Abstract
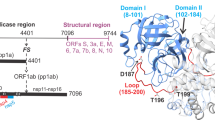
In the Gag-Pol polyprotein of HIV-1, the 99-amino acid protease is flanked at its N-terminus by a transframe region (TFR) composed of the transframe octapeptide (TFP) and 48 amino acids of the p6pol, separated by a protease cleavage site. The intact precursor (TFP-p6pol-PR) has very low dimer stability relative to that of the mature enzyme and exhibits negligible levels of stable tertiary structure. Thus, the TFR functions by destabilizing the native structure, unlike proregions found in zymogen forms of monomeric aspartic proteases. Cleavage at the p6pol-PR site to release a free N-terminus of protease is concomitant with the appearance of enzymatic activity and formation of a stable tertiary structure that is characteristic of the mature protease as demonstrated by nuclear magnetic resonance. The release of the mature protease from the precursor can either occur in two steps at pH values of 4 to 6 or in a single step above pH 6. The mature protease forms a dimer through a four-stranded β-sheet at the interface. Residues 1–4 of the mature protease from each subunit constitute the outer strands of the β-sheet, and are essential for maintaining the stability of the free protease but are not a prerequisite for the formation of tertiary structure and catalytic activity. Our experimental results provide the basis for the model proposed here for the regulation of the HIV-1 protease in the viral replication cycle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Wlodawer, A. & Erickson, J. Structure-based inhibitors of HIV-1 protease. Annu. Rev. Biochem. 62, 543– 585 (1993).
Oroszlan, S. & Luftig, R.B. Retroviral proteases. Curr. Top. Microbiol. Immunol. 157, 153– 185 (1990).
Candotti, D. et al. High variability of the gag/pol transframe region among HIV-1 isolates. C. R. Acad. Sci. 317, 183– 189 (1994).
Vogt, V.M. Proteolytic processing and particle maturation. Curr. Top. Microbiol. Immunol. 214, 95–131 ( 1996).
Wlodawer, A. & Vondrasek, J. Inhibitors of HIV-1 protease: a major success of structure-assisted drug design. Annu. Rev. Biophys. Biomol. Struct. 27, 249–284 (1998).
Condra, J.H. et al. In vivo emergence of HIV-1 variants resistant to multiple protease inhibitors. Nature 374, 569– 571 (1995).
Louis, J.M., Dyda, F., Nashed, N.T., Kimmel, A.R. & Davies, D.R. Hydrophilic peptides derived from the transframe region of Gag-Pol inhibit the HIV-1 protease. Biochemistry 37, 2105–2110 (1998).
Beissinger, M. et al. Sequence-specific resonance assignments of the 1H-NMR spectra and structural characterization in solution of the HIV-1 transframe protein p6. Eur. J. Biochem. 237, 383– 392 (1996).
Partin, K. et al. Deletion of sequences upstream of the proteinase improves the proteolytic processing of human immunodeficiency virus type 1. Proc. Natl. Acad. Sci. USA 88, 4776– 4780 (1991).
Zybarth, G. & Carter, C. Domains upstream of the protease (PR) in human immunodeficiency virus type 1 Gag-Pol influence PR autoprocessing. J. Virol. 69, 3878–3884 (1995).
Louis, J.M., Nashed, N.T., Parris, K.D., Kimmel, A.R. & Jerina, D.M. Kinetics and mechanism of autoprocessing of human immunodeficiency virus type 1 protease from an analog of the Gag- Pol polyprotein. Proc. Natl. Acad. Sci. USA 91, 7970–7974 (1994).
Co, E. et al. Proteolytic processing mechanisms of a miniprecursor of the aspartic protease of human immunodeficiency virus type 1. Biochemistry 33, 1248–1254 (1994).
Wondrak, E.M., Nashed, N.T., Haber, M.T., Jerina, D.M. & Louis, J.M. A transient precursor of the HIV-1 protease: isolation, characterization, and kinetics of maturation. J. Biol. Chem. 271, 4477–4481 (1996).
Louis, J.M. et al. Autoprocessing of the HIV-1 protease using purified wild-type and mutated fusion proteins expressed at high levels in Escherichia coli. Eur. J. Biochem. 199, 361–369 (1991).
Louis, J.M., Oroszlan, S. & Mora, P.T. Studies of the autoprocessing of the HIV-1 protease using cleavage site mutants. Adv. Exp. Med. Biol. 306 , 499–502 (1991).
Zybarth, G., Krausslich, H.G., Partin, K. & Carter, C. Proteolytic activity of novel human immunodeficiency virus type 1 proteinase proteins from a precursor with a blocking mutation at the N terminus of the PR domain. J. Virol. 68, 240– 250 (1994).
Tessmer, U. & Krausslich, H.G. Cleavage of human immunodeficiency virus type 1 proteinase from the N-terminally adjacent p6* protein is essential for efficient Gag polyprotein processing and viral infectivity. J. Virol. 72, 3459–3463 (1998).
Rose, J.R., Salto, R. & Craik, C.S. Regulation of autoproteolysis of the HIV-1 and HIV-2 proteases with engineered amino acid substitutions. J. Biol. Chem. 268, 11939–11945 (1993).
Mildner, A.M. et al. The HIV-1 protease as enzyme and substrate: mutagenesis of autolysis sites and generation of a stable mutant with retained kinetic properties. Biochemistry 33, 9405– 9413 (1994).
Wondrak, E.M. & Louis, J.M. Influence of flanking sequences on the dimer stability of human immunodeficiency virus type 1 protease. Biochemistry 35, 12957–12962 (1996).
Lindhofer, H., von der Helm, K. & Nitschko, H. In vivo processing of Pr160gag-pol from human immunodeficiency virus type 1 (HIV) in acutely infected, cultured human T- lymphocytes. Virology 214, 624–627 ( 1995).
Ho, D.D. et al. Characterization of human immunodeficiency virus type 1 variants with increased resistance to a C2-symmetric protease inhibitor. J. Virol. 68, 2016–2020 (1994).
Weber, I.T. Comparison of the crystal structures and intersubunit interactions of human immunodeficiency and Rous sarcoma virus proteases. J. Biol. Chem. 265, 10492–10496 ( 1990).
Lam, P.Y. et al. Rational design of potent, bioavailable, nonpeptide cyclic ureas as HIV protease inhibitors. Science 263, 380–384 (1994).
Wishart, D.S., Bigam, C.G., Holm, A., Hodges, R.S. & Sykes, B.D. 1H, 13C and 15N random coil NMR chemical shifts of the common amino acids. I. Investigations of nearest-neighbor effects. J. Biomol. NMR 5, 67–81 (1995).
Yamazaki, T. et al. Secondary structure and signal assignments of human-immunodeficiency-virus-1 protease complexed to a novel, structure-based inhibitor. Eur. J. Biochem. 219, 707–712 (1994).
Yamazaki, T. et al. Three-dimensional solution structure of the HIV-1 protease complexed with DMP323, a novel cyclic urea-type inhibitor, determined by nuclear magnetic resonance spectroscopy. Protein Sci. 5, 495–506 (1996).
Cherry, E. et al. Characterization of human immunodeficiency virus type-1 (HIV-1) particles that express protease-reverse transcriptase fusion proteins. J. Mol. Biol. 284, 43–56 (1998).
al-Janabi, J., Hartsuck, J.A. & Tang, J. Kinetics and mechanism of pepsinogen activation J. Biol. Chem. 247, 4628–4632 ( 1972).
Lin, X.L. et al. Enzymic activities of two-chain pepsinogen, two-chain pepsin, and the amino-terminal lobe of pepsinogen. J. Biol. Chem. 267, 17257–17263 (1992).
Khan, A.R. & James, M.N. Molecular mechanisms for the conversion of zymogens to active proteolytic enzymes. Protein Sci. 7, 815–836 (1998).
Peters, R.J. et al. Pro region C-terminus: protease active site interactions are critical in catalyzing the folding of alpha-lytic protease. Biochemistry 37, 12058–12067 ( 1998).
Hirel, P.H., Schmitter, M.J., Dessen, P., Fayat, G. & Blanquet, S. Extent of N-terminal methionine excision from Escherichia coli proteins is governed by the side-chain length of the penultimate amino acid. Proc. Natl. Acad. Sci. USA 86, 8247–8251 ( 1989).
Weber, I.T. et al. Crystallographic analysis of human immunodeficiency virus 1 protease with an analog of the conserved CA-p2 substrate—interactions with frequently occurring glutamic acid residue at P2' position of substrates. Eur. J Biochem. 249, 523– 300 (1997).
Bax, A. & Ikura, M. An efficient 3D NMR technique for correlating the proton and 15N backbone amide resonances with the alpha-carbon of the preceding residue in uniformly 15N/13C enriched proteins. J. Biomol. NMR 1, 99–104 (1991).
Wondrak, E.M., Louis, J.M., de Rocquigny, H., Chermann, J.C. & Roques, B.P. The gag precursor contains a specific HIV-1 protease cleavage site between the NC (P7) and P1 proteins. FEBS Lett. 333, 21–24 ( 1993).
Leis, J. et al. Standardized and simplified nomenclature for proteins common to all retroviruses. J.Virol. 62, 1808– 1809 (1988).
Acknowledgements
We wish to thank J. Hung for technical assistance, N.T. Nashed for discussions, L.K. Pannell for mass spectroscopic analyses, and H.R. Parikh for contract services. DMP323 was a generous gift from N. Hodge, DuPont Merck Pharmaceutical Company. This research was supported by the Intramural AIDS Targeted Program of the Office of the Director of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Louis, J., Clore, G. & Gronenborn, A. Autoprocessing of HIV-1 protease is tightly coupled to protein folding . Nat Struct Mol Biol 6, 868–875 (1999). https://doi.org/10.1038/12327
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/12327