Abstract
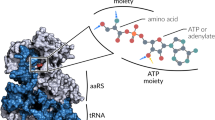
Accurate translation of the genetic code depends on the ability of aminoacyl-tRNA synthetases to distinguish between similar amino acids. In order to investigate the basis of amino acid recognition and to understand the role played by the zinc ion present in the active site of threonyl-tRNA synthetase, we have determined the crystal structures of complexes of an active truncated form of the enzyme with a threonyl adenylate analog or threonine. The zinc ion is directly involved in threonine recognition, forming a pentacoordinate intermediate with both the amino group and the side chain hydroxyl. Amino acid activation experiments reveal that the enzyme shows no activation of isosteric valine, and activates serine at a rate 1,000-fold less than that of cognate threonine. This study demonstrates that the zinc ion is neither strictly catalytic nor structural and suggests how the zinc ion ensures that only amino acids that possess a hydroxyl group attached to the β-position are activated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Meinnel, T., Mechulam, Y. & Blanquet, S. In Aminoacyl-tRNA synthetase: occurence, structure, and function (Söll, D. & RajBhandary, U.L. eds) 251–291 (ASM Press, Washington DC; 1995).
Fersht, A. Enzyme, structure and mechanism (W. H. Freeman and Company, New York; 1985).
Jakubowski, H. & Goldman, E. Microbiol. Rev. 56, 412–429 (1992).
Baldwin, A.N. & Berg, P. J. Biol. Chem. 241, 839–845 (1966).
Loftfield, R.B. & Vanderjagt, M.A. Biochem. J. 128, 1353–1356 (1972).
Schmidt, E. & Schimmel, P. Science 264, 265–267 (1994).
Nureki, O. et al. Science 280, 578–582 (1998).
Fersht, A.R. Science 280, 541 (1998).
Lin, L., Hale, S.P. & Schimmel, P. Nature 384, 33–34 (1996).
Silvian, L.F., Wang, J. & Steitz, T.A. Science 285, 1074–1077 (1999).
Nomanbhoy, T.K., Hendrickson, T.L. & Schimmel, P. Mol. Cell 4, 519–528 (1999).
Tsui, W.C. & Fersht, A.R. Nucleic Acids Res. 9, 4627–4637 (1981).
Sankaranarayanan, R. et al. Cell 97, 371–381 (1999).
Arnez, J.G. & Moras, D. Trends Biochem. Sci. 22, 211–216 (1997).
Alberts, I.L., Nadassy, K. & Wodak, S.J. Protein Sci. 7, 1700–1716 (1998).
Cavarelli, J. et al. EMBO J. 13, 327–337 (1994).
Belrhali, H. et al. Science 263, 1432–1436 (1994).
Arnez, J.G., Augustine, J.G., Moras, D. & Francklyn, C.S. Proc. Natl. Acad. Sci. 94, 7144–7149 (1997).
Schmitt, E. et al. EMBO J. 17, 5227–5237 (1998).
Eigner, E.A. & Loftfield, R.B. Methods Enzymol. 29, 601–619 (1974).
Igloi, G.L., von der Haar, F. & Cramer, F. Methods Enzymol. 59, 282–291 (1979).
Fersht, A.R., Shindler, J.S. & Tsui, W.C. Biochemistry 19, 5520–5524 (1980).
Christner, P., Carpousis, A., Harsch, M. & Rosenbloom, J. J. Biol. Chem. 250, 7623–7630 (1975).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Collaborative Computational Project Number 4. CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Evans, S.V. J. Mol. Graphics 11, 134–138 (1993).
Hodel, A., Kim, S.-H. & Brunger, A.T. Acta Crystallogr. A 48, 851–859 (1992).
Acknowledgements
We thank P. Schimmel for a critical reading of the manuscript. We would like to acknowledge the encouragement and support received from M. Springer, B. Ehresmann and C. Ehresmann. This work was supported by grants from an EEC project, CNRS, INSERM, ULP, and Ministère de la Recherche et de la Technologie.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sankaranarayanan, R., Dock-Bregeon, AC., Rees, B. et al. Zinc ion mediated amino acid discrimination by threonyl-tRNA synthetase. Nat Struct Mol Biol 7, 461–465 (2000). https://doi.org/10.1038/75856
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/75856
This article is cited by
-
Tyrosine-targeted covalent inhibition of a tRNA synthetase aided by zinc ion
Communications Biology (2023)
-
Specificity and catalysis hardwired at the RNA–protein interface in a translational proofreading enzyme
Nature Communications (2015)
-
Aminoacyl-tRNA synthetase dependent angiogenesis revealed by a bioengineered macrolide inhibitor
Scientific Reports (2015)
-
Structural basis for full-spectrum inhibition of translational functions on a tRNA synthetase
Nature Communications (2015)
-
Critical Role of Zinc Ion on E. coli Glutamyl-Queuosine-tRNAAsp Synthetase (Glu-Q-RS) Structure and Function
The Protein Journal (2014)