9 Å resolution. Biochemistry 29, 6609–6618 (1990).
Farber, G.K. An α/β-barrel full of evolutionary trouble. Curr. Opin. Struct. Biol. 3, 409–412 (1993).
Howard, A.J., Nielsen, C. & Xuong, N.H. Software for diffractometer with multiwire area detector. Meths. Enzymol. 114, 416–452 (1985).
Reeke, Jr. G.N. The ROCKS system of computer programs for macromolecular crystallography. J.Appl. Crystallogr. 17, 125–130 (1984).
Jones, T.A. FRODO: A graphics fitting program for macromolecules. In Computing Crystallography. (ed: Sayre D.) 303–317 (Clarendon Press, Oxford, 1982).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallographic R-factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Brünger, A.T. X-PLOR Manual, Version 3.1 (Yale University Press, Connecticut, USA, 1992).
The SERC (UK) Collaborative Computing Project no. 4, A suite of programs for protein crystallography, Daresbury Laboratories, Warrington WA4 4AD UK(1979).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjelgaard, M. Improved methods for the building of protein models in electron density maps and the location of errors in these. Acta Crystallogr. 47, 110–119 (1991).
Kleywegt, G.J. & Jones, T.A. Efficient rebuilding of protein structures. Acta Crystallogr. D52, 829–832 (1996).
Discover User Guide, version 3.1, (San Diego: Biosym Technologies, 1994).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A., Murphy, M.E.P. Raster3D Version 2.0. A program for photorealistic molecular graphics. Acta Crystallogr. D50, 869–673 (1994).
Drenth, J. in Principles of protein crystallography (ed. C.R. Cantor) 285–288 (New York: Springer Verlag, 1994).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Blom, N., Tétreault, S., Coulombe, R. et al. Novel active site in Escherichia coli fructose 1,6-bisphosphate aldolase. Nat Struct Mol Biol 3, 856–862 (1996). https://doi.org/10.1038/nsb1096-856
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1096-856
This article is cited by
-
Genetic susceptibility of δ-ALAD associated with lead (Pb) intoxication: sources of exposure, preventive measures, and treatment interventions
Environmental Science and Pollution Research (2021)
-
Structure-based prediction and identification of 4-epimerization activity of phosphate sugars in class II aldolases
Scientific Reports (2017)
-
DHAP-dependent aldolases from (hyper)thermophiles: biochemistry and applications
Extremophiles (2014)
-
Genome and low-iron response of an oceanic diatom adapted to chronic iron limitation
Genome Biology (2012)
-
Active-site remodelling in the bifunctional fructose-1,6-bisphosphate aldolase/phosphatase
Nature (2011)