Abstract
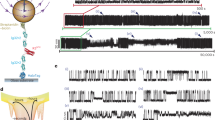
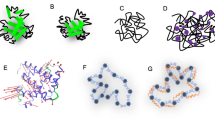
A novel off-resonance rotating-frame 15N NMR spin relaxation experiment is used to characterize conformational fluctuations with correlation times between 32 and 175 μs in the third fibronectin type III domain of human tenascin-C. Conformational fluctuations of contiguous regions of the β-sandwich structure of the type III domain may represent collective motions, such as transient twisting or breathing of the β-sheets. Flexibility of the loop containing the Arg-Gly-Asp (RGD) tripeptide may affect the accessibility of this motif in protein-protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Frauenfelder, H., Parak, F. & Young, R.D. Conformational substates in proteins. Annu. Rev. Biophys. Biophys. Chem. 17, 451–479 (1988).
Creighton, T.E. . Proteins. Structures and Molecular Properties 1–507 (W. H. Freeman & Co., New York; 1993).
Weber, G. Protein interactions 1–293 (Chapman and Hall, New-York; 1992).
Akke, M. & Palmer, A.G. Monitoring macromolecular motions on microsecond-millisecond time scales by R1ρ–R1 constant-relaxation-time NMR spectroscopy. J. Am. Chem. Soc. 118, 911–912 (1996).
Feher, V.A., Baldwin, E.P. & Dahlquist, F.W. Access of ligands to cavities within the core of a protein is rapid. Nature Struct. Biol. 3, 516–521 (1996).
Denisov, V.P., Peters, J., Hörlein, H.D. & Halle, B. Using buried water molecules to explore the energy landscape of proteins. Nature Struct. Biol. 3, 505–509 (1996).
Wagner, G. Characterization of the distribution of internal motions in the basic pancreatic trypsin inhibitor using a large number of internal NMR probes. Q. Rev. Biophys. 16, 1–57 (1983).
Szyperski, T., Luginbühl, P., Otting, G., Güntert, P. & Wüthrich, K. Protein dynamics studied by rotating frame 15N spin relaxation times. J. Biomol. NMR 3, 151–164 (1993).
Peng, J. & Wagner, G. Frequency spectrum of NH bonds in eglin c from spectral density mapping at multiple fields. Biochemistry 34, 16733–16752 (1995).
Phan, I.Q.H., Boyd, J. & Campbell, I.D. Dynamic studies of a fibronectin type I module pair at three frequencies: Anisotropic modelling and direct determination of Conformational exchange. J. Biomol. NMR 8, 369–378 (1996).
Wyss, D.F., Dayie, K.T. & Wagner, G. The counterreceptor binding site of human CD2 exhibits an extended surface patch with multiple conformations fluctuating with millisecond to microsecond motions. Prot. Sci. 6, 534–542 (1997).
Tolman, J.R., Flanagan, J.M., Kennedy, M.A. & Prestegard, J.H. NMR evidence for slow collective motions in cyanometmyoglobin. Nature Struct. Biol. 4, 292–297 (1997).
Orekhov, V.Y., Pervushin, K.V. & Arseniev, A.S. Backbone dynamics of (1–71)bacterioopsin studied by two-dimensional 1H-15N NMR spectroscopy. Eur. J. Biochem. 219, 887–896 (1994).
Tjandra, N., Kuboniwa, H., Ren, H. & Bax, A. Rotational dynamics of calcium-free calmodulin studied by 15N-NMR relaxation measurements. Eur. J. Biochem. 230, 1014–1024 (1995).
Mandel, A.M., Akke, M. & Palmer, A.G. Dynamics of ribonuclease H: Temperature dependence of motion on multiple time scales. Biochemistry 35, 16009–16023 (1996).
Ogata, K. et al. The cavity in the hydrophobic core of Myb DNA-binding domain is reserved for DNA recognition and trans-activation. Nature Struct. Biol. 3, 178–187 (1996).
Olejniczak, E.T., Zhou, M.M. & Fesik, S.W. Changes in the NMR-derived motional parameters of the insulin receptor substrate I phosphotyrosine binding domain upon binding to an interleukin 4 receptor phosphopeptide. Biochemistry 36, 4118–4124 (1997).
Leahy, D., Hendrickson, W.A., Aukhil, I. & Erickson, H.P. Structure of a fibronectin Type III domain from tenascin phased by MAD analysis of the selenomethionyl protein. Science 258, 987–991 (1992).
Main, A.L., Harvey, T.S., Baron, M., Boyd, J. & Campbell, I.D. The three-dimensional structure of the tenth type III module of fibronectin: An insight into RGD-mediated interactions. Cell 71, 671–678 (1992).
Campbell, I.D. & Spitzfaden, C. Building proteins with fibronectin type III modules. Structure 2, 333–337 (1994).
Bork, P., Downing, A.K., Kieffer, B. & Campbell, I.D. Structure and distribution of modules in extracellular proteins. Q. Rev. Biophys. 28, 119–167 (1996).
Erickson, H.P., Tenascin-C, tenascin-R and tenascin-X: a family of talented proteins in search of functions. Curr. Opin. Cell Biol. 5, 869–876 (1993).
Chiquet-Ehrismann, R., Tenascins, a growing family of extracellular matrix proteins. Experientia 51, 853–862 (1995).
Scholze, A., Götz, B. & Faissner, A. Glial cell interactions with tenascin-C: adhesion and repulsion to different tenascin-C domains is cell type related. Int. J. Devl. Neuroscience 14, 315–329 (1996).
Clore, G.M., Driscoll, P.C., Wingfield, P.T. & Gronenborn, A.M. Analysis of the backbone dynamics of interleukin-1β using two-dimensional inverse detected heteronuclear 15N-1H NMR spectroscopy. Biochemistry 29, 7387–7401 (1990).
Kay, I.E., Torchia, D.A. & Bax, A. Backbone dynamics of proteins as studied by nitrogen-15 inverse detected heteronuclear NMR spectroscopy: application to staphylococcal nuclease. Biochemistry 28, 8972–8979 (1989).
Le, H. & Oldfield, E. Correlation between 15N NMR chemical shifts in proteins and secondary structure. J. Biomol. NMR 4, 341–348 (1994).
de Dios, A.C., Pearson, J.G. & Oldfield, E . Secondary and tertiary structural effects on protein NMR chemical shifts: An ab initio approach. Science 260, 1491–1496 (1993).
Brüschweiler, R. & Wright, P.E. NMR order parameters of biomolecules: A new analytical representation and application to the Gaussian axial fluctuation model. J. Am. Chem. Soc. 116, 8426–8427 (1994).
Pedersen, T.G., Thomsen, N.K., Andersen, K.V., Madsen, J.C. & Poulsen, F.M. Determination of the rate constants k1 and k2 of the Linderstrøm-Lang model for protein amide hydrogen exchange. A study of the individual amides in hen egg-white lysozyme. J. Mol. Biol. 230, 651–660 (1993).
Clarke, J., Hamill, S.J. & Johnson, C.M. Folding and stability of a fibronectin type III domain of human tenascin. J. Mol. Biol. 270, 771–778 (1997).
Wendt, B. et al. Effect of amino acid substitutions and deletions on the thermal stability, the pH stability and unfolding by urea of bovine calbindin D9k. Eur. J. Biochem. 175, 439–445 (1988).
Beeser, S.A., Goldenberg, D.P. & Oas, T.G. Enhanced protein flexibility caused by a destabilizing amino acid replacement in BPTI. J. Mol. Biol. 269, 154–164 (1997).
Carr, P.A., Erickson, H.P. & Palmer, A.G. Backbone dynamics of homologous fibronectin type III cell adhesion domains from fibronectin and tenascin. Structure 5, 949–959 (1997).
Palmer, A.G., Williams, J. & McDermott, A. Nuclear magnetic resonance studies of biopolymer dynamics. J. Phys. Chem. 100, 13293–13310 (1996).
Schurr, J.M., Babcock, H.P. & Fujimoto, B.S. A test of the model-free formulas.Effects of anisotropic rotational diffusion and dimerization. J. Magn. Reson., Ser. B 105, 211–224 (1994).
Tjandra, N., Wingfield, P., Stahl, S. & Bax, A. Anisotropic rotational diffusion of perdeuterated HIV protease from 15N NMR relaxation measurements at two magnetic fields. J. Biomol. NMR 8, 273–284 (1996).
Mandel, A.M. & Palmer, A.G. Measurement of relaxation rate constants using constant-time accordion heteronuclear NMR spectroscopy. J. Magn. Reson., Ser. A 110, 62–72 (1994).
Brüschweiler, R., Liao, X. & Wright, P.E. Long-range motional restrictions in a multidomain zinc-finger protein from anisotropic tumbling. Science 268, 886–889 (1995).
Lee, L.K., Ranee, M., Chazin, W.J. & Palmer, A.G. Rotational diffusion anisotropy of proteins from simultaneous analysis of 15N and 13Cα nuclear spin relaxation. J. Biomol. NMR 9, 287–298 (1997).
Akke, M., Carr, P.A. & Palmer, A.G. Heteronuclear-correlation NMR spectroscopy with simultaneous isotope filtration, quadrature detection, and sensitivity enhancement using z rotations. J. Magn. Reson., Ser. B104, 298–302 (1994).
Davis, D.G., Perlman, M.E. & London, R.E. Direct measurements of the dissociation-rate constant for inhibitor-enzyme complexes via the T1ρ and T2 (CPMG) methods. J. Magn. Reson., Ser. B 104, 266–275 (1994).
Press, W.H., Flannery, B.P., Teukolsky, S.A. & Vetterling, W.T., Numerical Recipes. The Art of Scientific Computing 1-963 (Cambridge University Press, Cambridge; 1986).
Fushman, D., Cahill, S. & Cowburn, D. The main-chain dynamics of the dynamin pleckstrin homology (PH) domain in solution: analysis of 15N relaxation with monomer/dimer equilibration. J. Mol. Biol. 266, 173–194 (1997).
Johnson, M.L., Correia, J.J., Yphantis, D.A. & Halvorson, H.R. Analysis of data from the analytical ultracentrifuge by nonlinear least-squares techniques. Biophys. J. 36, 575–588 (1981).
Eftink, M.R. Use of multiple spectroscopic methods to monitor equilibrium unfolding of proteins. Meth. Enz. 259, 487–512 (1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Akke, M., Liu, J., Cavanagh, J. et al. Pervasive conformational fluctuations on microsecond time scales in a fibronectin type III domain. Nat Struct Mol Biol 5, 55–59 (1998). https://doi.org/10.1038/nsb0198-55
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0198-55
This article is cited by
-
Simultaneous determination of fast and slow dynamics in molecules using extreme CPMG relaxation dispersion experiments
Journal of Biomolecular NMR (2018)
-
Enhanced accuracy of kinetic information from CT-CPMG experiments by transverse rotating-frame spectroscopy
Journal of Biomolecular NMR (2013)
-
Resolving Conformational and Rotameric Exchange in Spin-Labeled Proteins Using Saturation Recovery EPR
Applied Magnetic Resonance (2010)
-
The second decade — into the third millenium
Nature Structural Biology (1998)
-
Protein dynamics from NMR
Nature Structural Biology (1998)