Abstract
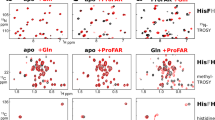
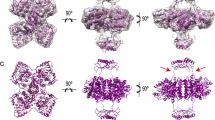
Phenylalanine hydroxylase converts phenylalanine to tyrosine, a rate-limiting step in phenylalanine catabolism and protein and neurotransmitter biosynthesis. It is tightly regulated by the substrates phenylalanine and tetrahydrobiopterin and by phosphorylation. We present the crystal structures of dephosphorylated and phosphorylated forms of a dimeric enzyme with catalytic and regulatory properties of the wild-type protein. The structures reveal a catalytic domain flexibly linked to a regulatory domain. The latter consists of an N-terminal autoregulatory sequence (containing Ser 16, which is the site of phosphorylation) that extends over the active site pocket, and an α-β sandwich core that is, unexpectedly, structurally related to both pterin dehydratase and the regulatory domains of metabolic enzymes. Phosphorylation has no major structural effects in the absence of phenylalanine, suggesting that phenylalanine and phosphorylation act in concert to activate the enzyme through a combination of intrasteric and possibly allosteric mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hufton, S.E., Jennings, I.G. & Cotton, R.G.H. Biochem. J. 311, 353– 366 (1995).
Kappock, T.J. & Caradonna, J.P. Chem. Rev. 96, 2659–2756 (1996).
Nowacki, P., Byck, S., Prevost, L. & Scriver, C.R. Nucleic Acids Res. 25, 139–142 ( 1997).
Kobe, B. et al. Protein Sci. 6, 1352–1357 (1997).
Goodwill, K.E. et al. Nature Struct. Biol. 4, 578– 585 (1997).
Erlandsen, H. et al. Nature Struct. Biol. 4, 995– 1000 (1997).
Fusetti, F., Erlandsen, H., Flatmark, T. & Stevens, R.C. J. Biol. Chem. 273, 16962–16967 (1998).
Fischer, R.S., Zhao, G. & Jensen, R.A. J. Gen. Microbiol. 137, 1293– 1301 (1991).
Holm, L. & Sander, C. Nucleic Acids Res. 22 , 3600–3609 (1994).
Schuller, D.J., Grant, G.A. & Banaszak, L.J. Nature Struct. Biol. 2, 69– 76 (1995).
Gallagher, D.T. et al. Structure 6, 465–475 (1998).
Lipscomb, W.N. Adv. Enzymol. Relat. Areas Mol. Biol. 68, 67– 151 (1994).
Cronk, J.D., Endrizzi, J.A. & Alber, T. Protein Sci. 269, 24657– 24665 (1994).
Lei, X.-D. & Kaufman, S. Proc. Natl. Acad. Sci. USA 95, 1500–1504 (1998).
Zhao, G., Xia, T., Song, J. & Jensen, R.A. Proc. Natl. Acad. Sci. USA 91, 1366–1370 ( 1994).
Johnson, L.N. & O'Reilly, M. Curr. Opin. Struct. Biol. 6, 762–769 (1996).
Canagarajah, B.J., Khokhlatchev, A., Cobb, M.H. & Goldsmith, E.J. Cell 90, 859–869 ( 1997).
Kobe, B. et al. EMBO J. 15, 6810–6821 (1996).
Kissinger, C.R. et al. Nature 378, 641–644 (1995).
Khan, A.R. & James, M.N.G. Protein Sci. 7, 815–836 (1998).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 ( 1997).
CCP4, Acta Crystallogr. D50, 760–763 (1994).
Furey, W. & Swaminathan, S. Methods Enzymol. 277, 590–620 (1997).
La Fortelle, E.de & Bricogne, G. Methods Enzymol. 276, 472–494 ( 1997).
Perrakis, A., Sixma, T.K., Wilson, K.S. & Lamzin, V.S. Acta Crystallogr. D53, 448–455 (1997).
Brünger, A.T., Kuriyan, J. & Karplus, M. Science 235, 458– 460 (1987).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A47, 110–119 ( 1991).
Merritt, E.A. & Murphy, M.E.P. Acta Crystallogr. D50, 869–873 (1994).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281– 296 (1991).
Kabsch, W. & Sander, C. Biopolymers 22, 2577–2637 (1983).
Acknowledgements
We thank J. Rossjohn and A. Oakley for help with synchrotron data collection, F. Katsis for the synthesis of heavy-atom substituted pterins, M. Parker for discussions, A. Perrakis for help with the wARP procedure, J. Hunt for automated density modification and refinement scripts, R. Read and B. Hazes for a version of SIGMAA, J. Varghese, B. Vandonkelaar, M. Lawrence and P. Colman for the xenon soaking device, personnel at the Photon Factory and DESY synchrotrons for help with data collection, and T. Teh for comments on the manuscript. We apologize to those whose work or original publication could not be cited because of space limitations. This work was supported by Australian Research Council (B.K.), W.M. Keck Foundation (R.C.S.) and NHMRC (R.G.H.C, B.E.K.); B.K. is a Wellcome Senior Research Fellow in Medical Science in Australia.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kobe, B., Jennings, I., House, C. et al. Structural basis of autoregulation of phenylalanine hydroxylase. Nat Struct Mol Biol 6, 442–448 (1999). https://doi.org/10.1038/8247
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/8247
This article is cited by
-
Structure of full-length wild-type human phenylalanine hydroxylase by small angle X-ray scattering reveals substrate-induced conformational stability
Scientific Reports (2019)
-
Structural mechanism for the arginine sensing and regulation of CASTOR1 in the mTORC1 signaling pathway
Cell Discovery (2016)
-
Stable preparations of tyrosine hydroxylase provide the solution structure of the full-length enzyme
Scientific Reports (2016)
-
Structural basis for ligand-dependent dimerization of phenylalanine hydroxylase regulatory domain
Scientific Reports (2016)
-
Crystal structure of a hypothetical protein, TTHA0829 from Thermus thermophilus HB8, composed of cystathionine-β-synthase (CBS) and aspartate-kinase chorismate-mutase tyrA (ACT) domains
Extremophiles (2016)