Abstract
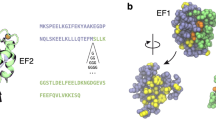
The cooperative binding of Ca2+ ions is an essential functional property of the EF-hand family of Ca2+-binding proteins. To understand how these proteins function, it is essential to characterize intermediate binding states in addition to the apo- and holo-proteins. The three-dimensional solution structure and fast time scale internal motional dynamics of the backbone have been determined for the half-saturated state of the N56A mutant of calbindin D9k with Ca2+ bound only in the N-terminal site. The extent of conformational reorganization and a loss of flexibility in the C-terminal EF-hand upon binding of an ion in the N-terminal EF-hand provide clear evidence of the importance of site–site interactions in this family of proteins, and demonstrates the strength of long-range effects in the cooperative EF-hand Ca2+-binding domain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Forsén, S., Kördel, J., Grundström, T. & Chazin, W.J. The molecular anatomy of a calcium-binding protein. Acc. Chem. Res. 26, 7–14 ( 1993).
Linse, S. et al. Electrostatic contributions to the binding of Ca2+ in calbindin D9k . Biochemistry 30, 154–162 (1991).
Linse, S., Jonsson, B. & Chazin, W.J. The effect of protein concentration on ion binding . Proc. Natl. Acad. Sci. USA 92, 4748– 4752 (1995).
Akke, M., Forsén, S. & Chazin, W.J. Molecular basis for co-operativity in Ca2+ binding to calbindin D9k. 1H nuclear magnetic resonance studies of (Cd2+)1-bovine calbindin D 9k . J. Mol. Biol. 220, 173– 189 (1991).
Akke, M., Forsén, S. & Chazin, W.J. Solution structure of (Cd2+)1-calbindin D9k reveals details of the stepwise structural changes along the Apo → (Ca2+)II 1 → (Ca2+)I,II 2 binding pathway. J. Mol. Biol. 252, 102– 121 (1995).
Carlström, G. & Chazin, W.J. Two-dimensional 1H nuclear magnetic resonance studies of the half-saturated (Ca2+)1 state of calbindin D9k. Further implications for the molecular basis of cooperative Ca2+ binding. J. Mol. Biol. 231, 415–430 (1993).
Wimberly, B., Thulin, E. & Chazin, W.J. Characterization of the N-terminal half-saturated state of calbindin D9k: NMR studies of the N56A mutant. Protein Sci. 4, 1045–1055 (1995).
Skelton, N.J., Kördel, J., Akke, M. & Chazin, W.J. Nuclear magnetic resonance studies of the internal dynamics in Apo (Cd2+)1 and (Ca2+)2 calbindin D 9k. The rates of amide proton exchange with solvent. J. Mol. Biol. 227, 1100–1117 ( 1992).
Akke, M., Skelton, N.J., Kordel, J., Palmer, A.G. III, & Chazin, W.J. Effects of ion binding on the backbone dynamics of calbindin D9k determined by 15N NMR relaxation. Biochemistry 32, 9832–9844 (1993).
Akke, M., Brüschweiler, R., Palmer, A.G. III. NMR order parameters and free energy: an analytical approach and its application to cooperative Ca2+ binding by calbindin D9k . J. Am. Chem. Soc. 115, 9832–9833 (1993).
Spassov, V. & Bashford, D. Electrostatic coupling to pH-titrating sites as a source of cooperativity in protein–ligand binding. Protein Sci. 7, 2012–2025 (1998).
Linse, S. & Chazin, W.J. Quantitative measurements of the cooperativity in an EF-hand protein with sequential calcium binding. Protein Sci. 4, 1038–1044 (1995).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures . J. Appl. Crystallogr. 26, 283– 291 (1993).
Kördel, J., Skelton, N.J., Akke, M. & Chazin, W.J. High-resolution structure of calcium-loaded calbindin D9k . J. Mol. Biol. 231, 711–734 ( 1993).
Skelton, N.J., Kördel, J. & Chazin, W.J. Determination of the solution structure of Apo calbindin D9k by NMR spectroscopy. J. Mol. Biol. 249 , 441–462 (1995).
Kroenke, C.D., Loria, J.P., Lee, L.K., Rance, M., & Palmer, A.G. III., Longitudinal and transverse 1H-15N dipolar/15N chemical shift anisotropy relaxation interference: unambiguous determination of rotational diffusion tensors and chemical exchange effects in biological macromolecules. J. Am. Chem. Soc. 120, 7905–7915 (1998).
Spyracopoulos, L., Gagne, S.M., Li, M.X. & Sykes, B.D. Dynamics and thermodynamics of the regulatory domain of human cardiac troponin C in the apo- and calcium-saturated states. Biochemistry 37, 18032– 18044 (1998).
Lipari, G. & Szabo, A. Model-free approach to the interpretation of nuclear magnetic resonance relaxation in macromolecules. 1. Theory and range of validity. J. Am. Chem. Soc. 104, 4546–4559 (1982).
Clore, G.M. et al. Deviations from simple two-parameter model-free approach to the interpretation of nitrogen-15 nuclear magnetic relaxation of proteins . J. Am. Chem. Soc. 112, 4989– 4991 (1990).
Yang, D. & Kay, L.E. Contributions to conformational entropy arising from bond vector fluctuations measured from NMR-derived order parameters: application to protein folding. J. Mol. Biol. 263, 369–382 (1996).
Li, Z., Raychaudhuri, S. & Wand, A.J. Insights into the local residual entropy of proteins provided by NMR relaxation. Protein Sci. 5, 2647–2650 (1996).
Bracken, C., Carr, P.A., Cavanagh, J., & Palmer, A.G., III . Temperature dependence of intramolecular dynamics of the basic leucine zipper of GCN4: implications for the entropy of association with DNA. J. Mol. Biol. 285, 2133–2146 (1999).
Gagne, S.M., Tsuda, S., Spyracopoulos, L., Kay, L.E. & Sykes, B.D. Backbone and methyl dynamics of the regulatory domain of troponin C: anisotropic rotational diffusion and contribution of conformational entropy to calcium affinity. J. Mol. Biol. 278, 667–686 (1998).
Kay, L.E., Muhandiram, D.R., Wolf, G., Shoelson, S.E. & Forman-Kay, J.D. Correlation between binding and dynamics at SH2 domain interfaces. Nature Struct. Biol. 5, 156–163 (1998).
Fisher, M.W.F., Majumdar, A. & Zuiderweg, E.R.P. Protein NMR relaxation: theory, applications and outlook. Prog. NMR Spectrosc. 33, 207– 272 (1998).
Cavanagh, J., Palmer, III,A.G., Fairbrother, W. & Skelton, N.J. Protein NMR spectroscopy. Principles and practice. (Academic Press, San Diego, California; 1996).
Skelton, N.J. et al. Practical aspects of two-dimensional proton-detected 15N spin relaxation measurements. J. Magn. Reson. 102, 253–264 (1993).
Gippert, G. New Computational Methods for 3D NMR Data Analysis and Protein Structure Determination in High-Dimensional Internal Coordinate Space. Ph.D. thesis, The Scripps Research Institute (1995).
Mäler, L., Potts, B.C. & Chazin, W.J. High resolution solution structure of apo calcyclin and structural variations in the S100 family of calcium-binding proteins. J. Biomol. NMR 13, 233–247 (1999).
Güntert, P., Braun, W. & Wüthrich, K. Efficient computation of three-dimensional protein structures in solution from nuclear magnetic resonance data using the program DIANA and the supporting programs CALIBA, HABAS and GLOMSA. J. Mol. Biol. 217, 517–530 ( 1991).
Güntert, P. & Wüthrich, K. Improved efficiency of protein structure calculations from NMR data using the program DIANA with redundant dihedral angle constraints. J. Biomol. NMR 1, 447–456 ( 1991).
Pearlman, D.A. et al. AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comp. Phys. Commun. 91, 1–41 (1995).
Smith, J.A., Gomez-Paloma, L., Case, D.A. & Chazin, W.J. Molecular dynamics docking driven by NMR-derived restraints to determine the structure of the calicheamicin gamma(I) oligosaccharide domain complexed to duplex DNA. Magn. Reson. Chem. 34, 147– 155 (1996).
Mandel, A.M., Akke, M., & Palmer, A.G., III. Backbone dynamics of Escherichia coli ribonuclease HI: correlations with structure and function in an active enzyme. J. Mol. Biol. 246, 144– 163 (1995).
Acknowledgements
This research was supported by grants from the National Institutes of Health to W.J.C. and to M.R., and a postdoctoral fellowship (to L.M.) from the Swedish National Science Research Council. We thank J. Chung for assistance with NMR experiments and A.G. Palmer, III, for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mäler, L., Blankenship, J., Rance, M. et al. Site–site communication in the EF-hand Ca2+-binding protein calbindin D9k. Nat Struct Mol Biol 7, 245–250 (2000). https://doi.org/10.1038/73369
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/73369
This article is cited by
-
IP3 receptor signaling and endothelial barrier function
Cellular and Molecular Life Sciences (2017)
-
NMR order parameters calculated in an expanding reference frame: identifying sites of short- and long-range motion
Journal of Biomolecular NMR (2011)
-
Protein proton–proton dynamics from amide proton spin flip rates
Journal of Biomolecular NMR (2009)
-
Dynamically driven protein allostery
Nature Structural & Molecular Biology (2006)
-
EF-hand protein dynamics and evolution of calcium signal transduction: an NMR view
JBIC Journal of Biological Inorganic Chemistry (2006)