Abstract
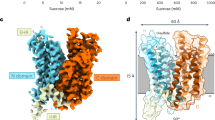
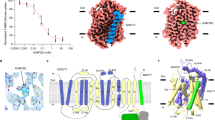
The X-ray structure of a sucrose-specific porin (ScrY) from Salmonella typhimurium has been determined by multiple isomorphous replacement at 2.4 Å resolution both in its uncomplexed form and with bound sucrose. ScrY is a noncrystallographic trimer of identical subunits, each with 413 structurally well-defined amino acids. A monomer is built up of 18 anti-parallel β-strands surrounding a hydrophilic pore, with a topology closely similar to that of maltoporin. Two non-overlapping sucrose-binding sites were identified in difference Fourier maps. The higher permeability for sucrose of ScrY as compared to maltoporin is mainly accounted for by differences in their pore-lining residues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lugtenberg, B. & Van Alphen, L. Molecular architecture and functioning of the outer membrane of Escherichia coli and other gram-negative bacteria. Biochim. Biophys. Acta 737, 51–115 (1983).
Bayer, M.E. & Bayer, M.H. in Bacterial Cell Wall (eds Ghuysen, J.-M. & Hakenbeck, R.) 447–462 (Elsevier Sci. B.V., Amsterdam; 1994).
Riley, M. & Labedan, B. in Escherichia coli and Salmonella (ed. Neidhardt, F.C.) 2118–2202 (ASM Press, Washington, D.C.; 1996).
Boos, W. & Lucht, J.M. in Escherichia coli and Salmonella (ed. Neidhardt, F.C.) 1175–1209 (ASM Press, Washington, D.C.; 1996).
Postma, P.W., Lengeler, J. & Jacobson, G.R. in Escherichia coli and Salmonella (ed. Neidhardt, F.C.) 1149–1174 (ASM Press, Washington, D.C.; 1996).
Benz, R. & Bauer, K. Permeation of hydrophilic molecules through the outer membrane of gram-negative bacteria. Eur.J.Biochem. 176, 1–19 (1988).
Welte, W., Nestel, U., Wacker, T. & Diederichs, K. Structure and function of the porin channel. Kidney Int. 48, 930–940 (1995).
Delcour, A.H., Adler, J., Kung, C. & Martinac, B. Membrane-derived oligosaccha-rides (MDO's) promote closing of an E. coli porin channel. FEBS Lett. 304, 216–220 (1992).
Welte, W., Diederichs, K., Przybylski, M., Glocker, M., Benz, R. & Breed, J. X-ray crystallographic and mass spectrometric structure determination and functional characterization of succinylated porin from R. capsulatus : Implications for ion selectivity and single-channel conductance. In Proceedings of the NATO Advanced workshop “New Methods for the Study of Molecular Aggregates” (eds Standing, K. & Ens, W.) in the press.
Weiss, M.S. et al. Molecular architecture and electrostatic properties of a bacterial porin. Science 254, 1627–1630 (1991).
Cowan, S.W. et al. Crystal structures explain functional properties of two E. coli porins. Nature 358, 727–733 (1992).
Kreusch, A., Neubüser, A., Schiltz, E., Weckesser, J. & Schulz, G.E. Structure of the membrane channel porin from Rhodopseudomonas blastica at 2.0 Å resolution. Protein Sci. 3, 58–63 (1994).
Hirsch, A., Diederichs, K., Breed, J., Saxena, K., Richter, O.-M., Ludwig, B. & Welte, W. The structure of porin from Paracoccus denitrificans at 3.1 Å resolution. FEBS Lett. 404, 208–210 (1997).
Eisenberg, G. & Dani, J.A. An Introduction to Molecular Architecture and Permeability of Ion Channels. Ann. Rev. Biophys. Biophys. Chem. 16, 205–226 (1987).
Glasstone, S., Laidler, K.J. & Eyring, H. The Theory of Rate Processes (McGraw Hill Book Comp., New York and London; 1941).
Frauenfelder, H. & Wolynes, P.G. Rate Theories and Puzzles of Hemeprotein Kinetics. Science 229, 337–345 (1985).
Rosenbusch, J.P. Characterization of the major envelope protein from Escherichia coli. J. Biol. Chem. 249, 8019–8029 (1974).
Birge, E. Bacterial and Bacteriophage Genetics (Springer Verlag Berlin; 1981).
Death, A., Notley, L. & Ferenci, T. Derepression of LamB protein facilitates outer membrane permeation of carbohydrates into Escherichia coli under conditions of nutrient stress. J. Bacteriol. 175, 1475–1483 (1993).
Benz, R., Schmid, A. & Vos-Scheperkeuter, G.H. Mechanism of sugar transport through the sugar-specific LamB channel of Escherichia coli outer membrane. J. Membrane Biol. 100, 21–29 (1987).
Schirmer, T., Keller, T.A., Wang, Y. & Rosenbusch, J.P. Structural basis for sugar translocation through maltoporin channels at 3.1 Å resolution. Science 267, 512–514 (1995).
Dutzler, R., Wang, Y.-F., Rizkallah, P.J., Rosenbusch, J.P. & Schirmer, T. Crystal structures of the various maltooligosaccharides bound to maltoporin reveal a specific sugar translocation pathway. Structure 4, 127–134 (1996).
Wang, Y.-F., Dutzler, R., Rizkallah, P.J., Rosenbusch, J.P. & Schirmer, T. Channel specificity: Structural basis for sugar discrimination and differential flux rates in maltoporin. J. Mol. Biol. 272, 56–63 (1997).
Meyer, J.E., Hofnung, M. & Schulz, G.E. Structure of maltoporin from Salmonella typhimurium ligated with a nitrophenyl-maltotrioside. J. Mol. Biol. 266, 761–75 (1997).
Schmid, K., Schupfner, M. & Schmitt, R. Plasmid-mediated uptake and metabolism of sucrose by Escherichia coli K-12. J. Bacteriol. 151, 68–76 (1982).
Schmid, K., Ebner, R., Jahreis, K., Lengeler, J.W. & Titgemeyer, F. A sugar-specific porin, ScrY, is involved in sucrose uptake in enteric bacteria. Mol. Microbiol. 5, 941–950 (1991).
Hardesty, C., Ferran, C. & DiRienzo, J.M. Plasmid-mediated sucrose metabolism in Escherichia coli: Characterization of scrY, the structural gene for a phosphoenol-pyruvate-dependent sucrose phosphotransferase system outer membrane porin. J. Bacteriol. 173, 449–456 (1991).
Schülein, K., Andersen, C. & Benz, R. The deletion of 70 amino acids near the N-terminal end of the sucrose-specific porin ScrY causes its functional similarity to LamB in vivo and in vitro. Mol. Microbiol. 17, 757–767 (1995).
Forst, D. et al. Crystallization and preliminary X-ray diffraction analysis of ScrY, a specific bacterial outer membrane porin. J. Mol. Biol. 229, 258–262 (1993).
Burley, S.K. & Petsko, G.A. Electrostatic interactions in aromatic oligopeptides contribute to protein stability. TIBTECH 7, 354–359 (1989).
Pebay-Peyroula, E., Garavito, R.M., Rosenbusch, J.P., Zulauf, M. & Timmins, P.A. Detergent structure in tetragonal crystals of OmpF porin. Structure 3, 1051–1059 (1995).
Schiffer, M., Chang, C.H. & Stevens, F.J. Transport proteins in bacteria: Common themes in their design. Science 258, 936–942 (1992).
Vyas, N.K. Atomic features of protein-carbohydrate interactions. Curr. Op. Struct. Biol. 1, 732–740 (1991).
Engelsen, S.B., du Penhoat, C.H. & Perez, S. Molecular Relaxation of Sucrose in aqueous Solution. J. Phys. Chem. 99, 13334–13351 (1995).
Brown, G.M. & Levy, H.A. Further Refinement of the Structure of Sucrose based on Neutron-Diffraction Data. Acta Crystallogr. B29, 790–797 (1973).
Immel, S. & Lichtenthaler, F.W. The Conformation of Sucrose in water: A Molecular Dynamics Approach. Liebigs Ann. 1995, 1925–1937 (1995).
Casset, F. et al. NMR, Molecular Modelling, and Crystallographic Studies of Lentil Lectin-Sucrose Interaction. J. Biol. Chem. 270, 25619–25628 (1995).
Tormo, J. et al. Crystal structure of a bacterial family-III cellulose-binding domain: a general mechanism for attachment to cellulose. EMBO J. 15, 5739–5751 (1996).
O'Reilly, M., Watson, K.A., Schinzel, R., Palm, D. & Johnson, L.N. Oligosaccharide substrate binding in Escherichia coli maltodextrin phosphorylase. Nature Struct. Biol. 4, 405–412 (1997).
Davies, G.J. et al. Structure Determination and Refinement of the Humicola inso-lens Endoglucanase V at 1.5 Å Resolution. Acta Crystallogr. D52, 7–17 (1996).
Ford, L.O., Johnson, L.N., Machin, P.A., Philips, D.C. & Tijan, R., Crystal Structure of a Lysozyme-Tetrasaccharide Lactone Complex. J. Mol. Biol. 88, 349–371 (1974).
Ng, K.K.-S., Drickamer, K. & Weis, W.I. Structural Analysis of Monosaccharide Recognition by Rat Liver Mannose-binding Protein. J. Biol. Chem. 271, 663–674 (1996).
Weis, W.I., Drickamer, K. & Hendrickson, W.A. Stucture of a C-type mannose-binding protein complexed with an oligosaccharide. Nature 360, 127–134 (1992).
Qian, M., Haser, R. & Payan, F. Carbohydrate binding sites in a pancreatic α-amylase-substrate complex, derived from X-ray structure analysis at 2.1 Å resolution. Protein Sci. 4, 747–755 (1995).
Shaanan, B., Lis, H. & Sharon, N. Structure of a Legume Lectin with an Ordered N-linked Carbohydrate in Complex with Lactose. Science 254, 862–866 (1991).
Francis, G., Brennan, L., Stretton, S. & Ferenci, T. Genetic mapping of starch- and lambda-receptor sites in maltoporin: identification of substitutions causing direct and indirect effects on binding sites by cysteine mutagenesis. Mol. Microbiol. 5, 2293–2301 (1991).
Benz, R., Francis, G., Nakae, T. & Ferenci, T. Investigation of the selectivity of maltoporin channels using LamB proteins: Mutations changing the maltodextrin binding site. Biochim. Biophys. Acta 1104, 299–307 (1992).
Meyer, J.E.W. & Schulz, G.E. Energy profile of maltooligosaccharide permeation through maltoporin as derived from the structure and from a statistical analysis of saccharide-protein interactions. Protein Sci. 6, 1084–1091 (1997).
Jordy, M., Andersen, C., Schülein, K., Ferenci, T. & Benz, R., Constants of Sugar Transport Through Two LamB Mutants of Escherichia coli : Comparison with Wild-type Maltoporin and LamB of Salmonella typhimurium. J. Mol. Biol. 259, 666–678 (1996).
Adam, G. & Delbrück, M. : Reduction of Dimensionality in Biological Diffusion Processes. in : Structural Chemistry & Molecular Biology 198–215 (eds Rich, A. & Davidson, N.) (Freeman, San Francisco; 1968).
Lupas, A. Coiled coils: New structures and new functions. TIBS 21, 375–382 (1996).
DeLano, W.L. & Brünger, A.T. Helix packing in Proteins: prediction and energetic analysis of dimeric, trimeric, and tetrameric GCN4 coiled coil structures. Proteins: Struct. Funct. Genet. 20, 105–123 (1994).
Engel, A.M., Cejka, Z., Lupas, A., Lottspeich, F. & Baumeister, W. Isolation and Cloning of Ompα, a coiled-coil protein spanning the periplasmic space of the ancestral eubacterium Thermotoga maritima. EMBO J. 11, 4369–4378 (1992).
Derouiche, R. et al. TolA central domain interacts with E. coli porins. EMBO J. 15, 6408–6415 (1996).
McLachlan, A.D. Coiled-Coil Structure of Murein Lipoprotein. Biochem. Soc. Transact. 6, 1353–1354 (1978).
Braun, V., Rotering, H., Ohms, J.-P. & Hagenmaeier, H. Conformational studies on Murein-Lipoprotein from the Outer Membrane of Escherichia coli. Eur. J Biochem. 70, 601–610 (1976).
Krylov, D., Mikhailenko, I. & Vinson, C. A thermodynamic scale for leucine zipper stability and dimerization specificity: e and g interhelical interactions. EMBO J. 13, 2849–2861 (1994).
Lavigne, P., Sönnichsen, F.D., Kay, C.M. & Hodges, R.S. Interhelical Salt Bridges, Coiled-Coil Stability, and Specificity of Dimerization. Science 271, 1136–1137 (1996).
Yang, A.S. & Honig, B. Electrostatic effects on protein stability. Curr. Op. Struct. Biol. 2, 40–45 (1992).
Lumb, K.J. & Kim, P.S. Measurement of Interhelical Electrostatic Interactions in the GCN4 Leucine Zipper. Science 268, 436–439 (1995).
Schülein, K., Schmid, K. & Benz, R. The sugar-specific outer membrane channel ScrY contains functional characteristics of general diffusion pores and substrate-specific porins. Mol. Microbiol. 5, 2233–2241 (1991).
Kabsch, W. Automatic Processing of Rotation Diffraction Data from Crystals of Initially Unknown Symmetry and Cell Constants. J. Appl. Cryst. 26, 795–800 (1993).
Diederichs, K. A comparison of some heavy-atom refinement and phasing programs. CCP4/ESF-EACBM Newsletters on Protein Crystallography 31, 23–30 (1994).
Kleywegt, G.J. Making the most of your search model. CCP4/ESF-EACBM Newsletter on Protein Crystallography 32, 32–36 (1996).
Collaborative Computational Project, Number 4 The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D50, 760–763 (1994).
Cowtan, K. ‘dm’: An Automated Procedure for Phase Improvement by Density Modification. CCP4/ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A.T. (1992) X-PLOR Version 3.1. A System for X-ray crystallography and NMR (Yale University Press, New Haven; 1992).
Brünger, A.T., Krukowski, A. & Erickson, J. Slow cooling-protocols for crystallographic refinement by simulated annealing. Acta Crystallogr. A46, 585–593 (1990).
Yeates, T.O. Simple statistics for intensity data from twinned specimens. Acta Crystallogr. A44, 142–144 (1988).
Gomis-Rüth, F.X. et al. Determination of Hemihedral Twinning and Initial Structural Analysis of Crystals of the Procarboxypeptidase A Ternary Complex. Acta Crystallogr. D51, 819–823 (1995).
Sheldrick, G.M. & Schneider, T.R. SHELXL: High-Resolution Refinement. Meth. Enz. 277, 319–343 (1997).
McLachlan, A.D. Gene duplication in the structural evolution of chymotrypsin. J. Mol. Biol. 128, 49–79 (1979).
Carson, M. Ribbon Models of Macromolecules, J. Mol. Graphics 5, 103–106 (1987).
Kraulis, P. MOLSCRIPT: A program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merrit, E.A. & Bacon, D.J. Raster3D: Photorealistic Molecular Graphics. Meth. Enz. 277, 505–524 (1997).
Kabsch, W. & Sanders, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Nicholls, A., Bharadwaj, R. & Honig, B. GRASP Graphical Representation and Analysis of Surface Properties. Biophys. J. 64, A166 (1993).
Wallace, A.C., Laskowski, R.A. & Thornton, J.M. LIGPLOT: A program to generate schematic diagrams of protein-ligand interactions. Protein Engng. 8, 127–134 (1995).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Forst, D., Welte, W., Wacker, T. et al. Structure of the sucrose-specific porin ScrY from Salmonella typhimurium and its complex with sucrose. Nat Struct Mol Biol 5, 37–46 (1998). https://doi.org/10.1038/nsb0198-37
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0198-37
This article is cited by
-
Binding of HasA by its transmembrane receptor HasR follows a conformational funnel mechanism
European Biophysics Journal (2020)
-
Statistical prediction of protein structural, localization and functional properties by the analysis of its fragment mass distributions after proteolytic cleavage
Scientific Reports (2016)
-
Correlated motions are a fundamental property of β-sheets
Nature Communications (2014)