Abstract
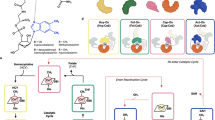
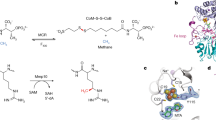
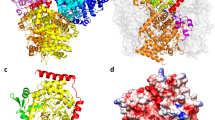
Biosynthesis of the corrin ring of vitamin B12 requires the action of six S-adenosyl-L-methionine (AdoMet) dependent transmethylases, closely related in sequence. The first X-ray structure of one of these, cobalt-precorrin-4 transmethylase, CbiF, from Bacillus megaterium has been determined to a resolution of 2.4 Å. CbiF contains two α/β domains forming a trough in which S-adenosyl-L-homocysteine (AdoHcy) binds. The location of AdoHcy and a number of conserved residues, helps define the precorrin binding site. A second crystal form determined at 3.1 Å resolution highlights the flexibility of two loops around this site. CbiF employs a unique mode of AdoHcy binding and represents a new class of transmethylase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stubbe, J. Binding site revealed for Nature's most beautiful cofactor. Science 266, 1663 (1994).
Scott, I. How nature synthesizes vatamin B12 - a survey of the last four billion years. Angew. Chem. 32, 1223-1376 ( 1993).
Battersby, A.R. How nature builds the pigments of life - The conquest of vitamin B12 . Science 264, 1551-1557 ( 1994).
Blanche, F. et al.,Vitamin B12 - How the problem of biosynthesis was solved. Angew. Chem. Int. Ed. Engl. 32, 1651-1653 (1995).
Debussche, L., Thibaut, D., Cameron, B., Crouzet, J. & Blanche, F. Biosynthesis of the corrin macrocycle of coenzyme-B 12 in Pseudomonas denitrificans. J. Bacteriol. 175, 7430-7440 (1993).
Lawerence, J.G. & Roth, J.R. The cobalamin (coenzyme B12 ) biosynthetic genes of Escherichia coli. J. Bacteriol. 177, 6371-6380 (1995 ).
Roth, J.R., Lawerence, J.G., Rubenfield, M., Kieffer-Higgins, S. & Church, G.M. Characterization of the cobalamin (vitamin B 12) biosynthetic genes of Salmonella typhimurium. J. Bacteriol. 175, 3303–3316 (1993).
Jordan, P.M. Highlights in haem biosynthesis. Curr. Opin. Struc. Biol. 4, 902–911 (1994).
Blanche, F., Parallels and decisive differences in vitamin B12 biosyntheses. Angew. Chem. Int. Ed. Engl. 32, 1651–1653 (1993).
Raux, E., Salmonella typhimurium cobalamin (vitamin B12) biosynthetic genes: Functional studies in S. typhiumurium and Escherichia coli. J. Bacteriol. 178, 753–767 (1996).
Raux, E., Thermes, C., Heathcote, P., Rambach, A. & Warren, M.J. A role for the Salmonella typhimurium cbiK in cobalamin (vitamin B12) and siroheme biosynthesis. J. Bacteriol. 179, 3203–3212 (1997).
Scott, I.A.,et al.Biosynthesis of vitamin B12: Factor IV, a new intermediate in the anarobic pathway. Proc. Natl. Acad. Sci. USA 93 , 14316–14319 (1996).
Martin-Verstraete, I., Debarbouille, M., Klier, A. & Rapoport, G. Levanase operon of Bacillus subtilus includes a fructose-specific phosphotransferase system regulating the expression of the operon. J. Mol. Biol. 214, 657–669 (1990).
Raux, E., Woodcock, S.C., Schubert, H.L., Wilson, K.S. & Warren, M.J. Cobalamin (vitamin B12) biosynthesis; Cloning, expression and crystallisation of the Bacillus megaterium S-adenosyl-L-methionine dependent cobalt-precorrin-4 transmethylase CbiF. Euro. J. Bacteriol. in the press (1998).
Oldfield, T.J. Real space refinement as a tool for model building. CCP4 Study Weekend: Macromolecular refinement (Dodson, E.J., Moore, M.H., Ralph, A. & Bailey, S., eds.) 67–74 (SERC Daresbury Laboratory, Warrington, UK.;1996).
Malone, T., Blumenthal, R.M. & Cheng, X. Structure-guided analysis reveals nine sequence motifs conserved among DNA amino-methyl-transferases, and suggests a catalytic mechanism for these enzymes . J. Mol. Biol. 253, 618– 632 (1995).
Hodel, A.E., Gershon, P.D., Shi, X. & Quiocho, F.A. The 1.85 Å structure of Vaccinia protein VP39: A bifunctional enzyme that participates in the modification of both mRNA ends. Cell 85, 247–256 (1996).
Blanche, F., Debussche, L., Thibaut, D., Crouzet, J. & Cameron, B. Purification and characterization of S-adenosyl-L-methionine: uroporphyrinogen III methyltransferase from Pseudomonas denitrificans. J. Bacteriol. 171, 4222–4231 (1989).
Drennan, C.L., Huang, S., Drummond, J.T., Matthews, R.G. & Ludwig, M.L. How a protein binds B12: A 3.0 Å X-ray structure of B12-binding domains of methionine synthase. Science 266, 1669–1674 ( 1994).
Roessner, C.A., et al.Expression of 9 Salmonella typhimurium enzymes for cobalamide synthesis . FEBS letters 301, 73– 78 (1992).
Woodcock, S.C. & Warren, M.J. Evidence for a covalent intermediate in the S-adenosyl-L-methionine-dependent transmethylation reaction caused by sirohaem synthase. Biochem. J. 313, 415 –421 (1996).
Woodcock, S.C., et al.The contribution of the CysGA and CysGB domains of siroheam synthase (CysG) towards cobalamin (vitamin B12) biosynthesis. Biochem. J. 330, 121–129 ( 1998).
Schluckebier, G., O'Gara, M., Saenger, W. & Cheng, X. Universal catalytic domain structure of AdoMet-dependent methyltransferases. J. Mol. Biol. 247, 16–20 ( 1995).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Bernstein, F.C., Koetzle, T.F., Williams, G.J.B., Meyer, E.J., Brice, M.D., Rogers, J.K., Kennard, O., Shimanouchi, T. & Tasumi, M. (1977).The protein data bank: a computer-based archival file for macromolecular structures. J. Mol. Biol. 112, 535–542.
Freyman, D.M., Keenan, R.J., Stoud, R.M. & Walter, P. The structure of the conserved GTPase domain of the signal recognition particle. Nature 385, 361–365 ( 1997).
Orengo, C.A., Michie, A.D., Jones, S., Jones, D.T., Swindells, M.B. & Thornton, J.M. CATH- a hierarchic classification of protein domain structures. Structure 5, 1093–1108 (1997).
Dixon, M.M., Huang, S., Matthews, R.G. & Ludwig, M. The structure of the C-terminal domain of methionine synthase: presenting S-adenosylmethionine for reductive methylastion of B12. Structure 4, 1263–1275 (1996).
Otwinowski, Z. processing of X-ray diffraction data collected in ossilation mode. Meth. Enz . 276, 307–326 ( 1991).
Otwinowski, Z. Maximum likelihood refinement of heavy atom parameters. Proceedings of the CCP4 Study Weekend (Wolf, W., Evans, P.R. & Leslie, A.G.W., eds) 80- 88 (SERC Daresbury Laboratory, Warrington, UK; 1991 ).
Cowtan, K. in CCP4 & ESF-EACBM Newsletter on Protein Crystallography 34- 38 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjelgaard, M. Improved methods for building protein models in electron density maps and location of errors in these models. Acta Crystallogr. A 47, 110-119 (1991).
Brunger, A.T. X-PLOR Version 3.1: A system for X-ray Crystallography and NMR (Yale University Press, New Haven, Connecticut, USA; 1992).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum likelihood method. Acta Crystallogr. D 53, 240-255 (1997).
Ramachandran, S. Conformations of polypeptides and proteins. Adv. Prot. Chem. 23, 283-437 (1968).
Laskowski, R.A., MacAuthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK - a program to check the sterochemical quality of protein structures . J. Appl. Crystallogr. 26, 283- 291 (1993).
Navaza, J. AMORE - an automated package for molecular replacement. Acta Crystallogr. A 50, 157-163 (1994).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities . J. Mol. Graph. 15, 133-138 (1997).
Kraulis, P.J. MOLSCRIPT - a program to produce both detailed and schematic plots of proteins structures . J. Appl. Crystallogr. 24, 946- 950 (1991).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281-296 (1991).
Acknowledgements
We gratefully acknowledge funding from the National Institutes of Health, the Wellcome Trust and the Biotechnology and Biological Sciences Research Council. We thank the Central Laboratory of the Research Council and the staff of the Daresbury Laboratory for the provision of synchrotron radiation facilities and the BBSRC for support of such usage through the Rolling Project Mode Time allocation to York.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schubert, H., Wilson, K., Raux, E. et al. The X-ray structure of a cobalamin biosynthetic enzyme, cobalt-precorrin-4 methyltransferase. Nat Struct Mol Biol 5, 585–592 (1998). https://doi.org/10.1038/846
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/846
This article is cited by
-
Binding studies of a putative C. pseudotuberculosis target protein from Vitamin B12 Metabolism
Scientific Reports (2019)
-
Crystal structure of putative CbiT from Methanocaldococcus jannaschii: an intermediate enzyme activity in cobalamin (vitamin B12) biosynthesis
BMC Structural Biology (2013)
-
Characterization of the cofactor-binding site in the SPOUT-fold methyltransferases by computational docking of S-adenosylmethionine to three crystal structures
BMC Bioinformatics (2003)
-
CysG structure reveals tetrapyrrole-binding features and novel regulation of siroheme biosynthesis
Nature Structural & Molecular Biology (2003)