Abstract
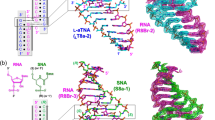
Here we report the 1.8Å X–ray structure of a 2:1 drug–DNA complex between distamycin A and an alternating B–DNA octamer duplex d(ICICICIC)2. The two distamycin A molecules are bound side by side with dyad symmetry in an antiparallel orientation in the expanded minor groove. The amides of each drug molecule are hydrogen bonded to the minor groove base atoms of only one DNA strand. The complex not only shows binding of two drug molecules, but the DNA duplex also exhibits striking low–high alternations in the helical twist angles, the sugar puckering and the phosphate conformations, providing the basis for a new model for an alternating B–DNA with a dinucleotide repeat.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wang, A.H.-J. & Teng, M. Crystallographic and Modeling Methods in Molecular Design. (ed. Bugg, C.E. & Ealick, S.E.) 123–150 (Springer-Verlag, New York, 1990).
Mitsuya, H. and Broder, S. Strategies for antiviral therapy in AIDS. Nature 325, 773–778 (1987).
Portugal, J. & Waring, M.J. Comparison of binding sites in DNA for berenil netropsin and distamycin. A footprint study. Eur. J. Biochem. 167, 281–289 (1987).
Pelton, J.G. & Wemmer, D.E. Structural modeling of the distamycin A-d(CGCGAATTCGCG)2 complex using 2D NMR and molecular mechanics. Biochemistry 27, 8088–8096 (1988).
Patel, D.J. & Shapiro, L. Molecular recognition in noncovalent antitumur agent-DNA complexes: NMR studies of the base and sequence dependent recognition of the DNA minor groove by netropsin. Biochimie 67, 887 (1985).
Coll, M., Frederick, C.A., Wang, A.H.-J. & Rich, A. A bifurcated hydrogen-bonded conformation in the d(A·T) base pair of the DNA dodecamer d(CGCAAATTTGCG) and its complex with distamycin. Proc. natn. Acad. Sci. U.S.A. 84, 8385–8389 (1987).
Kopka, M.L., Yoon, C., Goodsell, D., Pjura, P. and Dickerson, R.E., The Binding of an antitumur drug to DNA. Netropsin and CGCGAATTBrCGCG. J. molec. Biol. 183, 553–563 (1985).
Coll, M., Aymami, J., van der Marel, G.A., van Boom, J.H., Rick, A. & Wang, A.H.-J. Molecular structure of the netropsin-d(CGCGATATCGCG) complex: DNA conformation in an alternating AT segment. Biochemistry 28, 310–320 (1989).
Sriram, M., van der Marel, G.A., Roelen, H.L.R.F., van Boom, J.H. & Wang, A.H.-J. Structural consequences of a carcinogenic alkylation lesion on DNA: effect of O6-ethylguanine on the molecular structure of the d(CGC[e6G]AATTCGCG)-netropsin complex. Biochemistry 31, 11823–11834 (1992).
Tabernero, L., Verdaguer, N., Coll, M., Fita, I., van der Marel, G.A., van Boom, J.H., Rich, A. & Aymami, J. Molecular structure of the A-tract DNA dodecamer d(CGCAAATTTGCG) complexed with the minor groove binding drug netropsin. Biochemistry 32, 8403–8410 (1993).
Balendiran, K., Rao, S.T., Sekharudu, C.Y., Zon, G. and Sundaralingam, M. X-ray structure of the dodecamer d(CGCGTTAACGCG) and its netropsin complex. Helical parameters from three methods vary. Biochemistry (in the press).
Kopka, M.L. & Larsen, T.A. Netropsin and the lexitropsins. The search for sequence-specific minor groove-binding ligands in Nucleic Acid Targeted Drug Design (ed. Propst, C.L., & Perun, T.J.) 303–374 (Marcel Dekker, New York, 1992).
Pelton, J.G. & Wemmer, D.E. Structural characterization of a 2:1 distamycin A-d(CGCAAATTGGC) complex by two dimensional NMR. Proc. natn. Acad. Sci. U.S.A. 86, 5723–5727 (1989).
Pelton, J.G. & Wemmer, D.E. Binding modes of distamycin A with d(CGCAAATTTGCG)2 determined by two-dimensional NMR. J. Am. Chem. Soc. 112, 1393–1399 (1990).
Mrksich, M., Wade, W.S., Dwyer, T.J., Geierstanger, B.H., Wemmer, D.E. & Dervan, P.B. Antiparallel side-by-side dimeric motif for sequence-specific recognition in the minor groove of DNA by the designed peptide 1-methylimidazole-2-carbonamide netropsin. Proc. natn. Acad. Sci. U.S.A. 89, 7586–7590 (1992).
Wade, W.S., Mrksich, M. & Dervan, P.B. Design of pepetides that bind in the minor groove of DNA at 5′-(A,T)G(A,T)C(A,T)-3′ sequences by a dimeric side-by-side motif. J. Am. Chem. Soc. 114, 8783–8794 (1992).
Leslie, A.G.W, Arnott, S., Chandrasekaran, R. & Ratliff, R.L. Polymorphism of DNA double helices. J. molec. Biol. 143, 49–72 (1980).
Prive, G.G., Heinemann, U., Chandrasegaran, S., Kan, L.-S., Kopka, M.L. & Dickerson, R.E. Helix geometry, hydration, and G·A mismatch in a B-DNA decamer. Science 238, 498–504 (1987).
Cruse, W.B.T., Salisbury, S.A., Brown, T., Cosstick, R., Eckstein, F. & Kennard, O. Chiral phosphorothioate analogues of B-DNA. The crystal structure of Rp-d(Gp(S)CpGp(S)CpGp(S)C). J. molec. Biol. 192, 891–905 (1986).
Yoon, C, Prive, G.G., Goodsell, D.S. & Dickerson, R.E. Structure of an alternating-B DNA helix and its relationship to A-truct DNA. Proc. natn. Acad. Sci. U.S.A. 85, 6332–6336 (1988).
Altona, C. and Sundaralingam, M. Conformational analysis of the sugar ring in nucleotides and nulcleosides. A new description using the concept of pseudorotation. J. Am. Chem. Soc. 94, 8205–8211 (1972).
Fratini, A.V., Kopka, M.L., Drew, H.R. & Dickerson, R.E. Reversible bending and helix geometry in a B-DNA dodecamer: CGCGAATTBrCGCG. J. biol. Chem. 257, 14686–14707 (1982).
Bingman, C., Li, X., Zon, G. & Sundaralingam, M. Crystal and molecular structure of d(GTGCGCAC): investigation of the effects of base sequence on the conformation of octamer duplexes. Biochemistry 31, 12803–12812 (1992).
Bingman, C, Jain, S., Zon, G. & Sundaralingam, M. Crystal and molecular structure of the alternating dodecamer d(GCGTACGTACGC) in the A-DNA form: comparison with the isomorphous non-alternating dodecamer d(CCGTACGTACGG). Nucleic Acids Res. 20, 6637–6647 (1992).
Thota, N., Li, X., Bingman, C. & Sundaralingam, M. High resolution refinement of the hexagonal A-DNA octamer d(GTGTACAC) at 1.4Å. J. Acta Crystallogr. D49, 282–291 (1993).
Mitsui, Y. et al. Physical and Enzymatic studies on polyd(l-C)·poly d(l-C), an unusual double-helical DNA. Nature 288, 1166–1169 (1970).
Ramakrishnan, B. and Sundaralingam, M. High resolution crystal structure of the A-DNA decamer d(CCCGGCCGGG). Novel intermolecular base-paired G*(G·C) triplets. J. molec. Biol. 231, 431–444 (1993).
Takusagawa, F. The crystal structure of d(GTACGTAC) at 2.25Å resolution: Are the A-DNA's always unwounded approximately 10° at the C-G steps? J. Biomol. struc. dynamics 7, 795–809 (1990).
Prive, G.G., Yanagi, K. & Dickerson, R.E. Structure of the B-DNA decamer CCAACGTTGG and comparison with isomorphous decamers CCAAGATTGG and CCAGGCCTGG. J molec. Biol. 217, 177–199 (1991).
Lutter, L.C. Deoxyribonuclease I produces staggered cuts in the DNA of chromatin. J. molec. Biol. 117, 53–69 (1977).
Laskowski, M. Sr, Deoxyribonuclease I in The Enzymes Vol. IV, 3rd edit. (ed. Boyer, P.O.), 289–311 (Academic Press, London, 1971)
Scheffler, I.E., Elson, E.L. & Baldwin, R.L. Helix formation by dAT oligomers, I. Hairpin and straight-chain helices. J. moec. Biol. 36, 291–304 (1968).
Klug, A., Jack, A., Viswamitra, M.A., Kennard, O., Shakked, Z. & Steitz, T.A. A hypothesis on a specific sequence-dependent conformation of DNA and its relation to the binding of the lac-repressor protein. J. molec Biol. 131, 669–680 (1979).
Jones, T.A. Interactive computer graphics: FRODO in Methods Enz. 115, 157–171 (1985).
Brunger, A.T. XPLOR Manual, Version 3.0 (Yale University, New Haven, 1992).
Bernstein, F.C. et al. The protein data bank: a computer-based archieval file for macromolecule structures. J. molec. Biol. 112, 535–542 (1977).
Sundaralingam, M. Stereochemistry of nucleic acids and their constituents. IV. Allowed and preferred conformations of nucleosides, nucleoside mono-, di, tri-, tetraphosphates, nucleic acids and polynucleotides. Biopolymers 7, 821–860 (1969).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Chen, X., Ramakrishnan, B., Rao, S. et al. Binding of two distamycin A molecules in the minor groove of an alternating B–DNA duplex. Nat Struct Mol Biol 1, 169–175 (1994). https://doi.org/10.1038/nsb0394-169
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0394-169
This article is cited by
-
DNA-binding and antiviral activity of bis-netropsins containing clusters of lysine residues in the N-terminal region
Doklady Biochemistry and Biophysics (2004)
-
Interaction of the human topoisomerase I-DNA complex with oligo-1,3-thiazolecarboxamides and their oligonucleotide conjugates
Molecular Biology (2000)
-
Structural basis for G•C recognition in the DNA minor groove
Nature Structural Biology (1998)
-
Extending the recognition site of designed minor groove binding molecules
Nature Structural & Molecular Biology (1996)
-
Crystal structures of B-form DNA–RNA chimers complexed with distamycin
Nature Structural & Molecular Biology (1995)