Abstract
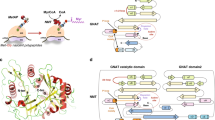
N-myristoyl transferase (NMT) catalyzes the transfer of the fatty acid myristate from myristoyl-CoA to the N-terminal glycine of substrate proteins, and is found only in eukaryotic cells. The enzyme in this study is the 451 amino acid protein produced by Candida albicans, a yeast responsible for the majority of systemic infections in immuno-compromised humans. NMT activity is essential for vegetative growth, and the structure was determined in order to assist in the discovery of a selective inhibitor of NMT which could be developed as an anti-fungal drug. NMT has no sequence homology with other protein sequences and has a novel α/β fold which shows internal twofold symmetry, which may be a result of gene duplication. On one face of the protein there is a long, curved, relatively uncharged groove, at the center of which is a deep pocket. The pocket floor is negatively charged due to the vicinity of the C-terminal carboxylate and a nearby conserved glutamic acid residue, which separates the pocket from a cavity. These observations, considered alongside the positions of residues whose mutation affects substrate binding and activity, suggest that the groove and pocket are the sites of substrate binding and the floor of the pocket is the catalytic center.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Bhatnagar, R.S. & Gordon, J.I. Understanding covalent modifications of proteins by lipids: where cell biology and biophysics mingle. Trends Cell Biol. 7, 14–20 (1997)
Boutin, J.A. Myristoylation Cell Signal 9, 15–35 (1997).
Rudnick, D.A., McWherter, C.A., Gokel, G.W. & Gordon, J.I., Myristoyl-CoA: Protein N-myristoyl transferase. Adv. Enzymol. 67, 375–430 (1993).
Johnson, D.R., Bhatnagar, R.S., Knoll, L.J. & Gordon, J.I. Genetic and biochemical studies of protein N-myristoylation. Ann. Rev. Biochem. 63, 869–914 (1994).
Boutin, J.A. et al. Myristoyl-CoA : protein N-myristoyl transferase activity in cancer cells. Purification and characterisation of a cytosolic isoform from the murine leukaemia cell line L1210. Eur. J. Biochem 214, 853–867 (1993).
Resh, M.D. Interaction of tyrosine kinase oncoproteins with cellular membranes. Biochem. Biophys. Acta 1115, 307–322 (1993).
Knoll, L.J., Russell Johnson, D., Bryant, M.L. & Gordon, J.I. “Functional Significance of Myristoyl Moiety in N-Myristoyl Proteins” in Lipid Modifications of Proteins eds. Casey, P.J. & Buss, J.E. Methods in Enzymology 250, 405–435 (1995).
Rudnick, D.A. et al. Kinetic and structural evidence for a sequential ordered Bi Bi mechanism of catalysis by Saccharomyces cerevisiae myristoyl-CoA:protein N-myristoyl transferase. J. Biol. Chem. 266, 9732–9739 (1991).
Rudnick, D.A., Johnson, R.L. & Gordon, J.I. Studies of the catalytic activities and substrate specificities of Saccharomyces cerevisiae myristoyl-coenzyme A: protein N-myristoyl transferase deletion mutants and human/yeast NMT chimeras in Escherichia coli and Saccharomyces cerevisiae. J. Biol. Chem. 267, 23852–23861 (1992).
Zhang, L., Jackson-Machelski, E. & Gordon, J.I. Biochemical studies of Saccharomyces cerevisiae myristoyl-coenzyme A : protein N-myristoyl transferase mutants. J. Biol. Chem. 271, 33131–33140 (1996).
Peseckis, S.M. & Resh, M.D. Fatty acyl transfer by human N-myristoyl transferase is dependent upon conserved cysteine and histidine residues J. Biol. Chem. 269, 30888–30892 (1994).
Duronio, R.J., Towler, D.A., Heuckeroth, R.O. & Gordon, J.I. Disruption of the yeast N-myristoyl transferase gene causes recessive lethality. Science 243, 796–800 (1989).
Duronio, R.J., Reed, S.I. & Gordon, J.I. Mutations of human myristoyl-CoA : protein N-myristoyl transferase cause temperature-sensitive myristic acid auxotrophy in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 89, 4129–4133 (1992).
Wiegand, R.C. et al. The Candida albicans myristoyl-CoA : protein N-myristoyl transferase gene. Isolation and expression in Saccharomyces cerevisiae and Escherichia coli. J. Biol. Chem. 267, 8591–8598 (1992).
Lodge, J.K., Johnson, R.L., Weinberg, R.A. & Gordon, J.I. Comparison of myristoyl-CoA : protein N-myristoyl transferases from three pathogenic fungi: Cryptococcus neoformans, Histoplasma capsulatum, and Candida albicans. J. Biol. Chem. 269, 2996–3009 (1994).
Ntwasa, M., Egerton, M. & Gay, N.J. Sequence and expression of Drosophila myristoyl-CoA : protein N-myristoyl transferase: evidence for proteolytic processing and membrane localisation. J Cell. Sci. 110, 149–156 (1997).
Rudnick, D.A., Lu, T., Jackson-Machelski, E., Hernandez, J.C., Li, Q., Gokel, G.W. & Gordon, J.I. Analogs of palmitoyl-CoA that are substrates for myristoyl-CoA : protein N-myristoyl transferase. Proc. Natl. Acad. Sci. USA 89, 10507–10511 (1992).
Duronio, R.J., Rudnick, D.A., Adams, S.P., Towler, D.A. & Gordon, J.I. Analyzing the substrate specificity of Saccharomyces cerevisiae myristoyl-CoA: protein N-myristoyl transferase by co-expressing it with mammalian G protein alpha subunits in Escherichia coli. J. Biol. Chem. 266, 10498–10504 (1991).
McWherter, C.A. et al. Scanning alanine mutagenesis and de-peptidization of a Candida albicans myristoyl-CoA : protein N-myristoyl transferase octapeptide substrate reveals three elements critical for molecular recognition. J. Biol. Chem. 272, 11874–11880 (1997).
Langner, C.A. et al. 4-oxatetradecanoic acid is fungicidal for Cryptococcus neoformans and inhibits replication of human immunodeficiency virus I. J. Biol. Chem. 267, 17159–17169 (1992).
Weinberg, R.A., McWherter, C.A., Freeman, S.K., Wood, D.C., Gordon, J.I. & Lee, S.C. Genetic studies reveal that myristoyl-CoA : protein N-myristoyl transferase is an essential enzyme in Candida albicans. Mol. Microbiol. 16, 241–250 (1995).
Nagaragan, S.R. et al. Conformationally constrained [p-(omega-aminoalkyl)phenacetyl]-L-seryl-L-lysyl dipeptide amides as potent peptidomimetic inhibitors of Candida albicans and human myristoyl-CoA : protein N-myristoyl transferase. J. Med. Chem. 40, 1422–1438 (1997).
de La Fortelle, E. & Bricogne, G. Maximum-Likelihood Heavy-Atom Parameter Refinement in the MIR and MAD Methods. Meth. Enz. 276, 472–494 (1997).
Holm, L. & Sander, C. New Structure - Novel Fold? Structure 5, 165–171 (1997).
Kleywegt, G.J. & Jones, T.A. Detecting folding motifs and similarities in protein structures. Meth. Enz. 277, 525–545 (1997).
Murray-Rust, J. et al. Topological similarities in TGFβ2, PDGF-BB and NGF define a superfamily of polypeptide growth factors. Structure 1, 153–159 (1993).
Cameron, O.D., Olin, B., Ridderstrom, M., Mannervik, B. & Jones, T.A. Crystal structure of human glyoxalase. 1. Evidence for gene duplication and domain swapping. EMBO J. 16, 3386–3395 (1997).
Conti, E., Franks, N.P. & Brick, P. Crystal structure of luciferase throws light on a superfamily of adenylate-forming enzymes. Structure 4, 287–298 (1996).
Engel, C. & Wierenga, R. The diverse world of coenzyme A binding proteins. Curr. Opin. Struct. Biol. 6, 790–797 (1996).
Nicholls, A., Sharp, K. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Kleywegt, G.J. & Jones, T.A. Detection, Delineation, Measurement and Display of Cavities in Macromolecular Structures. Acta Crystallogr. D50, 178–185 (1994).
Bairoch, A., Bucher, P. & Hofmann, K. The PROSITE database, its status in 1997. Nucleic Acids Res. 25, 217–221 (1997).
Wierenga, R.K., Drenth, J. & Schultz, G.E. Comparison of the three-dimensional protein and nucleotide structure of the FAD-binding domain of p-hydroxybenzoate hydroxylase with the FAD- as well as NADPH-binding domains of glutathione reductase. J. Mol. Biol. 167, 725–739 (1983).
Rudnick, D.A. et al. Structural and functional studies of Saccharomyces cerevisiae myristoyl-CoA : protein N-myristoyl transferase produced in Escherichia coli. Evidence for an acyl-enzyme intermediate. J. Biol. Chem. 265, 13370–13378 (1990).
Quilin, M., Baase, W. & Matthews, B. Collected Abstracts of the XVII Congress and General Assembly of the International Union of Crystallography, Seattle (Wa), USA. (ed. J.F. Griffin) C-215 Acta Crystallogr. A52 (1996).
Schiltz, M., Fourme, R., Broutin, I. & Prange, T. The catalytic site of serine proteinases as a specific binding cavity for xenon. Structure 3, 309–316 (1995).
Schiltz, M., Prangé, T. & Fourme, R. On the preparation and X-ray data collection of isomorphous xenon derivatives. J. Appl. Cryst. 27, 950–960 (1994).
Rocque, W.J., McWherter, C.A., Wood, D.C. & Gordon, J.I. A comparative analysis of the kinetic mechanism and peptide substrate specificity of human and Saccharomyces cerevisiae myristoyl-CoA : protein N-myristoyl transferase.J. Biol. Chem. 268, 9964–9971 (1993).
Mcllhinney, R.A., Patel, P.B. & McGlone, K. Characterization of a polyhistidine-tagged form of human myristoyl-CoA: protein N-myristoyl transferase produced in Escherichia coli. Eur. J. Biochem. 222, 137–146 (1994).
Bhatnagar, R.S., Jackson-Machelski, E., McWherter, C.A. & Gordon, J.I. Isothermal titration calorimetric studies of Saccharomyces cerevisiae myristoyl-CoA : protein N-myristoyl transferase. Determinants of binding energy and catalytic discrimination among acyl-CoA and peptide ligands. J. Biol. Chem. 269, 11045–11053 (1994).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Leslie, A.G.W. Molecular Data Processing Crystallographic Computing 5 (Moras, D., Podjarny, A. D. & Thierry, J. C., eds.) (Oxford University Press; 1990).
Collaborative Computing Project No. 4: The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D50, 760–763 (1994).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Otwinowski, Z., In Proceedings of the CCP4 Study Weekend: Data Collection and Processing, 29–30 January 1993. (Compiled by: L. Sawyer, N. Isaacs and S. Bailey) 56–62 (SERC Daresbury Laboratory, England; 1993).
Stowell, M.H.B. et al. A simple device for studying macromolecular crystals under moderate gas pressures (0.1–10 MPa). J. Appl. Crystallogr. 29, 608–613 (1996).
Sheldrick, G.M. Phase annealing in SHELX-90: direct methods for larger structures. Acta Crystallogr. A46, 467–473 (1990).
Bricogne, G. A Bayesian statistical theory of the phase problem. I. A multichannel maximum-entropy formalism for constructing generalised joint probability distributions of structure factors. Acta Crystallogr. A44, 517–545 (1988).
Kleywegt, G.J. & Jones, T.A. Use of non-crystallographic symmetry in protein structure refinement. Acta Crystallogr. D52, 842–857 (1996).
Jones, T.A., Zou, J.-Y., Cowan, S.W. and Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A.T., Kuriyan, J., Karplus M. Crystallographic R-factor refinement by molecular dynamics. Science 235, 458–460 (1988).
Brünger, A.T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Tronrud, D.E., Ten Eyck, L.F. & Matthews, B.W. An efficient general-purpose least-squares refinement program for macromolecular structures. Acta Crystallogr. A43, 489–501 (1987).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Weston, S., Camble, R., Colls, J. et al. Crystal structure of the anti-fungal target N-myristoyl transferase. Nat Struct Mol Biol 5, 213–221 (1998). https://doi.org/10.1038/nsb0398-213
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0398-213
This article is cited by
-
NMT1 and NMT2 are lysine myristoyltransferases regulating the ARF6 GTPase cycle
Nature Communications (2020)
-
High-resolution snapshots of human N-myristoyltransferase in action illuminate a mechanism promoting N-terminal Lys and Gly myristoylation
Nature Communications (2020)
-
A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain glycoside hydrolase family 3 β-N-acetylglucosaminidase
Scientific Reports (2015)
-
Protein myristoylation in health and disease
Journal of Chemical Biology (2010)
-
Homology modeling and molecular dynamics simulation of N-myristoyltransferase from protozoan parasites: active site characterization and insights into rational inhibitor design
Journal of Computer-Aided Molecular Design (2009)