Abstract
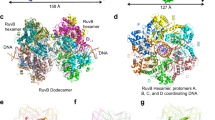
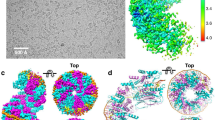
Here we present the crystal structure of the Escherichia coli protein RuvA bound to a key DNA intermediate in recombination, the Holliday junction. The structure, solved by isomorphous replacement and density modification at 6 Å resolution, reveals the molecular architecture at the heart of the branch migration and resolution reactions required to process Holliday intermediates into recombinant DNA molecules. It also reveals directly for the first time the structure of the Holliday junction. A single RuvA tetramer is bound to one face of a junction whose four DNA duplex arms are arranged in an open and essentially four-fold symmetric conformation. Protein–DNA contacts are mediated by two copies of a helix-hairpin-helix motif per RuvA subunit that contact the phosphate backbone in a very similar manner. The open structure of the junction stabilized by RuvA binding exposes a DNA surface that could be bound by the RuvC endonuclease to promote resolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
West, S.C. Annu. Rev. Genet. 31, 213–244 (1997).
Otsuji, N., Iyehara, H. & Hideshima, Y. J. Bacterial. 117, 337–344 (1974).
Lloyd, R.G., Benson, F.E. & Shurvinton, C.E. Mol. Gen. Genet. 194, 303–309 (1984).
Benson, F., Collier, S. & Lloyd, R.G. Mol. Gen. Genet. 225, 266–272 (1991).
Whitby, M.C., Bolt, E.L., Chan, S.N. & Lloyd, R.G. J. Mol. Biol. 264, 878–890 (1996).
Rafferty, J.B. et al. Science 274, 415–421 (1996).
Eggleston, A.K., Mitchell, A.H. & West, S.C. Cell 89, 607–617 (1997).
van Gool, A.J., Shah, R., Mezard, C. & West, S.C. EMBO. J. 17, 1838–1845 (1998).
Thayer, M.M., Ahern, H., Xing, D.X., Cunningham, R.P. & Tainer, J.A. EMBO J. 14, 4108–4120 (1995).
Doherty, A.J., Serpell, L.C. & Ponting, C.P. Nucleic Acids Res. 24, 2488–2497 (1996).
Rice, D.W., Rafferty, J.B., Artymiuk, P.J. & Lloyd, R.G. Curr. Opin. Struct. Biol. 7, 798–803 (1997).
Hargreaves, D. et al. Acta. Crystallogr., in the press.
Richmond, T.J., Finch, J.T., Rushton, B., Rhodes, D. & Klug, A. Nature 311, 532–537 (1984).
Pelletier, H., Sawaya, M.R., Kumar, A., Wilson, S.H. & Kraut, J. Science 264, 1891–1903 (1994).
Pelletier, H., Sawaya, M.R., Wolfle, W., Wilson, S.H. & Kraut, J. Biochemistry 35, 12742–12761 (1996).
Rafferty, J.B. et al. J. Mol. Biol. 278, 105–116 (1998).
Guo, F., Gopaul, D.N. & Van Duyne, G.D. Nature 389, 40–46 (1997).
Parsons, C.A., Tsaneva, I., Lloyd, R.G. & West, S.C. Proc. Nat. Acad. Sci. USA 89, 5452–5456 (1992).
Yu, X., West, S.C. & Egelman, E.H. J Mol. Biol. 266, 217–222 (1997).
Sharples, G.J., Benson, F.E., Illing, G.T. & Lloyd, R.G. Mol. Gen. Genet. 221, 219–226 (1990).
Mandal, T.N., Mahdi, A.A., Sharples, G.J. & Lloyd, R.G. J Bacteriol. 175, 4325–4334 (1993).
Ariyoshi, M. et al. Cell 78, 1063–1072 (1994).
Otwinowski, Z. & Minor, W. Meth. Enz. 276, 307–326 (1997).
CCP4. The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D50, 760–763 (1994).
Otwinowski, Z. Proceedings of the CCP4 Study Weekend: Isomorphous Replacement and Anomalous Scattering, p80 (SERC Daresbury Laboratory, Warrington, U.K., 1991).
Cowtan, K. Joint CCP4 and ESF-EACBM Newsletter and Protein Crystallography 31, 34–38 (1994).
Kleywegt, G.J. & Jones, T.A. Masks and Bones. Proceedings of the CCP4 Study Weekend: From First Map to Final Model, p59 (EPSRC Daresbury Laboratory, Warrington, UK., 1994).
Bennett, R.J., Dunderdale, H.J. & West, S.C. Cell 74, 1021–1031 (1993).
Jones, T.A. J. Appl. Crystallogr. 11, 268–270 (1978).
Ferrin, T.E., Huang, C.C., Jarvis, L.E. & Langridge, R. J Mol. Graphics 6, 13–27 (1988).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hargreaves, D., Rice, D., Sedelnikova, S. et al. Crystal structure of E.coli RuvA with bound DNA Holliday junction at 6 Å resolution. Nat Struct Mol Biol 5, 441–446 (1998). https://doi.org/10.1038/nsb0698-441
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0698-441
This article is cited by
-
DNA binding to RecD: role of the 1B domain in SF1B helicase activity
The EMBO Journal (2008)
-
Crystal structure of T4 endonuclease VII resolving a Holliday junction
Nature (2007)
-
The structural basis of Holliday junction resolution by T7 endonuclease I
Nature (2007)
-
Happy Hollidays: 40th anniversary of the Holliday junction
Nature Reviews Molecular Cell Biology (2004)
-
RuvAB-directed branch migration of individual Holliday junctions is impeded by sequence heterology
The EMBO Journal (2004)