Abstract
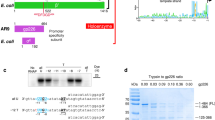
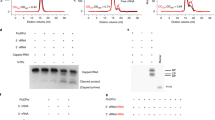
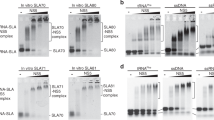
The 3′ end of brome mosaic virus RNA contains a tRNA-like sequence that directs its RNA synthesis. A stem loop structure in this sequence, stem loop C (SLC), was investigated using NMR, and correlated with its ability to direct RNA synthesis by its replicase. SLC consists of two discrete domains, a flexible stem with an internal loop and a rigid stem containing a 5′-AUA-3′ triloop. Efficient RNA synthesis requires the sequence on only one side of the flexible stem and a specific compact conformation of the triloop. A high resolution structure of the triloop places the 5′ adenine out in solution, and the 3′ adenine within the triloop, held tightly through stacking and unusual hydrogen bonds. This high resolution structure of an RNA promoter from a (+)-strand RNA virus provides new insights into how the RNA-dependent RNA polymerase binds to the RNA to initiate synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Buck, K.W. Comparison of the replication of positive-stranded RNA viruses of plants and animals. Adv. Virus Res. 47, 159– 251 (1996).
Hall, T.C., Rao, A.L.N., Pogue, G.P., Huntley, C.C. & Marsh, L. E. Replication, repair, and recombination of brome mosaic virus RNA. In New aspects of positive-strand RNA viruses (eds Brinton, M.A. & Heinz, F.X.) 47–54 (American Society for Microbiology, Washington, DC; 1990).
Ahlquist, P. Bromovirus RNA replication and transcription. Curr. Opin. Genet. Dev. 2, 71–76 ( 1992).
Ahlquist, P., Dasgupta, R. & Kaesberg, P. Near identity of 3′ RNA secondary structure in bromoviruses and cucumber mosaic virus. Cell 23, 183–189 (1981).
Miller, W.A., Bujarski, J.J., Dreher, T.W. & Hall, T.C. Minus-strand initiation by brome mosaic virus replicase within the 3′ tRNA-like structure of native and modified RNA templates. J. Mol. Biol. 187, 537–546 ( 1986).
Deutscher, M.P. In Enzymes of nucleic acid synthesis and modification vol. 2 RNA enzymes (ed. Jacob, S.T.) 159– 183 (CRC Press, Boca Ration; 1984).
Joshi, R.L., Joshi, S., Chapeville, F. & Haenni, A.L. tRNA-like structures of plant viral RNAs: conformational requirements for adenylation and aminoacylation. EMBO J. 2, 1123–1127 (1983).
Perret, V., Florentz, C., Dreher, T. & Giege, R. Structural analogies between the 3′ tRNA-like structure of brome mosaic virus RNA and yeast tRNATyr revealed by protection studies with yeast tyrosyl-tRNA synthetase. Eur. J. Biochem. 185, 331– 339 (1989).
Felden, B., Florentz, C., Giege, R. & Westhof, E. Solution structure of the 3′-end of brome mosaic virus genomic RNAs. Conformational mimicry with canonical tRNAs. J. Mol. Biol. 234, 508–531 (1994).
Rietveld, K., Pleij, C.W.A. & Bosch, L. Three-dimensional models of the tRNA-like 3′ termini of some plant viral RNAs. EMBO J. 2, 1079 –1085 (1983).
Dreher, T.W., Bujarski, J.J. & Hall, T.C. Mutant viral RNAs synthesized in vitro show altered aminoacylation and replicase template activities. Nature 311, 171–175 ( 1984).
Dreher, T.W. & Hall, T.C. Mutational analysis of the sequence and structural requirements in brome mosaic virus RNA for minus strand promoter activity. J. Mol. Biol. 201, 31– 40 (1988).
Chapman, M.R. & Kao, C.C. A minimal RNA promoter for minus-strand RNA synthesis by the brome mosaic virus polymerase complex. J. Mol. Biol. 286, 709–720 (1999).
Jaeger, J.A., Turner, D.H. & Zuker, M. Improved predictions of secondary structures for RNA . Proc. Natl. Acad. Sci. USA 86, 7706– 7710 (1989).
Milligan, J.F. & Uhlenbeck, O.C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 180, 51–62 (1989).
Wu, M. & Tinoco, I. Jr . RNA folding causes secondary structure rearrangement. Proc. Natl. Acad. Sci. USA 95, 11555–11560 (1998).
Rao, A.L.N. & Hall, T.C. Recombination and polymerase error facilitate restoration of infectivity in brome mosaic virus. J. Virol. 67, 969–979 ( 1993).
Petersheim, M. & Turner, D.H. Base-stacking and base-pairing contributions to helix stability: thermodynamics of double-helix formation with CCGG, CCGGp, CCGGAp, ACCGGp, CCGGUp, and ACCGGUp. Biochemistry 22, 256–263 (1983).
Riesner, D., Maass, G. Thiebe, R., Philippsen, P. & Zachau, H.G. The conformational transitions in yeast tRNAPhe as studied with tRNAPhe fragments. Eur. J. Biochem. 36, 76–88 ( 1973).
Freier, S.M., Kierzek, R., Jaeger, J.A., Sugimoto, N., Caruthers, M.H., Neilson, T. & Turner, D.H. Improved free-energy parameters for predictions of RNA stability. Proc. Natl. Acad. Sci. USA 83, 9373– 9377 (1986).
Turner, D.H., Sugimoto, N. & Freier, S.M. RNA structure prediction. Annu. Rev. Biophys. Chem. 17, 167–192 ( 1988).
Borer, P.N., Dengler, B., Tinoco, I. Jr. & Uhlenbeck, O.C. Stability of ribonucleic acid double-stranded helices. J. Mol. Biol. 86, 843–853 ( 1974).
Puglisi, J.D., Wyatt, J.R. & Tinoco, I. Jr . Solution conformation of an RNA hairpin loop. Biochemistry 29, 4215– 4226 (1990).
Davis, P.W., Thurmes, W. & Tinoco, I. Jr . Structure of a small RNA hairpin. Nucleic Acids Res. 21, 537–545 (1993).
Varani, G. & Tinoco, I. Jr . RNA structure and NMR spectroscopy Q. Rev. Biophys. 24, 479 –532 (1991).
Saenger, W. Principles of nucleic acid structure. (Springer Verlag, NY; 1984).
Lavery, R. & Sklenar, J. The definition of generalized helicoidal parameters and of axis curvature for irregular nucleic acids. J. Biomol. Struct. Dynam. 6, 63–91 (1988).
Lavery, R. & Sklenar, J. Defining the structure of irregular nucleic acids: conventions and principles. J. Biomol. Struct. Dynam. 6, 655–667 ( 1989).
Ahlquist, P., Dasgupta, R., and Kaesberg, P. Near identity of 3′ RNA secondary structure in bromoviruses and cucumber mosaic virus. Cell 23, 183–189 (1981).
Romero, J. Dzianott, A.M., and Bujarski, J.J. The nucleotide sequence and genome organization of the RNA2 and RNA3 segments in broad bean mottle virus. Virology 187, 671–681 ( 1992).
Siegel, R.W., Adkins, S. & Kao, C.C. Sequence-specific recognition of a subgenomic promoter by a viral RNA polymerase. Proc. Natl. Acad. Sci. USA 94, 11238–11243 (1997).
Sivakumaran, K., Kim, C.-H., Tayon Jr. R. & Kao, C.C. RNA Sequence and secondary structural determinants in a minimal viral promoter that directs replicase recognition and initiation of genomic plus-strand RNA synthesis. J. Mol. Biol. 294, 667– 682 (1999).
Hansen. J.L., Long, A.M. & Schultz, S.C. Structure of the RNA-dependent RNA polymerase of poliovirus . Structure 5, 1109–1122 (1997).
Lesburg, C.A., Cable, M.B., Ferrari, E., Hong, Z., Mannarino, A.F. & Weber, P.C. Crystal structure of the RNA-dependent RNA polymerase from hepatitis C virus reveals a fully encircled active site. Nature Struct. Biol. 6, 937–943 (1999).
Quadt, R. & Jaspars, E.M.J. Purification and characterization of brome mosaic virus RNA-dependent RNA polymerase. Virology 178, 189–194 (1990).
Batey, R.T., Inada, M., Kujawinski, E., Puglisi, J.D. & Williamson, J.R. Preparation of isotopically labeled ribonucleotides for multidimensional NMR spectroscopy of RNA. Nucleic Acids Res. 20, 4515–4523 (1992).
Puglisi, J.D. & Tinoco, I. Jr . Absorbance melting curves of RNA. Methods Enzymol. 180, 304 –325 (1989).
Plateau, P. & Gueron, M. Exchangeable proton NMR without base-line distortion, using strong-pulse sequences. J. Am. Chem. Soc. 104, 7310–7311 (1982).
Kay, L.E. Field gradient techniques in NMR spectroscopy. Curr. Opin. Struc. Biol. 5, 674–681 ( 1995).
Sklénar, V., Miyashiro, H., Zon, G. & Bax, A. Assignment of the 31P and 1H resonances in oligonucleotides by two-dimensional NMR spectroscopy. FEBS Lett., 208, 94– 98 (1986).
Varani, G. & Tinoco, I. Jr. Carbon assignments and heteronuclear coupling constants for an RNA oligonucleotide from natural abundance 13C-1H correlated experiments. J. Am. Chem. Soc. 113, 9349–9354 (1991).
Marino, J.P., Prestegard, J.H. & Crothers, D.M. Correlation of adenine H2/H8 resonances in uniformly 13C labeled RNAs by 2D HCCH-TOCSY: a new tool for 1H assignment. J. Am. Chem. Soc. 116, 2205–2206 (1994).
Brünger, A.T. X-PLOR version 3.1: a system for X-ray crystallography and NMR (Yale University Press, New Haven; 1993).
Wimberly, B.T. NMR derived structures of RNA loops: the conformation of eukaryotic 5S ribosomal loop E. (PhD thesis, University of California, Berkeley; 1992).
Varani, G., Aboul-ela, F. & Allain, F.H.-T. NMR investigation of RNA structure. Prog. NMR Spectrosc. 29, 51–127 (1996).
Sun, J.H., Adkins, S., Faurote, G. & Kao, C.C. Initiation of (−)-strand RNA synthesis catalyzed by the BMV RNA-dependent RNA polymerase: synthesis of oligonucleotides. Virology 226, 1– 12 (1996).
Acknowledgements
We thank B. Dengler for general lab management and D. Koh for DNA template synthesis. We also thank J. Pelton for NMR advice and H.-J. Park for his work in RNA sample preparation. Funding was provided by a National Institute of Health grant and a Department of Energy grant to I.T., and by a National Science Foundation grant to C.K.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, CH., Kao, C. & Tinoco, I. RNA motifs that determine specificity between a viral replicase and its promoter. Nat Struct Mol Biol 7, 415–423 (2000). https://doi.org/10.1038/75202
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/75202