Abstract
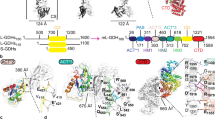
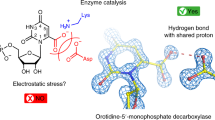
Adjacent cysteine residues can only form disulphide bridges in a distorted structure containing a cis–peptide link. Such bridges are extremely uncommon, identified so far in the acetyl choline receptor alone where the structure of the bridge is undetermined. Here we present the first molecular description of a disulphide bridge of this type in the quinoprotein methanol dehydrogenase from Methylobacterium extorquens. We show that this structure occurs in close proximity to the pyrrolo–quinoline quinone prosthetic group and a calcium ion in the active site of the enzyme. This unusual disulphide bridge appears to play a role in the electron transfer reaction mediated by methanol dehydrogenase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anthony, C. The bacterial oxidation of methane and methanol. Adv. microb. Physiol. 27, 113–210 (1986).
Anthony, C. Methanol dehydrogenase in gram-negative bacteria. in Principles and applications of quinoproteins (ed. Davidson, V.L.) 17–45 (Marcel Dekker, New York, 1993).
Anthony, C. The c-type cytochromes of methylotrophic bacteria. Biochim. biophys. Acta 1099, 1–15 (1992).
Anthony, C. & Zatman, L.J. The microbial oxidation of methanol: The prosthetic group of alcohol dehydrogenase of Pseudomonas sp.M27: A newoxidoreductase prosthetic group. Biochem. J. 104, 960–969 (1967).
Salisbury, S.A., Forrest, H.S., Cruse, W.B.T. & Kennard, O. A novel coenzyme from bacterial primary alcohol dehydrogenases. Nature 280, 843–844 (1979).
Duine, J.A., Frank, J. & Verwiel, P.E.J. Structure and activity of the prosthetic group of methanol dehydrogenase. Eur. J. Biochem. 108, 187–192 (1980).
Nunn, D.N., Day, D.J. & Anthony, C. The second subunit of methanol dehydrogenase of Methylobacterium extorquens AM1. Biochem. J. 260, 857–862 (1989).
Richardson, I.W. & Anthony, C. Characterization of mutant forms of the quinoprotein methanol dehydrogenase lacking an essential calcium ion. Biochem. J. 287, 709–715 (1992).
Anderson, D.J., Morris, C.J., Nunn, D.N., Anthony, C. & Lidstrom, M.E. Nucleotide sequence of the Methylobacterium extorquens AM1 moxF and moxJ genes involved in methanol oxidation. Gene. 90, 173–176 (1990).
Ramachandran, G.N. & Sasisekharan, V. Conformation of polypeptides and proteins. Adv. Protein Chem. 23, 283–437 (1968).
Chandrasekaran, R. & Balasubramanian, R. Stereochemical studies of cyclic peptides. VI. Energy calculations of the cyclic dipeptide cysteinyl-cysteine. Biochim. biophys. Acta 188, 1–9 (1969).
Capasso, S., Mattia, C. & Mazzarella, L. Structure of a cis-peptide unit: molecular conformation of the cyclic disulphide L-cysteinyl-L-cysteine. Acta Crystallog. B33, 2080–2083 (1977).
Mez, H.-C. Cyclo-L-cystine acetic acid. Cryst. Struct. Comm. 3, 657–660 (1993).
Mosckovitz, R. & Gershoni, J.M. Three possible disulphides in the acetylcholine receptor α-subunit. J. biol. Chem. 263, 1017–1022 (1988).
Ghosh, M., Harlos, K., Blake, C.C.F., Richardson, I. & Anthony, C. Crystallisation and preliminary crystallographic investigation of methanol dehydrogenase from Methylobacterium extorquens AM1. J. molec. Biol. 228, 302–305 (1992).
Adachi, O., Matsushita, K., Shinagawa, E. & Ameyama, M. Calcium in quinoprotein methanol dehydrogenase can be replaced by strontium. Agric. Biol. Chem. 54, 2833–2837 (1990).
Duine, J.A., Frank, J. & Jongejaw, J.A. Enzymology of quinoproteins. Adv. Enzym. 59, 169–212 (1987).
Frank, J. et al. On the mechanism of inhibition of methanol dehyrogenase by cyclopropane-derived inhibitors. Eur. J. Biochem. 184, 187–195 (1989).
Frank, J., Dijkstra, M., Balny, C., Verwiel, P.E.J. and Duine, J.A. Methanol dehydrogenase: mechanism of action. In PQQ and Quinoproteins (eds Jongejan, J.A. & Duine, J.A.) 13–22 (Kluwer Academic Publishers, Dordrecht, 1989).
Xia, X.A. et al. The 3-dimensional structures of methanol dehydrogenase from 2 methylotrophic bacteria at 2.6 Ångstrom resolution. J. biol. Chem. 267, 22289–22297 (1992).
Torchinsky, Y.M., Wittenberg, W. and Metzler, D. Methods for the quantitative determination of SH and S-S groups in proteins. In: Sulphur in Proteins, 113–132 (Pergamon, Oxford, 1981).
Day, D.J. & Anthony, C. Methanol dehydrogenase from Methylobacterium exorquens AM1. Meth. Enzym., 188, 210–216 (1990).
Jones, T.A. Interactive computer graphics: FRODO. Meth. Enzym. 115, 157–171 (1985).
Jones, T.A., Zuo, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building models in electron density maps and the location of errors in these models. Acta Crystallog. A47, 110–119 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Blake, C., Ghosh, M., Harlos, K. et al. The active site of methanol dehydrogenase contains a disulphide bridge between adjacent cysteine residues. Nat Struct Mol Biol 1, 102–105 (1994). https://doi.org/10.1038/nsb0294-102
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0294-102
This article is cited by
-
Bioinorganic insights of the PQQ-dependent alcohol dehydrogenases
JBIC Journal of Biological Inorganic Chemistry (2021)
-
The crystal structure of methanol dehydrogenase, a quinoprotein from the marine methylotrophic bacterium Methylophaga aminisulfidivorans MPT
Journal of Microbiology (2018)
-
PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference
Applied Microbiology and Biotechnology (2014)