Abstract
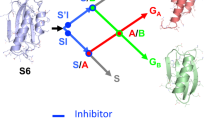
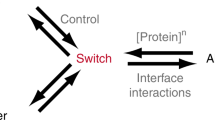
The amino-terminal domain of CD2 has the remarkable ability to fold in two ways: either as a monomer or as an intertwined, metastable dimer. Here we show that it is possible to differentially stabilize either fold by engineering the CD2 sequence, mimicking random mutagenesis events that could occur during molecular evolution. Crystal structures of a hinge-deletion mutant, which is stable as an intertwined dimer, reveal domain rotations that enable the protein to further assemble to a tetramer. These results demonstrate that a variety of folds can be adopted by a single polypeptide sequence, and provide guidance for the design of proteins capable of further assembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Murray, A.J., Lewis, S.J., Barclay, A.N. & Brady, R.L. Proc. Natl. Acad. Sci. USA 92, 7337–7341 (1995).
Jones, E.Y., Davies, S.J., Williams, A.F., Harlos, K. & Stuart, D.I. Nature 360, 232–239 (1992).
Bodian, D.L., Jones, E.Y., Harlos, K., Stuart, D.I. & Davis, S.J. Structure 2, 755–766 (1994).
Driscol, P.C., Cyster, G.J., Campbell, I.D. & Williams, A.F. Nature 353, 762–765 (1991).
Bennet, M.J., Schlunegger, M.P. & Eisenberg, D., Prot. Sci. 4, 2455–2468 (1995).
Peterson, A. & Seed, B. Nature 329, 400–403 (1987).
Arulanandam, A.R.N. et al. Proc. Natl. Acad. Sci. USA 90, 11613–11617 (1993).
Somoza, C., Driscoll, P.C., Cyster, J.G. & Williams, A.F. J. Exp. Med. 178, 549–558 (1993).
van der Merwe, P.A., McNamee, P.N., Davies, S.J. & Barclay, A.N. EMBO J. 12, 4945–4954 (1993).
Schlunegger, M.P., Bennet, M.J. & Eisenberg, D. (1997) Adv. Prot. Chem. 50, 61–122 (1997).
Barclay, A.N. et al. In The leukocyte antigen factsbook. pp. 54–62 (Academic Press, New York; 1993).
Chothia, C., Gelfand, I. & Kister, A. J. Mol. Biol. 278, 457–479 (1998).
Anderson, W.F., Ohlendorf, D.H., Tekeda, Y. & Mattews, B.W. Nature 290, 754–758 (1981).
Yan, Y. et al. Science 262, 2027-2030 (1993).
Tegoni, M., Ramoni, R., Bignetti, E., Spinelli, S. & Cambillau, C. Nature Struct. Biol. 3, 863–867 (1996).
Bianchet, M.A. et al. Nature Struct. Biol. 3, 934–939 (1996).
Otwinowski, Z. & Minor, W. Meth. Enz.. 276, 307–326 (1996).
Navaza J., Acta Crystallogr. A 50, 157–163 (1994).
Brünger, A.T. X-PLOR version 3.1: A system for X-ray crystallography and NMR (Yale University Press, New Haven, Connecticut; 1992).
Collaborative Computing Project Number 4 - programs for protein crystallography Acta Crystallogr. D 50, 760–763 (1994).
Lamzin, V.S. & Wilson, K.S. Acta Crystallogr. D 49, 129–147 (1993).
Acknowledgements
We thank A. N. Barclay (MRC CIU, University of Oxford), and S.J. Davies (IMM Oxford) for provision of CD2 mutants, and M.V. Hayes, A.R. Clarke and K.W. Wilkinson for general advice and discussions. We are grateful to the staff at Daresbury SRS for data collection facilities. A.J.M. was supported by a grant from the MRC, and J.G.H. from a studentship from the BBSRC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Murray, A., Head, J., Barker, J. et al. Engineering an intertwined form of CD2 for stability and assembly. Nat Struct Mol Biol 5, 778–782 (1998). https://doi.org/10.1038/1816
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/1816
This article is cited by
-
A five-residue motif for the design of domain swapping in proteins
Nature Communications (2019)
-
Two independently folding units of Plasmodium profilin suggest evolution via gene fusion
Cellular and Molecular Life Sciences (2015)
-
Molecular design principles underlying β-strand swapping in the adhesive dimerization of cadherins
Nature Structural & Molecular Biology (2011)
-
Backbone resonance assignment and order tensor estimation using residual dipolar couplings
Journal of Biomolecular NMR (2011)
-
HORI: a web server to compute Higher Order Residue Interactions in protein structures
BMC Bioinformatics (2010)