Abstract
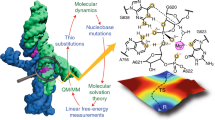
Helix packing is critical for RNA tertiary structure formation, although the rules for helix–helix association within structured RNAs are largely unknown. Docking of the substrate helix into the active site of the Tetrahymena group I ribozyme provides a model system to study this question. Using a novel chemogenetic method to analyze RNA structure in atomic detail, we report that complementary sets of noncanonical base pairs (a G·U wobble pair and two consecutively stacked sheared A·A pairs) create an RNA helix packing motif that is essential for 5′-splice site selection in the group I intron. This is likely to be a general motif for helix–helix interaction within the tertiary structures of many large RNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Strobel, S.A. & Doudna, J.A. RNA Seeing Double: Close-Packing of Helices in RNA Tertiary Structure. Trends Biochem. Sci. 22, 262–266 (1997).
Gutell, R.R., Larsen, N. & Woese, C.R. Lessons from an Evolving rRNA: 16S and 23S rRNA Structures from a Comparative Perspective. Microbiol. Rev. 58, 10–26 (1994).
Michel, F. & Westhof, E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J. Mol. Biol. 216, 585–610 (1990).
Strobel, S.A. & Cech, T.R. Minor groove recognition of the conserved G·U pair at the Tetrahymena ribozyme reaction site. Science 267, 675–679 (1995).
Waring, R.B., Towner, P., Minter, S.J. & Davies, R.W. Splice-site selection by a self-splicing RNA of Tetrahymena. Nature 321, 133–139 (1986).
Been, M.D. & Cech, T.R. One binding site determines sequence specificity of tetrahymena pre-rRNA self-splicing, trans-splicing, and RNA enzyme activity. Cell 47, 207–216 (1986).
Zaug, A.J., Been, M.D. & Cech, T.R., The Tetrahymena ribozyme acts like an RNA restriction endonuclease. Nature 324, 429–433 (1986).
Herschlag, D. Evidence for processivity and two-step binding of the RNA substrate from studies of J1/2 mutants of the Tetrahymena ribozyme. Biochemistry 31, 1386–1399 (1992).
Bevilacqua, P.C., Kierzek, R., Johnson, K.A. & Turner, D.H. Dynamics of ribozyme binding of substrate revealed by fluorescence-detected stopped-flow methods. Science 258, 1355–1358 (1992).
Strobel, S.A. & Cech, T.R. Translocation of an RNA duplex on a ribozyme. Nature Struct. Biol. 1, 13–17 (1994).
Murphy, F.L. & Cech, T.R. Alteration of substrate specificity for the endoribonucleoytic cleavage of RNA by the Tetrahymena ribozyme. Proc. Natl. Acad. Sci. U.S.A. 86, 9218–9222 (1989).
Doudna, J.A. & Szostak, J.W. RNA-catalysed synthesis of complementary-strand RNA. Nature 339, 519–522 (1989).
Strobel, S.A. & Cech, T.R. Exocyclic amine of the conserved G·U pair at the cleavage site of the Tetrahymena ribozyme contributes to 5′ -splice site selection and transition state stabilization. Biochemistry 35, 1201–1211 (1996).
Strobel, S.A. & Cech, T.R. Tertiary interactions with the internal guide sequence mediate docking of the P1 helix into the catalytic core of the Tetrahymena ribozyme. Biochemistry 32, 13593–13604 (1993).
Wang, J.-F., Downs, W.D. & Cech, T.R. Movement of the guide sequence during RNA catalysis by a group I ribozyme. Science 260, 504–508 (1993).
Cate, J.H. et al. Crystal structure of a group I ribozyme domain: principles of RNA packing. Science 273, 1678–1685 (1996).
Strobel, S.A. & Shetty, K. Defining the chemical groups essential for Tetrahymena group I intron function by nucleotide analog interference mapping. Proc. Natl. Acad. Sci. USA 94, 2903–2908 (1997).
Pyle, A.M., Murphy, F.L. & Cech, T.R. RNA substrate binding site in the catalytic core of the Tetrahymena ribozyme. Nature 358, 123–128 (1992).
Gaur, R.K. & Krupp, G. Modification interference approach to detect ribose moieties important for the optimal activity of a ribozyme. Nucleic Acids Res. 21, 21–26 (1993).
Conrad, F., Hanne, A., Gaur, R.K. & Krupp, G. Enzymatic synthesis of 2′ -modified nucleic acids: identification of important phosphate and ribose moieties in RNase P substrates. Nucleic Acids Res. 23, 1845–1853 (1995).
Eckstein, F. Nucleoside phosphorothioates. Ann. Rev. Biochem. 54, 367–402 (1985).
Arabshahi, A. & Frey, P.A. A simplified procedure for synthesizing nucleoside 1-thiotriphosphates: dATPαS, dGTPαS, UTPαS, and dTTPαS. Biochem. Biophys. Res. Com. 204, 150–155 (1994).
Beaudry, A.A. & Joyce, G.F. Directed evolution of an RNA enzyme. Science 257, 635–641 (1992).
Sousa, R. & Padilla, R. A mutant T7 RNA polymerase as a DNA polymerase. EMBO J. 14, 4609–4621 (1995).
Mei, R. & Herschlag, D. Mechanistic investigations of a ribozyme derived from the Tetrahymena group I intron. Insights into catalysis and the second step of self-splicing. Biochemistry 35, 5796–5809 (1996).
Gish, G. & Eckstein, F. DNA and RNA sequence determination based on phosphorothioate chemistry. Science 240, 1520–1522 (1988).
SantaLucia, J. & Turner, D.H. Structure of (rGGCGAGCC)2 in solution from NMR and restrained moleuclar dynamics. Biochemistry 32, 12612–12623 (1993).
Damberger, S.H. & Gutell, R.R. A comparative database of group I intron structures. Nucleic Acids Res. 22, 3508–3510 (1994).
Allain, F.H.T. & Varani, G. Structure of the P1 helix from group I self-splicing introns. J. Mol. Biol. 250, 333–353 (1995).
Pyle, A.M. et al. Replacement of the Conserved G·U with a G-C pair at the cleavage site of the Tetrahymena ribozyme decreases binding, reactivity, and fidelity. Biochemistry 33, 13856–13863 (1994).
Knitt, D.S., Narlikar, G.J. & Herschlag, D. Dissection of the role of the conserved G·U pair in group I RNA self-splicing. Biochemistry 33, 13864–13879 (1994).
Doudna, J.A., Cormack, B.P. & Szostak, J.W. RNA structure, not sequence, determines the 5′ splice-site specificity of a group I intron. Proc. Natl. Acad. Sci. USA 86, 7402–7406 (1989).
Gautheret, D., Konings, D. & Gutell, R.R. A major family of motifs involving G·A mismatches in ribosomal RNA. J. Mol. Biol. 242, 1–8 (1994).
SantaLucia, J., Kierzek, R. & Turner, D.H. Effects of GA mismatches on the structure and thermodynamics of RNA internal loops. Biochemistry 29, 8813–8819 (1990).
Gautheret, D., Konings, D. & Gutell, R.R. G·U base pairing motifs in ribosomal RNA. RNA 1, 807–814 (1995).
Murphy, F.L. & Cech, T.R. GAAA tetraloop and conserved bulge stabilize tertiary structure of a group I intron domain. J. Mol. Biol. 236, 49–63 (1994).
Pley, H.W., Flaherty, K.M. & McKay, D.B. Model for an RNA tertiary interaction from the structure of an intermolecular complex between a GAAA tetraloop and an RNA helix. Nature 372, 111–113 (1994).
Tanner, M.A. & Cech, T.R. Activity and thermostability of the small self-splicing group I intron in the pre-tRNAIle of the purple bacterium Azoarcus. RNA 2, 74–83 (1996).
Costa, M. & Michel, F. Frequent use of the same tertiary motif by self-folding RNAs. EMBO J. 14, 1276–1285 (1995).
Christian, E.L. & Yarus, M. Metal coordination sites that contribute to structure and catalysis in the group I intron from Tetrahymena. Biochemistry 32, 4475–4480 (1993).
Carson, M. RIBBONS 2.0 Manual (University of Alabama at Birmingham, Alabama, USA; 1991).
Nicholls, A., GRASP: graphical respresentation and analysis of surface properties (Columbia University, New York, USA; 1993).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Strobel, S., Ortoleva-Donnelly, L., Ryder, S. et al. Complementary sets of noncanonical base pairs mediate RNA helix packing in the group I intron active site. Nat Struct Mol Biol 5, 60–66 (1998). https://doi.org/10.1038/nsb0198-60
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0198-60
This article is cited by
-
Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing
Nature Catalysis (2023)
-
Backbone and nucleobase contacts to glucosamine-6-phosphate in the glmS ribozyme
Nature Structural & Molecular Biology (2006)
-
Crystal structure of a self-splicing group I intron with both exons
Nature (2004)
-
Non-Watson Crick base pairs might stabilize RNA structural motifs in ribozymes — a comparative study of group-I intron structures
Journal of Biosciences (2003)
-
A tertiary interaction that links active-site domains to the 5′ splice site of a group II intron
Nature (2000)